Abstract
Boric acid (BA) has antimicrobial properties and is used to combat bacterial infections, including Enterobacteria. However, the molecular mechanisms and cellular responses to BA are still unknown. This genomics study aims to provide new information on the genes and molecular mechanisms related to the antimicrobial effect of BA in Escherichia coli. The Keio collection of E. coli was used to screen 3985 single-gene knockout strains in order to identify mutant strains that were sensitive or hypersensitive to BA at certain concentrations. The mutant strains were exposed to different concentrations of BA ranging from 0 to 120 mM in LB media. Through genome-wide screens, 92 mutants were identified that were relatively sensitive to BA at least at one concentration tested. The related biological processes in the particular cellular system were listed. This study demonstrates that intrinsic BA resistance is the result of various mechanisms acting together. Additionally, we identified eighteen out of ninety-two mutant strains (Delta_aceF, aroK, cheZ, dinJ, galS, garP, glxK, nohA, talB, torR, trmU, trpR, yddE, yfeS, ygaV, ylaC, yoaC, yohN) that exhibited sensitivity using other methods. To increase sensitivity to BA, we constructed double and triple knockout mutants of the selected sensitive mutants. In certain instances, engineered double and triple mutants exhibited significantly amplified effects. Overall, our analysis of these findings offers further understanding of the mechanisms behind BA toxicity and intrinsic resistance in E. coli.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The molecular mechanisms underlying boron toxicity, as well as the physiological causes and cellular processes involved, are not yet well understood. Boron toxicity has been linked to the chemical properties of boron, which tightly binds to the cis-diol groups of certain biomolecules, including ribose, Ado-Met, and NADH [1, 2]. Boron requirement and toxicity are intriguing because although some living organisms require or benefit from a small amount of boron, it becomes toxic above a certain level [3, 4]. Therefore, in many parts of the world where the boron level is high, the risk of boron toxicity for living organisms is a concern [5]. Boron is also used as a food preservative, insecticide, and treatment for vaginal diseases caused by Saccharomyces and Candida [6, 7]. Boron has significant effects on a variety of organisms. One of these effects is its role as a micronutrient in plant growth, as demonstrated by Warrington in 1923 [8]. Furthermore, boron is vital for plant structure and cell walls, making it an essential element [9]. It has also been reported to be essential for some species of Cyanobacteria and Bacillus boroniphilus [10, 11].
Studies have shown that boron may contribute to promoting certain metabolic activities, as indicated by various investigations [12,13,14]. Several important studies conducted with yeast have demonstrated that a membrane protein actively transports boron [15]. Furthermore, boron has been shown to be necessary for certain animals and single-cell eukaryotes [16, 17]. The essentiality of boron for certain organisms has been increasingly considered over time. The study found that plants with mutations in NIP5:1 are more susceptible to boron deficiency in root and branch development due to the lack of boron uptake [18]. Furthermore, it has been suggested that neutral boric acid [B(OH)3] can passively diffuse in addition to facilitating acid transport into the cell through the NIP5:1 protein concentration gradient along the plasma membrane [19]. Although there have been discussions on the existence of regulatory mechanisms in bacteria regarding intracellular boron levels, no experimental studies have been conducted yet. Only a few suggestions have been made regarding the variation of intracellular boron levels among different species [20]. To date, no genes or proteins related to boron resistance or efflux functions have been identified in the bacterial world.
Bacteria adapt to environmental changes or extreme conditions through the regulation of their genes, proteins, and metabolites. Understanding the dynamics of this regulatory network is a key goal of biology. Escherichia coli is a commonly used model organism in microbiological studies and is one of the most well-characterized organisms in this field [21]. The K12 strains E. coli MG1655 [22] and E. coli W3110 [23] have been extensively used for genomic studies. We utilized the Keio mutant collection, an important example of new generation genomic research in E. coli [24]. This mutant collection can be used to study the effects of any chemical or agent that can kill a bacterium at a certain dose. In this study, we screened the Keio mutant line to identify the genes conferring intrinsic resistance to boric acid (BA) in E. coli. The roles of some genes in intrinsic BA resistance were discovered and discussed.
Materials and Methods
Escherichia coli Strains and Plasmids
The Keio mutant line, derived from the starting E. coli strain BW25113 [25], was used in this study [24]. For complementation purposes, pCA24N plasmids containing the target genes were employed [26]. An empty pCA24N plasmid, which contains a chloramphenicol resistance (CmR), was used as a control. The open reading frame (ORF) regions cloned into this vector are transcribed by the T5-lac promoter, which is inducible by isopropyl-β-D-thiogalactoside (IPTG) [26]. The plasmids and strains used in this study are listed in Table 1.
Genome-Wide Screening of Intrinsic BA Resistance-Associated Genes with the Replicator
The Keio mutant strains are stored in 96 deep well plates with 15% glycerol at − 80 °C. They were then transferred to 96 deep well plates containing 1 ml of Luria Bertani (LB) medium per well using a cryoreplicator consisting of 96 pins. The plates were incubated at 37 °C. Next, all strains were transferred with the replicator to deep well plates containing 50 μg/ml of kanamycin in LB medium. After incubation for 3 h, the strains were plated on LB agar medium with varying BA concentrations (0, 25, 50, 80, 100, 120 mM). The mutant strains were then incubated for 3 days to grow.
Sequential Spotting
The mutant strains that were identified as sensitive to BA during screening studies underwent a sequential spot test. A single colony was inoculated into LB agar containing 50 μg/ml kanamycin without BA from the mutant strain’s stock. After confirming colony purity, the mutants were grown in LB at 37 °C until they reached a density of OD600 0.5. They were then serially diluted (1/2, 1/4, 1/8, and 1/16). The reproductive status of the mutants was assessed by spotting 5-μl cultures onto LB agar medium with varying concentrations of BA. As the concentration of BA increased, it became toxic to the bacterium, leading to reduced bacterial growth. The plates were incubated for 72 h at 37 °C, and bacterial growth was recorded in tabular form.
Determination of MIC
To determine the minimum inhibitory concentration (MIC) of the mutants [27], single colony cultures were grown on LB agar. The mutants were then inoculated onto LB agar medium containing 50 μg/ml kanamycin. After overnight growth, a colony was selected and inoculated into 5 ml of LB liquid medium with kanamycin (50 μg/ml) and left to incubate overnight at 37 °C. Sterile microplates were used to aliquot LB broth medium containing 50 μg/ml kanamycin and different concentrations of BA. The mutant strains, which were sensitive, were inoculated into a broth medium with an initial OD600 value of 0.05 and incubated at 37 °C. Daily monitoring was conducted to observe growth, and the MIC value was determined for each strain.
Complementation of Sensitive Mutant Strains
To obtain a sufficient concentration of the recombinant plasmid vector containing the target gene (pCA24N::target gene), E. coli strains carrying the plasmids were cultivated and then the recombinant vectors carrying the target gene were isolated using a plasmid isolation kit (GeneAll 101–150). Then, the recombinant plasmid was transformed using the heat shock method into the particular mutant (E. coli BW25113::Δgene) [28]. The susceptibility of the cells that regained the gene was reassessed by growing them in LB broth with varying BA concentrations, 100 μM IPTG, and 25 μg/ml chloramphenicol. The assessment was conducted using the MIC test described above.
