Abstract
Bladder cancer (BC) is one of the most often reported malignancies globally, with a high recurrence rate and associated morbidity and mortality, especially in advanced BC. There has been a surge in the number of molecular targets revealed for BC prognosis and treatment. However, there is still a great need to discover novel biomarkers. Consequently, the current study investigated biomarkers that might indicate the progression of bladder cancer. In this study, bioinformatics analysis was done on a single GEO dataset, and TCGA-BLCA information was connected with differentially expressed genes (DEGs). The levels of mRNA and protein expression were validated using qRT-PCR. According to our findings, CRYAB, ECM1, ALDOB, AOC, GPX3, IGFBP7, AQP2, LASS2, TMEM176A, GALNT1, and LASS2 were highly enriched in cell division, identical protein binding, and developmental process in bladder cancer patients. In addition, among the highly differentiated genes, ECM1, GALNT1, LASS2, and GPX3 showed significant molecular alterations in BC, which are crucial for marker identification. Moreover, the mRNA, CNVs, and protein levels of ECM1, GALNT1, LASS2, and GPX3 were significantly increased in BC patients. Our predictions and analysis studies stated that these four genes act as urine biomarkers and played a crucial role in disease prognosis and the therapeutic process of bladder cancer. Our outcomes showed that these four novel urine biomarkers have the potential to provide innovative diagnostics, early predictions, and disease targets, ultimately improving the BC patient’s prognosis.
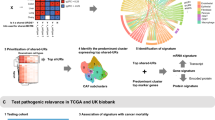
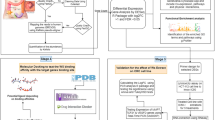
Graphical Abstract










