Abstract
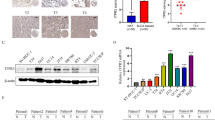
Exposure to cadmium (Cd), a non-essential heavy metal, leads to the malignant transformation of urothelial cells and promotes bladder cancer (BC) development, but the mechanisms are unclear. Therefore, we aimed to explore the possible molecules associated with Cd-related BC. By analyzing and mining biological big data in public databases, we screened genes associated with the malignant transformation of uroepithelial cells caused by Cd and further screened the key gene associated with BC prognosis from these genes. The possible roles of the key gene in BC progression were then further explored through biological big data analysis and cellular experiments. Data mining yielded a total of 387 malignant transformation-related genes, which were enriched in multiple cancer-related signaling pathways, such as cytokine-cytokine receptor interaction, Toll-like receptor signaling pathway, and Jak-STAT signaling pathway. Further screening identified Fibronectin 1 (FN1) as the key gene. High expression of FN1 was associated with the advanced pathologic stage, T stage, N stage, and M stage and predicted an unfavorable outcome in BC patients. FN1 expression was positively associated with the infiltration of several types of immune cells, particularly tumor-associated macrophages and cancer-associated fibroblasts. Overexpression of FN1 could be detected in Cd-treated urothelial cells and BC cell lines. Interestingly, silencing of FN1 impaired the proliferation and invasive capacity of BC cells. In conclusion, the present study provides new insight into the mechanism of Cd-related BC. FN1 might be a prognostic marker and therapeutic target for BC. Future studies are needed to confirm these results.



Similar content being viewed by others
Abbreviations
- Cd :
-
Cadmium
- BC :
-
Bladder cancer
- DEGs :
-
Differentially expressed genes
- GEO :
-
Gene Expression Omnibus
- GO :
-
Gene Ontology
- KEGG :
-
Kyoto Encyclopedia of Genes and Genomes
- PPI :
-
Protein–protein interaction
- TCGA :
-
The Cancer Genome Atlas
- DMEM :
-
Dulbecco’s modified Eagle’s medium
- FN1 :
-
Fibronectin 1
- FDR :
-
False discovery rate
- CAFs :
-
Cancer-associated fibroblasts
- TME :
-
Tumor microenvironment
- TAMs :
-
Tumor-associated macrophages
References
Antoni S, Ferlay J, Soerjomataram I, Znaor A, Jemal A, Bray F (2017) Bladder cancer incidence and mortality: a global overview and recent trends. Eur Urol 71(1):96–108. https://doi.org/10.1016/j.eururo.2016.06.010
Richters A, Aben KKH, Kiemeney L (2019) The global burden of urinary bladder cancer: an update. World J Urol. https://doi.org/10.1007/s00345-019-02984-4
Malats N, Real FX (2015) Epidemiology of bladder cancer. Hematol Oncol Clin North Am 29(2):177–189, vii. https://doi.org/10.1016/j.hoc.2014.10.001
Cooley LF, McLaughlin KA, Meeks JJ (2020) Genomic and therapeutic landscape of non-muscle-invasive bladder cancer. Urol Clin North Am 47(1):35–46. https://doi.org/10.1016/j.ucl.2019.09.006
Soza-Ried C, Bustamante E, Caglevic C, Rolfo C, Sirera R, Marsiglia H (2019) Oncogenic role of arsenic exposure in lung cancer: a forgotten risk factor. Crit Rev Oncol Hematol 139:128–133. https://doi.org/10.1016/j.critrevonc.2019.01.012
Liao LM, Friesen MC, Xiang YB, Cai H, Koh DH, Ji BT, Yang G, Li HL, Locke SJ, Rothman N, Zheng W, Gao YT, Shu XO, Purdue MP (2016) Occupational lead exposure and associations with selected cancers: the Shanghai Men’s and Women’s Health Study Cohorts. Environ Health Perspect 124(1):97–103. https://doi.org/10.1289/ehp.1408171
Charoenngam N, Ponvilawan B, Ungprasert P (2021) Higher zinc intake is associated with decreased risk of lung cancer. J Evid Based Med 14(3):185–187. https://doi.org/10.1111/jebm.12448
Banerjee M, Yaddanapudi K, States JC (2022) Zinc supplementation prevents mitotic accumulation in human keratinocyte cell lines upon environmentally relevant arsenic exposure. Toxicol Appl Pharmacol 454:116255. https://doi.org/10.1016/j.taap.2022.116255
Kadkol S, Diamond AM (2020) The interaction between dietary selenium intake and genetics in determining cancer risk and outcome. Nutrients 12 (8). https://doi.org/10.3390/nu12082424