BA Sensitivity Analysis of Double and Triple Mutants
In the BW25113 background, we generated double and triple mutants via P1 transduction [29]. The aim of the study was to observe the mutants and determine if their susceptibility to BA is increased by the knockout of identified genes. The protocols published by Silhavy et al. (1984) were followed [30]. To generate a double mutant, the kanamycin gene cassette was deleted through FLP recombination using plasmid pCP20. Kanamycin-resistant (KmR) transductants were obtained after overnight incubation at 37 °C. We obtained double and triple mutants and confirmed the absence of the target genes using PCR.
Data Mining, Analysis, and Interpretation
The activities of knockout genes in susceptible mutant strains were investigated using the EcoCyc at http://ecocyc.org/ [31] and UniProt databases, and the corresponding UniProt IDs of the proteins were obtained (https://www.uniprot.org/). The knockout genes of sensitive mutant strains were classified according to their systems and subsystems using the EcoCyc Database and Omics Dashboard (Pathway Tools, https://ecocyc.org/dashboard/dashboard-intro.shtml?orgid=ECOLI) [31]. Enrichment of the selected gene hits was carried out using the DAVID database (https://david.ncifcrf.gov/tools.jsp) [32, 33] with an FDR of 0.1 and a cut-off score of 2 SD from the mean. The STRING database (https://string-db.org/) [34] was also used to explore the direct or indirect relationships between the knockout genes of the identified susceptible mutant strains.
Results
Genome-Wide Screening of the Mutants for BA Sensitivity
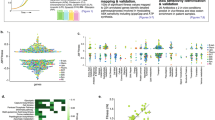
A total of 3985 mutants from the Keio collection were screened in triplicate across six different BA concentrations, generating an initial dataset of nearly 71,730 data points (23,910 × 3). An example from one microplate is shown in Fig. 1. The mutants that showed reduced or no growth at specific BA concentrations were recorded. Out of 3985 mutants, 92 sensitive mutants were determined after comparing them to the wild type and surrounding mutants. The table presents gene information and the concentrations of BA (in mM) that were tested for the strains. The dark gray color indicates good growth, while the color scale that fades towards white represents a decrease in growth, with white indicating no growth. “Wt” denotes the control strain (wild type), while the other strains are the sensitive mutants identified by their 4-letter gene names in Table 2.
Effect of 50 mM BA on K-12 BW25113ΔtorR ΩKmR mutant E. coli strain. LB plates without any BA (left panel) and LB plates with 50 mM BA (right panel) were used to culture 96 mutants from the plate in Keio collection. Among them, ΔtorR mutant failed to grow on the plate with 50 mM BA. Further investigation on pure colonies confirmed its hypersensitivity
The mutants, identified through screening studies, were categorized into three sensitivities: highly sensitive (HS), moderately sensitive (MS), and low sensitive (LS). The mutants’ responses to different concentrations of BA determined their sensitivity levels. Mutants that ceased reproduction or exhibited limited growth at 25 mM were classified as highly sensitive (HS), at 50 mM as moderately sensitive (MS), and at 80 mM as low sensitive (LS) (Table 2).
The study found that the absence of certain genes caused the BA-sensitive phenotype. A decrease in colony size or lack of reproduction of mutants indicates BA sensitivity. The study identified 92 genes associated with BA sensitivity (Table 2). These genes were classified based on their activity using Pathway Tools and the EcoCyc databases. It was observed that one gene was involved in more than one system or subsystem (Table 3).
Assessment of BA Sensitivity and Resistance Using Sequential Spotting and MIC Tests
Ninety-two mutants underwent spot tests to determine their sensitivity to BA compared to the wild-type strain. The results confirmed that 18 mutants, namely ΔaceF, ΔtorR, ΔdinJ, ΔtrmU, ΔcheZ, ΔylaC, ΔyoaC, ΔyohN, ΔglxK, ΔyfeS, ΔaroK, ΔygaV, ΔgalS, ΔtrpR, ΔtalB, ΔgarP, ΔnohA, and ΔyddE, exhibited relatively higher sensitivity (Fig. 2).
Spot tests to determine the BA sensitivity of the 18 mutant strains. The mutants were subjected to sequential spotting under different concentrations of BA, as described in the “Materials and Methods” section. Five different dilutions of the bacterial culture (1/1, 1/2, 1/4, 1/8, 1/16) and 13 different BA concentrations were used. The growth of the mutants and the wild-type strain is illustrated, with the particular strain name listed below each picture
MIC (minimum inhibitory concentration) values were determined using the microtube dilution method to compare the resistance levels of the mutant strains in solid and liquid media. The MIC test for the mutants was conducted in LB broth medium containing various BA concentrations (0, 25, 30, 40, 45, 50, 60, 65, 70, 80, 90, 100 mM). The reproductive status was monitored daily. Table 4 presents the MIC values of the 18 sensitive mutants to BA and the control strain. The control strain had a MIC value of 100 mM BA, while the mutants had reduced MIC values ranging from 40 to 80 mM BA, confirming their sensitivity or hypersensitivity (Table 4).
Complementation of BA-Sensitive Mutant Strains
To conduct the complementation experiments, we transformed the target gene into the corresponding mutants listed in Table 1 using the recombinant plasmid. For each mutant, we also prepared controls consisting of cells containing the empty plasmid pCA24N without any gene insert. We assessed the sensitivity to BA using the MIC test and monitored reproductive status daily. The complementation experiments revealed that the mutants ΔglxK, ΔcheZ, ΔtalB, ΔgalS, and ΔnohA did not exhibit complementation. However, it was demonstrated that the mutants ΔtrmU, ΔaceF, ΔtorR, ΔdinJ, ΔylaC, ΔyoaC, ΔyohN, ΔyfeS, ΔygaV, ΔtrpR, ΔgarP, and ΔyddE regained their genes and resistance levels to BA. This confirms that the BA-sensitive phenotype of the mutant is attributed to the specific gene of interest (Table 5).
Testing the Sensitivity of Double and Triple Knockout Mutants to BA
Double and triple mutants were generated using P1 transductions. The mutants were assessed for their sensitivity to BA, and it was observed that some mutants displayed more pronounced effects. The results are presented in Table 6. The double and triple mutants obtained exhibited significantly higher sensitivity compared to the individual single mutants. For example, the garP-ylaC and yoaC-yfeS double mutants demonstrated a sensitivity of approximately 50 mM, while each parent mutant exhibited a sensitivity ranging from 50 to 80 mM. Moreover, the generated triple mutants displayed even greater sensitivity, reaching down to 25 mM BA, compared to the individual mutants, as observed in the cases of ΔyoaCgarPylaC and ΔgarPyoaCyfeS. These results confirm the BA sensitivity of the specific mutants. Additionally, we suggest that these sensitive triple mutants can be employed in other genomic library selection studies, using other BA-tolerant bacteria, to identify more candidate genes related to the effect of BA on bacteria.