Similar content being viewed by others
Data Availability
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- ALDOB :
-
Aldolase B enzyme
- AOC :
-
Allene oxide cyclase
- AQP2 :
-
Aquaporin 2
- BC :
-
Bladder cancer
- cDNA :
-
Complementary deoxyribonucleic acid
- CNVs :
-
Copy number variation
- CRYAB :
-
Alpha-crystallin B chain
- DEG :
-
Differentially expressed genes
- ECM1 :
-
Extracellular matrix protein 1
- EV :
-
Extracellular vesicles
- GalNAc :
-
N-acetyl-d-galactosamine
- GALNT1 :
-
N-acetylgalactosaminyltransferase 1
- GEO :
-
Gene Expression Omnibus
- GPX3 :
-
Glutathione peroxidase 3
- IGFBP7 :
-
Insulin-like growth factor-binding protein 7
- KEGG :
-
Kyoto Encyclopedia of Genes and Genomes
- LASS2 :
-
LAG1 longevity-assurance homolog 2
- MIBC :
-
Muscle-invasive bladder cancer
- mRNA :
-
Messenger RNA
- NGS :
-
Next-generation sequencing
- NIMBC :
-
Non-muscle-invasive bladder carcinomas
- NMP22 :
-
Nuclear matrix protein 22
- PPI :
-
Protein-protein interaction networks
- qRT-PCR :
-
Real-time quantitative reverse transcription-polymerase chain reaction
- STRING :
-
Search Tool for the Retrieval of Interacting Genes/Proteins
- TCGA :
-
The Cancer Genome Atlas
- TCGA-BLCA :
-
Cancer genome atlas bladder cancer
- TMEM176A :
-
Human transmembrane protein 176A
- UALCAN :
-
User-friendly, interactive web resource for analyzing cancer transcriptome data
References
Vartolomei, M.D., Porav-Hodade, D., Ferro, M., Mathieu, R., Abufaraj, M., Foerster, B., et al. (Eds). (2018). Prognostic role of pretreatment neutrophil-to-lymphocyte ratio (NLR) in patients with non–muscle-invasive bladder cancer (NMIBC): A systematic review and meta-analysis. Urologic Oncology: Seminars and Original Investigations; Elsevier.
Chen, L., Li, W., Li, Z., Song, Y., Zhao, J., Chen, Z., et al. (2021). circNUDT21 promotes bladder cancer progression by modulating the miR-16–1–3p/MDM2/p53 axis. Molecular Therapy. Nucleic Acids, 26, 625–36.
Oeyen, E., Hoekx, L., De Wachter, S., Baldewijns, M., Ameye, F., & Mertens, I. (2019). Bladder cancer diagnosis and follow-up: The current status and possible role of extracellular vesicles. International Journal of Molecular Sciences, 20(4), 821.
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R. L., Torre, L. A., & Jemal, A. (2018). Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 68(6), 394–424.
Batista, R., Vinagre, J., Prazeres, H., Sampaio, C., Peralta, P., Conceição, P., et al. (2019). Validation of a novel, sensitive, and specific urine-based test for recurrence surveillance of patients with non-muscle-invasive bladder cancer in a comprehensive multicenter study. Frontiers in Genetics, 10, 1237.
Ng, K., Stenzl, A., Sharma, A., Vasdev, N., (Eds). (2021). Urinary biomarkers in bladder cancer: A review of the current landscape and future directions. Urologic Oncology: Seminars and Original Investigations; Elsevier.
Parker, J., & Spiess, P. E. (2011). Current and emerging bladder cancer urinary biomarkers. The Scientific World Journal, 11, 1103–12.
Bhat, A., & Ritch, C. R. (2019). Urinary biomarkers in bladder cancer: Where do we stand? Current Opinion in Urology, 29(3), 203–9.
di Meo, N. A., Loizzo, D., Pandolfo, S. D., Autorino, R., Ferro, M., Porta, C., et al. (2022). Metabolomic approaches for detection and identification of biomarkers and altered pathways in bladder cancer. International Journal of Molecular Sciences, 23(8), 4173.
Mohsenzadegan, M., Razmi, M., Vafaei, S., Abolhasani, M., Madjd, Z., SaeednejadZanjani, L., et al. (2022). Co-expression of cancer-testis antigens of MAGE-A6 and MAGE-A11 is associated with tumor aggressiveness in patients with bladder cancer. Scientific Reports, 12(1), 1–16.
Ward, D. G., Baxter, L., Gordon, N. S., Ott, S., Savage, R. S., Beggs, A. D., et al. (2016). Multiplex PCR and next generation sequencing for the non-invasive detection of bladder cancer. PLoS One, 11(2), e0149756.
Pardini, B., Cordero, F., Naccarati, A., Viberti, C., Birolo, G., Oderda, M., et al. (2018). microRNA profiles in urine by next-generation sequencing can stratify bladder cancer subtypes. Oncotarget, 9(29), 20658.
Speck-Planche, A., Kleandrova, V. V., Luan, F., & Cordeiro, M. N. D. S. (2013). Unified multi-target approach for the rational in silico design of anti-bladder cancer agents. Anti-Cancer Agents in Medicinal Chemistry, 13(5), 791–800.
Robertson, A. G., Kim, J., Al-Ahmadie, H., Bellmunt, J., Guo, G., Cherniack, A. D., et al. (2017). Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell, 171(3), 540–56.e25. https://doi.org/10.1016/j.cell.2017.09.007
Weinstein, J. N., Collisson, E. A., Mills, G. B., Shaw, K. R., Ozenberger, B. A., Ellrott, K., et al. (2013). The cancer genome atlas pan-cancer analysis project. Nature Genetics, 45(10), 1113–20.
Ware, A. P., Kabekkodu, S. P., Chawla, A., Paul, B., & Satyamoorthy, K. (2022). Diagnostic and prognostic potential clustered miRNAs in bladder cancer. 3 Biotech, 12(8), 1–15.
Milan, T., & Wilhelm, B. T. (2017). Mining cancer transcriptomes: Bioinformatic tools and the remaining challenges. Molecular Diagnosis & Therapy., 21(3), 249–258. https://doi.org/10.1007/s40291-017-0264-1
Ahn, J.-H., Kang, C.-K., Kim, E.-M., Kim, A.-R., & Kim, A. (2022). Proteomics for early detection of non-muscle-invasive bladder cancer: Clinically useful urine protein biomarkers. Life (Basel, Switzerland), 12(3), 395.
Zhang, C., Berndt-Paetz, M., & Neuhaus, J. (2020). Identification of key biomarkers in bladder cancer: Evidence from a bioinformatics analysis. Diagnostics (Basel), 10(2), 66.
Zhang, Y., Fang, L., Zang, Y., & Xu, Z. (2018). Identification of core genes and key pathways via integrated analysis of gene expression and DNA methylation profiles in bladder cancer. Medical Science Monitor: International Medical Journal of Experimental and Clinical Research, 24, 3024.
Oliveira, MCd., Caires, H. R., Oliveira, M. J., Fraga, A., Vasconcelos, M. H., & Ribeiro, R. (2020). Urinary biomarkers in bladder cancer: Where do we stand and potential role of extracellular vesicles. Cancers (Basel), 12(6), 1400.
Liu, Y.-R., Ortiz-Bonilla, C. J., & Lee, Y.-F. (2018). Extracellular vesicles in bladder cancer: Biomarkers and beyond. International Journal of Molecular Sciences, 19(9), 2822.
Song, Y., Jin, D., Ou, N., Luo, Z., Chen, G., Chen, J., et al. (2020). Gene expression profiles identified novel urine biomarkers for diagnosis and prognosis of high-grade bladder urothelial carcinoma. Frontiers in Oncology, 10, 394.
Wang, J., Guo, M., Zhou, X., Ding, Z., Chen, X., Jiao, Y., et al. (2020). Angiogenesis related gene expression significantly associated with the prognostic role of an urothelial bladder carcinoma. Translational Andrology and Urology, 9(5), 2200.
Chang, C., Worley, B. L., Phaëton, R., & Hempel, N. (2020). Extracellular glutathione peroxidase GPx3 and its role in cancer. Cancers (Basel), 12(8), 2197.
Reszka, E., Lesicka, M., Wieczorek, E., Jabłońska, E., Janasik, B., Stępnik, M., et al. (2020). Dysregulation of redox status in urinary bladder cancer patients. Cancers (Basel), 12(5), 1296.
Zhou, S., Liu, R., Yuan, K., Yi, T., Zhao, X., Huang, C., et al. (2013). Proteomics analysis of tumor microenvironment: Implications of metabolic and oxidative stresses in tumorigenesis. Mass Spectrometry Reviews, 32(4), 267–311.
Zhang, L., Lv, B., Shi, X., & Gao, G. (2020). High expression of N-acetylgalactosaminyltransferase 1 (GALNT1) associated with invasion, metastasis, and proliferation in osteosarcoma. Medical Science Monitor, 26, e927837-1.
Perez, A., Loizaga, A., Arceo, R., Lacasa, I., Rabade, A., Zorroza, K., et al. (2014). A pilot study on the potential of RNA-associated to urinary vesicles as a suitable non-invasive source for diagnostic purposes in bladder cancer. Cancers (Basel), 6(1), 179–92.
Urabe, F., Kimura, T., Ito, K., Yamamoto, Y., Tsuzuki, S., Miki, J., et al. (2021). Urinary extracellular vesicles: A rising star in bladder cancer management. Translational Andrology and Urology, 10(4), 1878.
Ruan, H., Wang, T., Yang, C., Jin, G., Gu, D., Deng, X., et al. (2016). Co-expression of LASS2 and TGF-β1 predicts poor prognosis in hepatocellular carcinoma. Scientific Reports, 6(1), 1–10.
Wang, H., Wang, J., Zuo, Y., Ding, M., Yan, R., Yang, D., et al. (2012). Expression and prognostic significance of a new tumor metastasis suppressor gene LASS2 in human bladder carcinoma. Medical Oncology, 29(3), 1921–7.
Wang, H., Zuo, Y., Ding, M., Ke, C., Yan, R., Zhan, H., et al. (2017). LASS2 inhibits growth and invasion of bladder cancer by regulating ATPase activity. Oncology Letters, 13(2), 661–8.
Author information
Authors and Affiliations
Contributions
RR: funding acquisition, in silico and in vivo experiments, software, data analysis. HW: investigation, data curation, formal analysis. LX: methodology, project administration; SM: conceptualization, editing the manuscript. XY: funding, project administration, supervision drafts the manuscript.
Corresponding author
Ethics declarations
Ethical Approval
This study involved animals and ethics approved by the Department of Urology, Third Hospital of Shanxi Medical University Hospital, China, with ethics number YXLL-2021-066.
Consent to Participate
All authors have their consent to participate.
Consent for Publication
All authors have their consent to publish their work.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Figure S1
Tissue specificity of differentially expressed genes. A) GALNT1, B) ECM1, C) CERS2 and D) GPX3. (PNG 152 kb)