Garcia-Esquinas E, Pollan M, Tellez-Plaza M, Francesconi KA, Goessler W, Guallar E, Umans JG, Yeh J, Best LG, Navas-Acien A (2014) Cadmium exposure and cancer mortality in a prospective cohort: the strong heart study. Environ Health Perspect 122(4):363–370. https://doi.org/10.1289/ehp.1306587
Souza-Arroyo V, Fabian JJ, Bucio-Ortiz L, Miranda-Labra RU, Gomez-Quiroz LE, Gutierrez-Ruiz MC (2022) The mechanism of the cadmium-induced toxicity and cellular response in the liver. Toxicology 480:153339. https://doi.org/10.1016/j.tox.2022.153339
Koedrith P, Kim H, Weon JI, Seo YR (2013) Toxicogenomic approaches for understanding molecular mechanisms of heavy metal mutagenicity and carcinogenicity. Int J Hyg Environ Health 216(5):587–598. https://doi.org/10.1016/j.ijheh.2013.02.010
Andersson EM, Sandsveden M, Forsgard N, Sallsten G, Manjer J, Engstrom G, Barregard L (2021) Is cadmium a risk factor for breast cancer - results from a nested case-control study using data from the Malmo Diet and Cancer Study. Cancer Epidemiol Biomarkers Prev 30(9):1744–1752. https://doi.org/10.1158/1055-9965.EPI-21-0181
Xu Y, Mu W, Li J, Ba Q, Wang H (2021) Chronic cadmium exposure at environmental-relevant level accelerates the development of hepatotoxicity to hepatocarcinogenesis. Sci Total Environ 783:146958. https://doi.org/10.1016/j.scitotenv.2021.146958
Shi H, Sun X, Kong A, Ma H, Xie Y, Cheng D, Wong CKC, Zhou Y, Gu J (2021) Cadmium induces epithelial-mesenchymal transition and migration of renal cancer cells by increasing PGE2 through a cAMP/PKA-COX2 dependent mechanism. Ecotoxicol Environ Saf 207:111480. https://doi.org/10.1016/j.ecoenv.2020.111480
Awadalla A, Mortada WI, Abol-Enein H, Shokeir AA (2020) Correlation between blood levels of cadmium and lead and the expression of microRNA-21 in Egyptian bladder cancer patients. Heliyon 6(12):e05642. https://doi.org/10.1016/j.heliyon.2020.e05642
Hoggarth ZE, Osowski DB, Freeberg BA, Garrett SH, Sens DA, Sens MA, Zhou XD, Zhang KK, Somji S (2018) The urothelial cell line UROtsa transformed by arsenite and cadmium display basal characteristics associated with muscle invasive urothelial cancers. PLoS ONE 13(12):e0207877. https://doi.org/10.1371/journal.pone.0207877
Wu B, Jiang X, Huang Y, Ying X, Zhang H, Liu B, Li Z, Qi D, Ji W, Cai X (2021) Integrated analysis of mRNA-m(6)A-protein profiles reveals novel insights into the mechanisms for cadmium-induced urothelial transformation. Biomarkers 26(6):499–507. https://doi.org/10.1080/1354750X.2021.1913513
Chang JT, Nevins JR (2006) GATHER: a systems approach to interpreting genomic signatures. Bioinformatics 22(23):2926–2933. https://doi.org/10.1093/bioinformatics/btl483
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613. https://doi.org/10.1093/nar/gky1131
Vasaikar SV, Straub P, Wang J, Zhang B (2018) LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res 46(D1):D956–D963. https://doi.org/10.1093/nar/gkx1090
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6(269):l1. https://doi.org/10.1126/scisignal.2004088
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, Varambally S (2017) UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 19(8):649–658. https://doi.org/10.1016/j.neo.2017.05.002
Ma Q, Chen Y, Xiao F, Hao Y, Song Z, Zhang J, Okuda K, Um SW, Silva M, Shimada Y, Si C, Liang C (2021) A signature of estimate-stromal-immune score-based genes associated with the prognosis of lung adenocarcinoma. Transl Lung Cancer Res 10(3):1484–1500. https://doi.org/10.21037/tlcr-21-223
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, Li B, Liu XS (2017) TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res 77(21):e108–e110. https://doi.org/10.1158/0008-5472.CAN-17-0307
Hu ZW, Sun W, Wen YH, Ma RQ, Chen L, Chen WQ, Lei WB, Wen WP (2022) CD69 and SBK1 as potential predictors of responses to PD-1/PD-L1 blockade cancer immunotherapy in lung cancer and melanoma. Front Immunol 13:952059. https://doi.org/10.3389/fimmu.2022.952059
Feng S, Yuan W, Sun Z, Guo X, Ling J, Chang A, Zhao H, Zhuo X (2022) SPP1 as a key gene in the lymph node metastasis and a potential predictor of poor prognosis in head and neck carcinoma. J Oral Pathol Med. https://doi.org/10.1111/jop.13333