Discussion
BA-Sensitive Mutants of E. coli Were Identified Through Genome-Wide Screens, Spot, and MIC Tests
Boron is beneficial to living organisms in small amounts but can become toxic beyond a certain threshold. Additionally, boron exhibits antibacterial activity, although the molecular mechanisms behind this are unknown. Several publications have reported on the function of genes related to boron resistance and transport in yeast and plants [15, 35,36,37]. However, the molecular mechanisms underlying boron resistance or sensitivity in other organisms and bacteria remain unclear.
Advancements in genomics and transcriptomics have facilitated the utilization of functional genomic-based approaches to comprehend the global metabolic changes resulting from genotypic and/or environmental variations [38,39,40,41,42]. Our study employed a functional genomics approach to investigate the impact of BA on E. coli. A total of 3985 mutants were screened for sensitivity in the presence of increasing levels of BA.
The objective of this study was to investigate the genes and mechanisms responsible for intrinsic BA resistance in E. coli, a model bacterium. To achieve this, the Keio mutant line, consisting of single mutants of nonessential genes belonging to E. coli, was used [24]. The genome-wide screening of the mutants identified 92 knockout strains that showed increased susceptibility to BA, revealing genes involved in intrinsic BA resistance (Table 2). In addition, sequential spot tests and MIC experiments were conducted on these 92 mutants to determine their sensitivity to BA. The results showed that 18 of the mutants were particularly sensitive under all the methods tested (Fig. 2). The mutants’ sensitivity to BA ranged from most hypersensitive to sensitive in the following order: ΔyoaC > ΔtorR > ΔglxK > ΔylaC > ΔaroK > ΔyfeS > ΔaceF > ΔdinJ > ΔcheZ > ΔtrmU > ΔygaV > ΔyohN > ΔgalS > ΔtalB > ΔtrpR > ΔyddE > ΔnohA > ΔgarP. These mutants were found to be sensitive to BA following genome-wide screening, sequential spot, and MIC experiments.
Complementation Studies and Generation of Triple Mutants Confirmed the BA-Sensitive Phenotype of the Particular Mutants
Complementation experiments were performed to investigate whether the sensitivity to BA was due to the knockout gene. The mutant strains, along with proper control strains, were subject to BA spot tests after transformation with the corresponding gene cloned into an expression plasmid. It was demonstrated that the mutants ΔtrmU, ΔaceF, ΔtorR, ΔdinJ, ΔylaC, ΔyoaC, ΔyohN, ΔyfeS, ΔygaV, ΔtrpR, ΔgarP, and ΔyddE were complemented with their respective genes (Table 5). This confirms that the BA-sensitive phenotype of each mutant is due to the deleted gene. Further studies with these mutants are recommended.
To elaborate on the particular genes’ effect on intrinsic BA resistance, we generated double and triple knockouts to test for increased BA sensitivity. We created double mutants of ΔgarPylaC and ΔyoaCyfeS, and from these, we generated three triple mutants: ΔylaCgarPyoaC, ΔylaCgarPyfeS, and ΔyfeSyoaCgarP (Table 6). The mutants with the yoaC knockout exhibited the most sensitive phenotype to BA. The single, double, and triple mutants, where yoaC is deleted, had BA sensitivity levels of 45 mM, 40 mM, and 25 mM, respectively. YoaC is particularly noteworthy for its effect on intrinsic BA resistance in E. coli. Further research is advised on this gene with respect to BA response. There is no information available about yoaC in the literature, and the function of this gene is unknown. Investigating the involvement of YoaC in BA stress could be a good starting point to elucidate the physiological function of YoaC in E. coli and other bacteria.
Pathway and Network Analyses of the Genes of the Sensitive Mutants Provide Insights into Intrinsic Resistance to BA
It is important to note that different culturing conditions may yield different results, which could lead to the addition or omission of some genes specifically related to the studied process. With these limitations in mind, we discuss the corresponding genes in BA-sensitive mutants in terms of the affected systems. Our focus is primarily on the systems expected to exhibit high sensitivity, along with any other significant findings and noteworthy genes that did not display a response. To gain a broader perspective, we analyzed all 92 genes using Omics Dashboard and String approaches, with particular attention paid to the 18 genes in detail.
Using Pathway Tools and the EcoCyc database, we mapped several gene hits to multiple systems and subsystems (Table 3 and Fig. 3). Our findings suggest that various cellular mechanisms, along with the corresponding physiological responses of E. coli, are involved in BA toxicity and resistance. This conclusion is supported by the identification of genes across multiple cellular systems, as shown in Table 3.
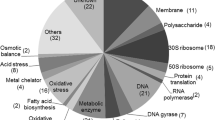
Strain numbers of subsystems. Each gene hit, which is associated with a BA-sensitive mutant strain, is mapped to specific cellular processes. It is important to note that gene hits may be associated with more than one process. The gene list was evaluated using Omics Dashboard (Pathway Tools) and the EcoCyc databases
Out of the 92 BA-sensitive gene hits, a significant number were assigned to various cellular processes, including “Regulation,” such as sigma factor regulons, signal transduction pathways, transcription factors, and transcription factor regulons. Hits were observed in various categories, including “Cell exterior,” “Cell Wall Biogenesis-Organization Proteins,” “Flagellar Proteins,” “Lipopolysaccharide Metabolism Proteins,” “Outer Membrane Proteins,” “Periplasmic Proteins,” “Pilus Proteins,” “Plasma Membrane Proteins,” and “Transport Proteins.” Additionally, hits were found in the “Central dogma” category, which covers DNA Metabolism, Protein Metabolism, RNA Metabolism, Transcription Proteins, and Translation Proteins. The “Response to stimulus” category includes hits related to Detoxification Proteins, Other Proteins involved in Stimulus Response, Proteins Involved in Response to Cold, Proteins Involved in Response to DNA Damage, Proteins Involved in Response to Osmotic Stress, Proteins Involved in Response to Starvation, Proteins Involved in Response to pH, and others.
Overall, we observed 62, 34, 26, and 19 sensitive hits in the categories of “Regulation,” “Cell exterior,” “Central dogma,” and “Response to stimulus,” respectively (Fig. 3). These findings suggest that BA has an impact on several cellular processes to varying degrees.
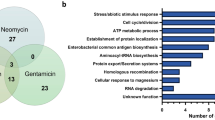
Enrichment analysis was performed on the 92 BA-sensitive gene hits using the DAVID pathway database. The analysis revealed that proteins related to exoribonuclease II activity were the most significantly affected among the BA-sensitive gene mutants, with an enrichment score of over 50. Additionally, highly affected protein groups include those associated with RNA metabolism and the regulation of chemotaxis, with fold enrichment values above 30 and 20, respectively (Fig. 4). Mutants lacking genes in RNA processing exhibit sensitivity to BA, suggesting that BA may target the processing and maturation of tRNAs and possibly other RNAs. Further research is suggested to investigate the effect of BA on bacteria in relation to RNA.