Figure S2
Normalized expression of selected differential marker genes (CRYAB, ALDOB, IGFBP7, RHCG and TMEM176A) in Bladder cancer. A) mRNA expression pattern of CRYAB, ALDOB, IGFBP7, RHCG and TMEM176A marker genes (as a box-whisker plot) in normal and primary tumor samples. B) Expression pattern of the top marker genes based on tumor subgroups based on race. (PNG 49 kb)

Figure S3
A) Molecular alterations for top differential expressed genes. B) Copy number variations of the DEGs. (PNG 93 kb)

Figure S4
mRNA expression profile in bladder cancer and other cancer types. Pan cancer gene expression profile of A) CRYAB, B) ALDOB, C) IGFBP7, and D) TMEM176A. (PNG 69 kb)

Figure S5
Epigenetic analysis of marker genes in BLCA. A) Methylation pattern of the top marker genes (CRYAB, IGFBP7, ALDOB and TMEM176A) in normal and primary tumor samples. B) Expression pattern of the CRYAB, IGFBP7, ALDOB and TMEM176A marker genes based on tumor subgroups based on race. (PNG 46 kb)

Figure S6
Survival analysis of marker genes in BLCA. A) The Kaplan–Meier plotters based on bladder cancer datasets showed patients with high CRYAB expression have poor overall survival, B) IGFBP7, C) ALDOB and D) TMEM176A. (PNG 47 kb)
Table S1
Supplementary Material S1 (3.19 MB xls)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ren, R., Wang, H., Xie, L. et al. Identify Potential Urine Biomarkers for Bladder Cancer Prognosis Using NGS Data Analysis and Experimental Validation. Appl Biochem Biotechnol 195, 2947–2964 (2023). https://doi.org/10.1007/s12010-022-04234-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-022-04234-7