Abstract
Staphylococcus aureus is a major cause of hospital-associated infections worldwide. The organism’s ability to form biofilms has led to resistance against current treatment options such as beta-lactams, glycopeptides, and daptomycin. The ArlRS two-component system is a crucial regulatory system necessary for S. aureus autolysis, biofilm formation, capsule synthesis, and virulence. This study aims to investigate the role of the arlR deletion mutant in the detection and activation of S. aureus. We created an arlR deleted mutant and complementary strains and characterized their impact on the strains using partial growth measurement. The quantitative real-time PCR was performed to determine the expression of icaA, and the microscopic images of adherent cells were captured at the optical density of 600 to determine the primary bacterial adhesion. The biofilm formation assay was utilized to investigate the number of adherent cells using crystal violet staining. Eventually, the Triton X-100 autolysis assay was used to determine the influence of arlR on the cell autolytic activities. Our findings indicate that the deletion of arlR reduced the transcriptional expression of icaA but not icaR in the ica operon, leading to decrease in polysaccharide intercellular adhesin (PIA) synthesis. Compared to the wild-type and the complementary mutants, the arlR mutant exhibited decreased in biofilm production but increased autolysis. It concluded that the S. aureus response regulatory ArlR influences biofilm formation, agglutination, and autolysis. This work has significantly expanded our knowledge of the ArlRS two-component regulatory system and could aid in the development of novel antimicrobial strategies against S. aureus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococcus aureus, also known as golden staph, is a gram-positive bacterium categorized as one of the ESKAPE pathogens, the highly virulent antibiotic-resistant bacterial strains, resulting in significant morbidity and mortality rates worldwide [1]. S. aureus can cause a variety of infections ranging from mild to severe, including blood septicemia, thrombophlebitis, infectious endocarditis [2,3,4] osteomyelitis, or device-related infections (DRI) [5], such as intravascular catheters, prosthetic joints, vascular grafts, and pacemakers [6].
Staphylococcus aureus has emerged as a leading cause of hospital-associated infections and the most life-threatening multidrug-resistant opportunistic pathogens worldwide [7, 8]. Methicillin-resistant S. aureus (MRSA), also known as oxacillin-resistant S. aureus, is the most virulent form of S. aureus, which has reached a particular epidemic and healthcare threat globally [9, 10]. MRSA is defined as an oxacillin minimum inhibitory concentration (MIC) greater than or equal to 4 mg/L [11]. MRSA has developed resistance to current treatment options such as -lactams, glycopeptides, and daptomycin due to the organism’s capacity to produce biofilms [12, 13], commonly known as the two-component regulatory system (TCS), which functions as a sensory system for S. aureus [14]. The success of S. aureus in invading humans highly depends on various virulence factors, including exoenzymes, toxins, and adhesions that make the bacterial adhere to the mucous membranes or invade the immune system and adapt to dynamic environmental stimuli to survive [15, 16]. Multiple virulence factors are controlled by the two-component regulatory system (TCS). Most of the S. aureus strains possess 16 TCSs, one of which is vital for bacterial viability [17, 18].
The ArlRS TCS has been involved in various pathogenic processes, such as autolysis, biofilm formation, virulence, and capsule synthesis, which plays a vital role in the regulation of clumping and adherence [19, 20]. ArlRS consists of a membrane-bound histidine kinase ArlS that responds to different environmental signals, auto-phosphorylates a conserved histidine residue, and a cognate response regulator ArlR which its aspartate residue receives the phosphate and affects the transcription of a subset of target genes [21, 22]. The arlR was found to activate the production of MgrA responsible to controls the virulence factors and the intracellular adhesion gene cluster (ica operon) [23, 24]. Additionally, it serves as a direct activator for abcA, a gene that encodes an ATP-dependent transporter associated with cell wall autolysis and -lactam antibiotics resistance [25]. ArlR-MgrA directly controls the cascade that represses the biofilm formation regulator Rbf and upregulates SarX, a Sar transcriptional regulatory protein family member that regulates ica expression [26]. It has been reported that MgrA knockout in USA300 resulted in just a slight reduction of MIC for oxacillin (from 64 to 32 mg/L), but arlRS knockout in USA300 results in a significantly decreased MIC value (4 mg/L) [10]. Moreover, the arlR mutant has an essential clumping defect for staphylococcal infection. Polysaccharide intercellular adhesion (PIA) is a common element of staphylococcal biofilm [27, 28]. The ArlR plays a vital role in regulating biofilm formation, which can directly bind to the promoter region between icaR and icaADBC, enhancing the expression of icaA and PIA production [29].