Enrichment of the corresponding genes in BA-sensitive mutants g. Clusters with three or more genes were considered (p value < 0.05). The DAVID gene functional classification (version 6.8) database was employed to assess the enrichment using E. coli K-12 genome. Only the particular processes with three or more genes were included
“Central Dogma and Cell Exterior Proteins”
Twenty-six mutants, classified in the “Central Dogma,” exhibited sensitivity to BA (Table 3). For instance, trmU (mnmA) is a gene involved in RNA metabolism. TrmU catalyzes the formation of the 2-thiouridine modification [43, 44] in the nucleoside 5-methylamino-methyl-2-thiourridine (mnm5s2U34) found in tRNAGln, tRNALys, and tRNAGlu. These modifications regulate selective mRNA translation and have an impact on protein expression. Several studies have identified factors involved in mRNA splicing processes. One such study found that BA causes a reversible inhibition effect in pre-mRNA splicing [45], suggesting that BA interacts with RNA in the cell.
Our study identified 34 genes associated with the “Cell exterior” system (Fig. 2). One of these genes, YlaC, is an inner membrane protein (UniProt P0AASO) [46] that has been found to be highly sensitive to BA. Another gene, the yohN (rcnB) gene, encodes a protein involved in the nickel cobalt transport system [47, 48]. Additionally, a study showed that mutant strains lacking the rcnB gene were more susceptible to LL-37 and Magainin 2 antimicrobial peptides (AMPs) than the wild type [49]. The reason for the increased sensitivity of the rcnB mutant to BA remains unknown and requires further investigation.
The GarP protein, which belongs to the MFS (Major Facilitator Superfamily) transporter family, is involved in the transport of galactarate, glucarate, and glycerate [50, 51]. Biologically, it is classified within the scope of “anion and transmembrane transport.” GarP (UniProt P0AA80) is located in the inner membrane of the cell and contains 12 transmembrane domains. A study investigated the genes required for the intrinsic multidrug resistance of E. coli and identified garP as one of the genes involved. The study demonstrated that the garP mutant is more susceptible to chloramphenicol than the control strain [52], indicating a relationship between garP and antibiotic resistance. Our findings regarding the BA sensitivity of the garP mutant provide new information to the existing literature.
“Degradation”
Among the nine genes classified under degradation, the aceF gene appears to be the most sensitive. AceF is a part of the pyruvate dehydrogenase complex and is located in the cytosol. The pyruvate dehydrogenase enzyme is in charge of oxidatively decarboxylating pyruvate to produce acetyl-CoA. The AceF protein is specifically involved in the core region (E1) of the pyruvate dehydrogenase multi-enzyme complex and interacts with the 24-subunit [53, 54]. The deletion of aceF may have led to a reduction in energy production, potentially impacting the cells’ response to BA.
The glxK gene encodes glycerate kinase activity, which is involved in respiration. The product of this reaction was initially identified as 3-phosphoglycerate [55]. GlxK catalyzes the phosphorylation of D-glycerate and is involved in glycerolipid, amino acid, and glucose metabolism. The potential relationship between BA and glycerate could be a starting point for further investigations.
“Regulation”
The regulation category consists of 62 genes, including the torR mutant, which encodes a transcriptional regulator and belongs to the TorS/TorR two-component regulatory system. Initially, torR was found to regulate trimethylamine N-oxide reductase genes [56]. Another study discovered that TorR proteins accumulate at the poles of older cells and require interaction with DnaK and MreB proteins. This regulation is cell cycle-dependent and regulates the expression of many genes [57]. The TorS/TorR binary system has been linked to E. coli’s response to alkaline stress [58]. Furthermore, the torR gene has been shown to contribute to phosphomycin tolerance in E. coli (EHEC) O157:H7 and K12 strains [56, 59]. Therefore, there is a correlation between TorR activity and the response to antibacterial compounds and stress. It is worth noting that the torR mutant of E. coli is sensitive to BA.
YfeS is a conserved protein with an unknown function. The yfeS and yfeK genes are organized in the same operon and are controlled by the sigma 24 promoter (yfeKp sigma 24). Sigma 24 (rpoE, Sigma E factor) is involved in the expression of genes that encode membrane and periplasmic proteins in response to heat shock and other stresses. Sigma E triggers the transcription of many genes associated with heat shock and unfolded proteins. Heat shock also induces Sigma E’s expression [60]. Additionally, Sigma E is involved in resistance to zinc, cadmium, and copper [61,62,63]. Therefore, the association of Sigma E with stress and its binding site in the promoter of the yfeS gene can serve as an initial clue for investigating the relationship between BA stress and yfeS.
The galS gene encodes a protein with DNA-binding transcriptional dual regulatory activity. GalS, also known as the galactose isorepressor, is a DNA-binding transcription factor that represses the transcription of operons involved in the transport and catabolism of D-galactose [64,65,66,67,68]. Further research is needed to investigate any potential link between galactose and BA. TrpR, also known as a tryptophan transcriptional repressor, negatively regulates trp expression [69, 70]. The TrpR regulon is involved in tryptophan biosynthesis, transport, and regulation [71,72,73,74]. BA may be effective in tryptophan synthesis and potentially other aromatic amino acids, which requires further investigation in the future.
“Virulence Related”
Among the 10 virulence-related genes, the dinJ gene, YafQ-DinJ, is involved in the toxin-antitoxin system and encodes a polypeptide chain with DNA-binding transcriptional repressor activity. It is intriguing to find that a gene involved in the toxin-antitoxin system may be linked to intrinsic BA resistance. DinJ serves as the antitoxin component of the toxin-antitoxin (TA) module known as DinJ-YafQ. In this system, YafQ is a stable toxin, while DinJ is an unstable antitoxin. YafQ acts as a specific mRNA endoribonuclease that inhibits bacterial growth by impeding translation elongation under stress conditions. It has been shown to hinder protein synthesis, reduce growth rate, and inhibit colony growth. DinJ acts as an antitoxin that neutralizes YafQ’s activity by binding to it [75]. A study involving mutants of these two genes reported a decrease in biofilm formation [76]. Bacterial toxin-antitoxin (TA) systems have been associated with adaptation to stress conditions [77, 78]. E. coli harbors TA modules encoded by at least five gene pairs, namely, relBE, mazEF, chpBIK, yefM-yoeB, and dinJ-yafQ modules [78]. The DinJ-YafQ system has been confirmed as an active TA module [79]. The absence of the DinJ protein in a stressful environment has been shown to elevate RpoS levels, subsequently affecting the cell’s stress response [80]. The combination of yafQ and dinJ genes appears to be strongly linked, as shown by the STRING analysis (Fig. 5). The reason that the dinJ mutant is sensitive could be because of the increase in the activity of YafQ under BA stress. It may be a survival strategy for the bacterial population to keep some cells alive at the expense of sacrificing many others during environmental BA stress. However, further research is needed to confirm this interpretation, using various experimental approaches.
String analysis of the BA-sensitive gene hits. The predicted functional associations between the genes are shown by lines, where the number of the lines represents the linkage based on curated databases, gene-fusions, co-expression, and experimental evidence. STRING (version 10.5) with medium confidence score of 0.4 was used
“Response to Stimulus”
cheY and cheZ genes are among the 19 genes classified as response to stimulus. The cheZ gene encodes the CheZ protein, which plays a role in chemotaxis. CheZ functions as a catalyst for the dephosphorylation of CheY, a response regulator involved in the chemotactic signaling pathway of E. coli [81, 82]. Chemotaxis refers to the positive or negative reaction of cells or free-moving organisms to a chemical stimulus.