Goswami CP, Nakshatri H (2014) PROGgeneV2: enhancements on the existing database. BMC Cancer 14:970. https://doi.org/10.1186/1471-2407-14-970
Liu N, Zhang GD, Bai P, Su L, Tian H, He M (2022) Eight hub genes as potential biomarkers for breast cancer diagnosis and prognosis: a TCGA-based study. World J Clin Oncol 13(8):675–688. https://doi.org/10.5306/wjco.v13.i8.675
Nouri Y, Weinkove R, Perret R (2021) T-cell intrinsic Toll-like receptor signaling: implications for cancer immunotherapy and CAR T-cells. J Immunother Cancer 9 (11). https://doi.org/10.1136/jitc-2021-003065
Zhu J, Li Y, Lv X (2022) IL4I1 enhances PD-L1 expression through JAK/STAT signaling pathway in lung adenocarcinoma. Immunogenetics. https://doi.org/10.1007/s00251-022-01275-4
Amundson SA, Smilenov LB (2010) Integration of biological knowledge and gene expression data for biomarker selection: FN1 as a potential predictor of radiation resistance in head and neck cancer. Cancer Biol Ther 10(12):1252–1255. https://doi.org/10.4161/cbt.10.12.13731
Jiang K, Liu H, Xie D, Xiao Q (2019) Differentially expressed genes ASPN, COL1A1, FN1, VCAN and MUC5AC are potential prognostic biomarkers for gastric cancer. Oncol Lett 17(3):3191–3202. https://doi.org/10.3892/ol.2019.9952
Wang S, Gao B, Yang H, Liu X, Wu X, Wang W (2019) MicroRNA-432 is downregulated in cervical cancer and directly targets FN1 to inhibit cell proliferation and invasion. Oncol Lett 18(2):1475–1482. https://doi.org/10.3892/ol.2019.10403
Xie Y, Liu C, Qin Y, Chen J, Fang J (2019) Knockdown of IRE1a suppresses metastatic potential of colon cancer cells through inhibiting FN1-Src/FAK-GTPases signaling. Int J Biochem Cell Biol 114:105572. https://doi.org/10.1016/j.biocel.2019.105572
Chen Z, Tao Q, Qiao B, Zhang L (2019) Silencing of LINC01116 suppresses the development of oral squamous cell carcinoma by up-regulating microRNA-136 to inhibit FN1. Cancer Manag Res 11:6043–6059. https://doi.org/10.2147/CMAR.S197583
Wang D, Ye Q, Gu H, Chen Z (2022) The role of lipid metabolism in tumor immune microenvironment and potential therapeutic strategies. Front Oncol 12:984560. https://doi.org/10.3389/fonc.2022.984560
Xiao C, Tian H, Zheng Y, Yang Z, Li S, Fan T, Xu J, Bai G, Liu J, Deng Z, Li C, He J (2022) Glycolysis in tumor microenvironment as a target to improve cancer immunotherapy. Front Cell Dev Biol 10:1013885. https://doi.org/10.3389/fcell.2022.1013885
Perez S, Rius-Perez S (2022) Macrophage polarization and reprogramming in acute inflammation: a redox perspective. Antioxidants (Basel) 11 (7). https://doi.org/10.3390/antiox11071394
Chabeli MS, Wang X, Yinghao L, Chen C, Yang C, Shou Y, Wang S, Chen K (2022) Similarities between wound re-epithelialization and metastasis in ESCC and the crucial involvement of macrophages: a review. Cancer Treat Res Commun 32:100621. https://doi.org/10.1016/j.ctarc.2022.100621
Qin R, Ren W, Ya G, Wang B, He J, Ren S, Jiang L, Zhao S (2022) Role of chemokines in the crosstalk between tumor and tumor-associated macrophages. Clin Exp Med. https://doi.org/10.1007/s10238-022-00888-z
Wu C, Gu J, Gu H, Zhang X, Zhang X, Ji R (2022) The recent advances of cancer associated fibroblasts in cancer progression and therapy. Front Oncol 12:1008843. https://doi.org/10.3389/fonc.2022.1008843
Yamamoto Y, Kasashima H, Fukui Y, Tsujio G, Yashiro M, Maeda K (2022) The heterogeneity of cancer-associated fibroblast subpopulations: their origins, biomarkers, and roles in the tumor microenvironment. Cancer Sci. https://doi.org/10.1111/cas.15609
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by LZ, YW, MS, and AC. The first draft of the manuscript was written by LZ, WZ, and YZ. All authors read, reviewed, and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for Publication
All authors approved for this publication.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, L., Wang, Y., Song, M. et al. Fibronectin 1 as a Key Gene in the Genesis and Progression of Cadmium-Related Bladder Cancer. Biol Trace Elem Res 201, 4349–4359 (2023). https://doi.org/10.1007/s12011-022-03510-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12011-022-03510-1