One study examined motile, but often nonchemotactic (che) mutants of E. coli. The study evaluated six genes, two of which appear to belong to the “CheA” class (cheA and cheW), and four of which appear to belong to the “CheB” class (cheX, cheB, cheY, and cheZ). Mutations in the cheA, cheW, cheX, and cheY genes did not result in any functional impairment, while mutations in the cheB and cheZ genes led to a high rate of reduced mobility. These results indicate that these two processes are likely involved in regulating variations in flagellar movement in response to chemotactic stimuli [83]. In addition, the flgC gene encodes FlgC protein, one of the four proteins composing the flagellar basal body [84]. This complex is involved in interactions with components that determine the direction of the motor, as well as with CheY and CheZ chemotaxis proteins [85]. The STRING analysis in Fig. 5 reveals a well-connected network among the cheZ, cheY, and flgC genes, suggesting that the intrinsic resistance of E. coli to BA may be somehow related to its response to stimuli.
“Biyosynthesis”
Within the biosynthesis classification, there are 12 genes, including aroK, aroB, metL, and guaA genes. Additionally, the wcaJ, galU, and otsA genes are well linked, as shown by the STRING analysis in Fig. 5. The shikimate kinase and chorismate pathway play a role in producing aromatic amino acids such as tryptophan, tyrosine, and phenylalanine. AroK has been shown to possess shikimate kinase activity [86,87,88]. In a study, the A133P mutant of AroK was found to be linked to resistance to the mecillinam antibiotic [89]. Additionally, the aroK deletion mutant was found to be more sensitive to protamine, a cationic antibiotic peptide (CAMP). The researchers also suggested that AroK enhances the production of aromatic metabolites, which function as signaling molecules [90]. The aroK gene has also been associated with sensitivity to methyl methanesulfonate [91]. According to a study conducted in biofilm and planktonic growth media, the aroK gene was found to be important for survival in competitive planktonic growth conditions [92]. Our study reveals that aroK mutants are sensitive to BA.
Furthermore, in the study by Uluisik et al. (2011), it was demonstrated that boric acid stress affects the amino acid control mechanism in yeast through eIF2a phosphorylation, which is dependent on Gcn2 kinase [93]. These findings indicate that BA has an impact on amino acid metabolism, which is also supported by the results of our study.
Conclusion
The Keio mutant line is a collection that enables the study of the phenotypes of thousands of non-essential gene mutants of E. coli in different environments. Our study conducted a genome-wide screening of 3985 E. coli mutants under varying concentrations of BA in the media, using the Keio collection to directly target the genes involved in intrinsic BA resistance. Bioinformatic analyses were conducted using the 92 BA-sensitive gene hits to identify processes and pathways associated with intrinsic BA resistance. The results showed that the “Regulation,” “Cell exterior,” and “Central Dogma” systems were relatively more affected by BA exposure. Notably, the functions of exoribonuclease II, RNA metabolic processes, and the regulation of chemotaxis activities were highlighted. In conclusion, our study indicates that intrinsic BA resistance in E. coli is achieved through multiple mechanisms working together within the cell. These findings offer valuable insights that can be used to generate new hypotheses for the scientific community investigating the mechanisms of BA on cellular life.
Data Availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Bérubé M, Dowlut M, Hall DG (2008) Benzoboroxoles as efficient glycopyranoside-binding agents in physiological conditions: structure and selectivity of complex formation. J Org Chem 73(17):6471–6479. https://doi.org/10.1021/jo800788s
Zhang Q, Tang N, Brock JWC, Mottaz HM, Ames JM, Baynes JW, Smith RD, Metz TO (2007) Enrichment and analysis of nonenzymatically glycated peptides: boronate affinity chromatography coupled with electron-transfer dissociation mass spectrometry. J Proteome Res 6(6):2323–2330. https://doi.org/10.1021/pr070112q
Nable RO, Bañuelos GS, Paull JG (1997) Boron toxicity. Plant Soil 193(2):181–198. https://doi.org/10.1023/A:1004272227886
Biţă A, Scorei IR, Bălşeanu TA, Ciocîlteu MV, Bejenaru C, Radu A, Bejenaru LE, Rău G, Mogoşanu GD, Neamţu J, Benner SA (2022) New insights into boron essentiality in humans and animals. Int J Mol Sci 23:16. https://doi.org/10.3390/ijms23169147
Çöl M, Çöl C (2003) Environmental boron contamination in waters of Hisarcik area in the Kutahya Province of Turkey. Food Chem Toxicol 41(10):1417–1420. https://doi.org/10.1016/S0278-6915(03)00160-1
Habes D, Morakchi S, Aribi N, Farine J-P, Soltani N (2006) Boric acid toxicity to the German cockroach, Blattella germanica: alterations in midgut structure, and acetylcholinesterase and glutathione S-transferase activity. Pestic Biochem Physiol 84:17–24. https://doi.org/10.1016/j.pestbp.2005.05.002
Nielsen FH (2004) Dietary fat composition modifies the effect of boron on bone characteristics and plasma lipids in rats. BioFactors 20(3):161–171. https://doi.org/10.1002/biof.5520200305
Warington K (1923) The effect of boric acid and borax on the broad bean and certain other plants. Annals of Botany, 37(148), 629–672. https://www.jstor.org/stable/43236455
O’neill MA, Ishii T, Albersheim P, Darvill AG (2004) Rhamnogalacturonan II: structure and function of a borate cross-linked cell wall pectic polysaccharide. Annu Rev PlantBiol 55:109–148. https://doi.org/10.1146/annurev.arplant.55.031903.141750
Ahmed I, Yokota A, Fujiwara T (2007) A novel highly boron tolerant bacterium, Bacillus boroniphilus sp. nov., isolated from soil, that requires boron for its growth. Extremophiles 11(2):217–224. https://doi.org/10.1007/s00792-006-0027-0
Mateo P, Bonilla I, Fernandez-Valiente E, Sanchez-Maeso E (1986) Essentiality of boron for dinitrogen fixation in Anabaena sp. PCC 7119. Plant Physiol, 81, 430–433. https://academic.oup.com/plphys/article/81/2/430/6083812.
Bolaños L, Lukaszewski K, Bonilla I, Blevins D (2004) Why boron? Plant Physiol Biochem 42(11):907–912. https://doi.org/10.1016/j.plaphy.2004.11.002
Brown PH, Bellaloui N, Wimmer MA, Bassil ES, Ruiz J, Hu H, Pfeffer H, Dannel F, Römheld V (2002) Boron in plant biology. Plant Biol 4(2):205–223. https://doi.org/10.1055/S-2002-25740
Dordas C, Brown PH (2000) Permeability of boric acid across lipid bilayers and factors affecting it. J Membr Biol 175:95–105
Kaya A, Karakaya HC, Fomenko DE, Gladyshev VN, Koc A (2009) Identification of a novel system for boron transport: Atr1 is a main boron exporter in yeast. Mol Cell Biol 29(13):3665–3674. https://doi.org/10.1128/MCB.01646-08
Lewin J (1966) Boron as a growth requirement for diatoms. J Phycol 2(4):160–163. https://doi.org/10.1111/j.1529-8817.1966.tb04616.x
Rowe RI, Eckhert CD (1999) Boron is required for zebrafish embryogenesis. J Exp Biol 202(12):1649–1654. https://doi.org/10.1242/jeb.202.12.1649
Takano J, Miwa K, Fujiwara T (2008) Boron transport mechanisms: collaboration of channels and transporters. Trends Plant Sci. https://doi.org/10.1016/j.tplants.2008.05.007
Frommer WB, von Wirén N (2002) Ping-pong with boron. Nature 420(6913):282–283. https://doi.org/10.1038/420282a
Miwa H, Fujiwara T (2009) Isolation and identification of boron-accumulating bacteria from contaminated soils and active sludge. Soil Sci Plant Nutr 55(5):643–646. https://doi.org/10.1111/j.1747-0765.2009.00402.x
Fux CA, Shirtliff M, Stoodley P, Costerton JW (2006) Can laboratory reference strains mirror “real-world” pathogenesis? Trends Microbiol 12(2):58–63. https://doi.org/10.1016/j.tim.2004.11.001
Blattner FR, Plunkett G, Bloch CA, Perna NT, Burland V, Riley M, Collado-Vides J, Glasner JD, Rode CK, Mayhew GF, Gregor J, Davis NW, Kirkpatrick HA, Goeden MA, Rose DJ, Mau B, Shao Y (1997) The complete genome sequence of Escherichia coli K-12. Science 277(5331):1453–1462. https://doi.org/10.1126/science.277.5331.1453
Hayashi K, Morooka N, Yamamoto Y, Fujita K, Isono K, Choi S, Ohtsubo E, Baba T, Wanner BL, Mori H, Horiuchi T (2006) Highly accurate genome sequences of Escherichia coli K-12 strains MG1655 and W3110. Mol Syst Biol 2:1. https://doi.org/10.1038/msb4100049
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol, 2(1), 2006.0008. https://doi.org/10.1038/MSB4100050
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97(12):6640–6645. https://doi.org/10.1073/PNAS.120163297
Kitagawa M, Ara T, Arifuzzaman M, Ioka-Nakamichi T, Inamoto E, Toyonaga H, Mori H (2005) Complete set of ORF clones of Escherichia coli ASKA library (A Complete Set of E. coli K-12 ORF Archive): unique resources for biological research. DNA Res 12(5):291–299. https://doi.org/10.1093/dnares/dsi012
Andrews JM (2001) Determination of minimum inhibitory concentrations. J Antimicrob Chemother 48(1):5–16. https://doi.org/10.1093/jac/48.suppl_1.5
Froger A, Hall JE (2007) Transformation of plasmid DNA into E. coli using the heat shock method. J Visualized Exp 6:e253. https://doi.org/10.3791/253
Dibek E, Sezer Kürkçü M, Çiftçi BH, Çöl B (2020) P1 transdüksiyon yöntemi ile birden fazla gen bakımından mutant olan Escherichia coli suşlarının elde edilmesi. Bitlis Eren Üniv Fen Bilimleri Dergisi 9(1):110–119. https://doi.org/10.17798/bitlisfen.588763
Silhavy TJ, Berman ML, Enquist LW (1984) Experiments with gene fusions. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Keseler IM, Mackie A, Santos-Zavaleta A, Billington R, Bonavides-Martínez C, Caspi R, Fulcher C, Gama-Castro S, Kothari A, Krummenacker M, Latendresse M, Muñiz-Rascado L, Ong Q, Paley S, Peralta-Gil M, Subhraveti P, Velázquez-Ramírez DA, Weaver D, Collado-Vides J, Karp PD (2017) The EcoCyc database: reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res 45(D1):D543–D550. https://doi.org/10.1093/nar/gkw1003
Sherman BT, Hao M, Qiu J, Jiao X, Baseler MW, Lane HC, Imamichi T, Chang W (2022) DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 50(W1):W216–W221. https://doi.org/10.1093/nar/gkac194
Huang DW, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13. https://doi.org/10.1093/nar/gkn923
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Jensen LJ, Von Mering C (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368. https://doi.org/10.1093/nar/gkw937
Miwa K, Takano J, Omori H, Seki M, Shinozaki K, Fujiwara T (2007) Plants tolerant of high boron levels. Science 318(5855):1417–1417. https://doi.org/10.1126/science.1146634
Sutton T, Baumann U, Hayes J, Collins NC, Shi B-J, Schnurbusch T, Hay A, Mayo G, Pallotta M, Tester M, Langridge P (2007) Boron-toxicity tolerance in barley arising from efflux transporter amplification. Science 318(5855):1446–1449. https://doi.org/10.1126/science.1146853
Takano J, Miwa K, Yuan L, von Wirén N, Fujiwara T (2005) Endocytosis and degradation of BOR1, a boron transporter of Arabidopsis thaliana, regulated by boron availability. Proc Natl Acad Sci 102(34):12276–12281. https://doi.org/10.1073/pnas.0502060102
Lockhart DJ, Winzeler EA (2000) Genomics, gene expression and DNA arrays. Nature 405(6788):827–836. https://doi.org/10.1038/35015701
Richmond CS, Glasner JD, Mau R, Jin H, Blattner FR (1999) Genome-wide expression profiling in Escherichia coli K-12. Nucleic Acids Research, 27(19), 3821–3835. https://academic.oup.com/nar/article/27/19/3821/1055563
Tao H, Bausch C, Richmond C, Blattner FR, Conway T (1999) Functional genomics: expression analysis of Escherichia coli growing on minimal and rich media. J Bacteriol 181(20):6425–6440
Wei Y, Lee J-M, Richmond C, Blattner FR, Rafalski JA, Larossa RA (2001) High-density microarray-mediated gene expression profiling of Escherichia coli. J Bacteriol 183(2):545–556. https://doi.org/10.1128/JB.183.2.545-556.2001
Yoon SH, Han MJ, Lee SY, Jeong KJ, Yoo JS (2003) Combined transcriptome and proteome analysis of Escherichia coli during high cell density culture. Biotechnol Bioeng 81(7):753–767. https://doi.org/10.1002/BIT.10626
Ikeuchi Y, Shigi N, Kato J, Nishimura A, Suzuki T (2006) Mechanistic insights into sulfur relay by multiple sulfur mediators involved in thiouridine biosynthesis at tRNA wobble positions. Mol Cell 21(1):97–108. https://doi.org/10.1016/j.molcel.2005.11.001
Kambampati R, Lauhon CT (2003) MnmA and IscS are required for in vitro 2-thiouridine biosynthesis in Escherichia coli. Biochemistry. https://doi.org/10.1021/bi026536
Shomron N, Ast G (2003) Boric acid reversibly inhibits the second step of pre-mRNA splicing. FEBS Lett 552(2–3):219–224. https://doi.org/10.1016/s0014-5793(03)00928-1
Arends J, Thomanek N, Kuhlmann K, Marcus K, Narberhaus F (2016) In vivo trapping of FtsH substrates by label-free quantitative proteomics. Proteomics 16(24):3161–3172. https://doi.org/10.1002/PMIC.201600316
Blériot C, Effantin G, Lagarde F, Mandrand-Berthelot M-A, Rodrigue AS (2011) RcnB is a periplasmic protein essential for maintaining intracellular Ni and Co concentrations in Escherichia coli. J Bacteriol 193(15):3785–3793. https://doi.org/10.1128/JB.05032-11
Blériot C, Gault M, Gueguen E, Arnoux P, Pignol D, Marie-André Mandrand-Berthelot BCD, Rodrigue A (2014) Cu binding by the Escherichia coli metal-efflux accessory protein RcnB. Metallomics 6(8):1400–1409. https://doi.org/10.1039/c4mt00036f
Kozlowska J, Vermeer LS, Rogers GB, Rehnnuma N, Amos S-BTA, Koller G, McArthur M, Bruce KD, Mason AJ (2014) Combined systems approaches reveal highly plastic responses to antimicrobial peptide challenge in Escherichia coli. PLoS Pathog 10(5):e1004104. https://doi.org/10.1371/journal.ppat.1004104
Pao SS, Paulsen IT, Saier MH (1998) Major facilitator superfamily. Microbiol Mol Biol Rev 62(1):1–34. https://doi.org/10.1128/MMBR.62.1.1-34.1998
Monterrubio R, Baldoma L, Obradors N, Aguilar J, Badia J (2000) A common regulator for the operons encoding the enzymes involved in D-galactarate, D-glucarate, and D-glycerate utilization in Escherichia coli. J Bacteriol 182(9):2672–2674. https://doi.org/10.1128/JB.182.9.2672-2674.2000
Duo M, Hou S, Ren D (2008) Identifying Escherichia coli genes involved in intrinsic multidrug resistance. Appl Microbiol Biotechnol 81(4):731–741. https://doi.org/10.1007/s00253-008-1709-6
Wagenknecht T, Grassucci R, Schaak D (1990) Cryoelectron microscopy of frozen-hydrated alpha-ketoacid dehydrogenase complexes from Escherichia coli. J Biol Chem 265(36):22402–22408. https://doi.org/10.1016/S0021-9258(18)45719-5
Yang HC, Hainfeld JF, Wall JS, Frey PA (1985) Quaternary structure of pyruvate dehydrogenase complex from Escherichia coli. J Biol Chem 260(30):16049–16051. https://doi.org/10.1016/S0021-9258(17)36196-3
Doughty CC, Hayashi JA, Guenther HL (1966) Purification and properties of D-glycerate 3-kinase from Escherichia coli. J Biol Chem 241(3):568–572
Simon G, Mejean V, Jourlin C, Chippaux M, Pascal M-C (1994) The torR gene of Escherichia coli encodes a response regulator protein involved in the expression of the trimethylamine N-oxide reductase genes. J Bacteriol 176(18):5601–5606
Yao Y, Fan L, Shi Y, Odsbu I, Morigen (2016) A spatial control for correct timing of gene expression during the Escherichia coli cell cycle. Genes 8(1):1–20. https://doi.org/10.3390/genes8010001
Bordi C, Théraulaz L, Méjean V, Jourlin-Castelli C (2003) Anticipating an alkaline stress through the Tor phosphorelay system in Escherichia coli. Mol Microbiol 48(1):211–223. https://doi.org/10.1046/j.1365-2958.2003.03428.x
Kurabayashi K, Hirakawa Y, Tanimoto K, Tomita H, Hirakawa H (2015) Identification of a second two-component signal transduction system that controls fosfomycin tolerance and glycerol-3-phosphate uptake. J Bacteriol 197(5):861–871. https://doi.org/10.1128/JB.02491-14
Ades SE, Grigorova IL, Gross CA (2003) Regulation of the alternative sigma factor sigma(E) during initiation, adaptation, and shutoff of the extracytoplasmic heat shock response in Escherichia coli. J Bacteriol 185(8):2512–2519. https://doi.org/10.1128/JB.185.8.2512-2519.2003
Egler M, Grosse C, Grass G, Nies DH (2005) Role of the extracytoplasmic function protein family sigma factor RpoE in metal resistance of Escherichia coli. J Bacteriol 187(7):2297–2307. https://doi.org/10.1128/JB.187.7.2297-2307.2005
Hiratsu K, Amemura M, Nashimoto H, Shinagawa H, Makino K (1995) The rpoE gene of Escherichia coli, which encodes sigma E, is essential for bacterial growth at high temperature. J Bacteriol 177(10):2918–2922. https://doi.org/10.1128/jb.177.10.2918-2922.1995
Rouvière PE, De Las Peñas A, Mecsas J, Lu CZ, Rudd KE, Gross CA (1995) rpoE, the gene encoding the second heat-shock sigma factor, sigma E. Escherichia coli EMBO J 14(5):1032–1042. https://doi.org/10.1002/j.1460-2075.1995.tb07084.x
Geanacopoulos M, Adhya S (1997) Functional characterization of roles of GalR and GalS as regulators of the gal regulon. J Bacteriol 179(1):228–234. https://doi.org/10.1128/jb.179.1.228-234.1997
Semsey S, Krishna S, Sneppen K, Adhya S (2007) Signal integration in the galactose network of Escherichia coli. Mol Microbiol 65(2):465–476. https://doi.org/10.1111/j.1365-2958.2007.05798.x
Weickert MJ, Adhya S (1992) Isorepressor of the gal regulon in Escherichia coli. J Mol Biol 226(1):69–83. https://doi.org/10.1016/0022-2836(92)90125-4
Weickert MJ, Adhya S (1993) Control of transcription of gal repressor and isorepressor genes in Escherichia coli. J Bacteriol 175(1):251–258. https://doi.org/10.1128/jb.175.1.251-258.1993
Weickert MJ, Adhya S (1993) The galactose regulon of Escherichia coli. Mol Microbiol 10(2):245–251. https://doi.org/10.1111/j.1365-2958.1993.tb01950.x
Gunsalus RP, Yanofsky C (1980) Nucleotide sequence and expression of Escherichia coli trpR, the structural gene for the trp aporepressor. Proc Natl Acad Sci 77(12):7117–7121. https://doi.org/10.1073/pnas.77.12.7117
Squires CL, Lee FD, Yanofsky C (1975) Interaction of the trp repressor and RNA polymerase with the trp operon. J Mol Biol 92(1):93–111. https://doi.org/10.1016/0022-2836(75)90093-5
Heatwole VM, Somerville RL (1991) The tryptophan-specific permease gene, mtr, is differentially regulated by the tryptophan and tyrosine repressors in Escherichia coli K-12. J Bacteriol 173(11):3601–3604. https://doi.org/10.1128/jb.173.11.3601-3604.1991
Lawley B, Pittard AJ (1994) Regulation of aroL expression by TyrR protein and Trp repressor in Escherichia coli K-12. J Bacteriol 176(22):6921–6930. https://doi.org/10.1128/jb.176.22.6921-6930.1994
Sarsero JP, Wookey PJ, Pittard AJ (1991) Regulation of expression of the Escherichia coli K-12 mtr gene by TyrR protein and Trp repressor. J Bacteriol 173(13):4133–4143. https://doi.org/10.1128/jb.173.13.4133-4143.1991
Otwinowski Z, Schevitz RW, Zhang R-G, Lawson CL, Joachimiak A, Marmorstein RQ, Luisi BF, Sigler PB (1988) Crystal structure of trp represser/operator complex at atomic resolution. Nature 335(6188):321–329. https://doi.org/10.1038/335321a0
Prysak MH, Mozdzierz CJ, Cook AM, Zhu L, Zhang Y, Inouye M, Woychik NA (2009) Bacterial toxin YafQ is an endoribonuclease that associates with the ribosome and blocks translation elongation through sequence-specific and frame-dependent mRNA cleavage. Mol Microbiol 71(5):1071–1087. https://doi.org/10.1111/j.1365-2958.2008.06572.x
Harrison JJ, Wade WD, Akierman S, Vacchi-Suzzi C, Stremick CA, Turner RJ, Ceri H (2009) The chromosomal toxin gene yafQ is a determinant of multidrug tolerance for Escherichia coli growing in a biofilm. Antimicrob Agents Chemother 53(6):2253–2258. https://doi.org/10.1128/AAC.00043-09
Buts L, Lah J, Dao-Thi M-H, Wyns L, Loris R (2005) Toxin–antitoxin modules as bacterial metabolic stress managers. Trends Biochem Sci 30(12):672–679. https://doi.org/10.1016/j.tibs.2005.10.004
Gerdes K, Christensen SK, Løbner-Olesen A (2005) Prokaryotic toxin–antitoxin stress response loci. Nat Rev Microbiol 3(5):371–382. https://doi.org/10.1038/nrmicro1147
Motiejūnaitė R, Armalytė J, Markuckas A, Sužiedėlienė E (2007) Escherichia coli dinJ-yafQ genes act as a toxin-antitoxin module. FEMS Microbiol Lett 268(1):112–119. https://doi.org/10.1111/j.1574-6968.2006.00563.x
Hu Y, Benedik MJ, Wood TK (2012) Antitoxin DinJ influences the general stress response through transcript stabilizer CspE. Environ Microbiol 14(3):669–679. https://doi.org/10.1111/j.1462-2920.2011.02618.x
Silversmith RE, Levin MD, Schilling E, Bourret RB (2008) Kinetic characterization of catalysis by the chemotaxis phosphatase CheZ. J Biol Chem 283(2):756–765. https://doi.org/10.1074/jbc.M704400200
Zhao R, Collins EJ, Bourret RB, Silversmith RE (2002) Structure and catalytic mechanism of the E. coli chemotaxis phosphatase CheZ. Nat Struct Biol 9:570–575. https://doi.org/10.1038/nsb816
Parkinson JS (1978) Complementation analysis and deletion mapping of Escherichia coli mutants defective in chemotaxis. J Bacteriol 135(1):45–53. https://doi.org/10.1128/jb.135.1.45-53.1978
Jones CJ, Macnab RM, Okino H, Aizawa S-I (1990) Stoichiometric analysis of the flagellar hook-(basal-body) complex of Salmonella typhimurium. J Mol Biol 212(2):377–387. https://doi.org/10.1016/0022-2836(90)90132-6
Sockett H, Yamaguchi S, Kihara M, Irikura VM, Macnab RM (1992) Molecular analysis of the flagellar switch protein FliM of Salmonella typhimurium. J Bacteriol 174(3):793–806. https://doi.org/10.1128/jb.174.3.793-806.1992
Berlyn MB, Giles NH (1969) Organization of enzymes in the polyaromatic synthetic pathway: separability in bacteria. J Bacteriol 99(1):222–230. https://doi.org/10.1128/jb.99.1.222-230.1969
Ely B, Pittard J (1979) Aromatic amino acid biosynthesis: regulation of shikimate kinase in Escherichia coli K-12. J Bacteriol 138(3):933–943. https://doi.org/10.1128/jb.138.3.933-943.1979
DeFeyter RC, Pittard J (1986) Purification and properties of shikimate kinase II from Escherichia coli K-12. J Bacteriol 165(1):331–333. https://doi.org/10.1128/jb.165.1.331-333.1986
Vinella D, Gagny B, Joseleau-Petit D, D’Ari R, Cashel M (1996) Mecillinam resistance in Escherichia coli is conferred by loss of a second activity of the AroK protein. J Bacteriol 178(13):3818–3828. https://doi.org/10.1128/jb.178.13.3818-3828.1996
Weatherspoon-Griffin N, Yang D, Kong W, Hua Z, Shi Y (2014) The CpxR/CpxA two-component regulatory system up-regulates the multidrug resistance cascade to facilitate Escherichia coli resistance to a model antimicrobial peptide. J Biol Chem 289(47):32571–32582. https://doi.org/10.1074/jbc.M114.565762
Rooney JP, George AD, Patil A, Begley U, Bessette E, Zappala MR, Huang X, Conklin DS, Cunningham RP, Begley TJ (2009) Systems based mapping demonstrates that recovery from alkylation damage requires DNA repair, RNA processing, and translation associated networks. Genomics 93(1):42–51. https://doi.org/10.1016/j.ygeno.2008.09.001
Junker LM, Peters JE, Hay AG (2006) Global analysis of candidate genes important for fitness in a competitive biofilm using DNA-array-based transposon mapping. Microbiol (Read Engl) 152(Pt 8):2233–2245. https://doi.org/10.1099/mic.0.28767-0
Uluisik I, Kaya A, Fomenko DE, Karakaya HC, Carlson BA, Gladyshev VN, Koc A (2011) Boron stress activates the general amino acid control mechanism and inhibits protein synthesis. PLoS ONE 6(11):e27772. https://doi.org/10.1371/journal.pone.002777
Acknowledgements
We thank the Scientific and Technological Research Council of Turkey (TUBİTAK). We would like to express our gratitude to Prof. Dr. Timothy J. Larson for his assistance and support with P1 transduction experiments. The National BioResource Project [NIG] of Japan provided the Keio collection used in this study, for which we are thankful.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK). This study was funded by the Scientific and Technological Research Council of Turkey (TUBİTAK) 114Z987 project.
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of BÇ, MSK, and ED.
All authors have given approval to the final version of the manuscript. Details of the contributions of each author: design (BÇ) and supervision (BÇ) of the study, culturing and isolation of Escherichia coli strains and plasmids (MSK, ED), genome-wide screening of intrinsic boric acid resistance associated genes with the replicator (MSK, ED), sequential spotting (MSK, ED), determination of MIC (MSK), complementation of sensitive mutant strains (MSK), synthetic phenotype analysis of double and triple deletion mutants (MSK, ED), data mining, analysis and interpretation (MSK, BÇ), drafting (MSK, BÇ), and revision (MSK, BÇ) of the manuscript. Abbreviation of author names: BÇ Bekir Çöl; MSK Merve Sezer Kürkçü; ED Esra Dibek
Corresponding author
Ethics declarations
Ethics Approval
The study does not require ethics approval.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Çöl, B., Kürkçü, M.S. & Di̇bek, E. Genome-Wide Screens Identify Genes Responsible for Intrinsic Boric Acid Resistance in Escherichia coli. Biol Trace Elem Res (2024). https://doi.org/10.1007/s12011-024-04129-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12011-024-04129-0