The synthesis of PIA is performed via the ica locus, which has an operon composed of four open frames named icaA, icaD, icaB, and icaC. The icaR is likewise a component of the ica operon and is located upstream but in the opposite location [30, 31]. Our previous study showed structural insights into the activation and recognition of ArlR, and ArlR recognizes a 20 bp imperfect inverted repeat sequence located at the ica operon [32]. Nonetheless, the downstream gene regulatory mechanisms of arlR in S. aureus are poorly studied. In this research, we have introduced a novel approach to investigate the role of the arlR deletion mutant in detecting and activating S. aureus. We have created the strains mutant and complementary mutant, evaluated the effects of the arlR in the growth measurement, determined the influence of arlR on the cell autolytic activities, and assessed the intracellular adhesion cells agglutination. This study provides an in-depth understanding of the ArlR response regulator and its functions, offering unique opportunities to combat antibiotic resistance and directly target pathogen virulence. The outcomes of this study may also pave the way for further research into targeting the ArlR regulatory system as a novel target for fighting S. aureus.
Materials and Methods
Bacterial Strains and Primers
The bacterial strains used in this study are listed in Table 1. S.aureus were cultured in Trypticase soy broth (TSB) (Oxoid) at 37 °C. Escherichia coli cells were cultured in Luria-Bertani (LB) (Oxoid) medium. Cloning of S. aureus chromosomal DNA fragments was performed in Escherichia coli stains DH5α using plasmid PKOR1 and vector pLi50 [33].
Construction of the arlR Mutant Strains
A 1000 bp upstream and downstream region of the arlR were amplified by homologous recombinant form S. aureus genome DNA using BIO-RAD C1000 Touch PCR thermal cycler, ligated to temperature-sensitive shuttle plasmid PKOR1 through fusion PCR and Gateway cloning technology using primers. The recombinant plasmid was extracted and transformed into a restriction system-defective S. aureus RN4220 by electroporation. The target strain was electrophoresed with a modified plasmid. Positive colonies were broth incubated overnight at 30 °C with 2 mL TSB containing 10 µg/mL chloramphenicol (Sigma). The strains were diluted by a ratio of 1:100 into a total volume of 50 mL TSB containing 10 µg/mL chloramphenicol and incubated overnight at 42 °C. The cultivation process was repeated twice. The cultures were diluted before plating 100 µL on TSB plate (10 µg/mL chloramphenicol) by incubating overnight at 42 °C. A single colony was resuspended with 2 mL TSB and incubated overnight at 30 °C without any additive antibiotics to facilitate plasmid excision. This culture’s 100 folded serial dilutions were spread on the TSB containing 1 µg/mL tetracycline (Sigma) and incubated for 24 h at 37 °C. The transformants were screened for chloramphenicol-sensitive colonies, which grown on the TSB plates not on TSB plate containing 10 µg/mL tetracycline. A single colony was isolated and re-cultured on the TSB plates and then inoculated into 5 mL TSB and incubated overnight with continuous shaking at 37 °C. The deleted mutants were identified by PCR using the primers attB1-UP-F and attB2-Down-R.
Creation of the arlR Complement
The arlR gene pair and its native promoter were amplified with the primer pairs of parlR-Kpn1-F and parlR-HindIII-R using S. aureus genome DNA. The PCR product was gel-purified and digested by KpnI and HindIII at 37 °C overnight. The digested fragment was cloned into vector pLi50 and then introduced into E.coli DH5α to generate the plasmid pLi50-arlR. The restriction defective RN4220 was electroporated by plasmid after genetic modification in RN4220. The recombinant plasmid was transduced into the S. aureus △arlR.
Quantitative PCR
For qPCR experiments, arlR deleted mutant and complementary strains were activated using 5 mL TSB and incubating overnight at 37 °C. Following incubation, 50 µL cultures were transferred to 50 mL TSB and incubated at 37 °C for 12 h then lysed for 30 min at 37 °C with 100 µL lysostaphin (Sigma). The total RNA was extracted by Qiagen RNeasy Mini kit, and 300 ng of RNA was verified by the A260/A280 ratio of 2.0–2.1. Then the high-capacity cDNA reverse transcription kit (Bio-Rad) was used for the first-strand cDNA synthesis. The qPCR experiments were performed using the Power SYBR Green PCR Master Mix (Bio-Rad) to determine the expression of icaA via Bio-Rad CFX Connect Real-Time PCR Detection System by following conditions: 95 °C for 10 min, 95 °C for 10 s, 60 °C for 15 s, and 72 °C for 20 s with 39 cycles. All qRT-PCR primers used for the qRT-PCR (Table 2) were verified using normal PCR. 16 S-RNA was used as a reference gene to standardize all the data.
Growth Curves of arlR Wild-Type, Delete Mutant, and Complementation Strains
The strains were incubated overnight with 5 mL of TSB at 37 °C before being diluted 1:100. Using the TSB medium to adjust an OD600 to 0.15, cultures were grown at 37 °C with 200 rpm of continuous shaking. The optical density was determined by measuring the OD600 every 30 min for 10 h.
Scanning Electron Microscopy (SEM) Images
The bacterial cultures were grown in 5 mL TSB at 37 °C overnight and then were diluted 1:100 with TSB in 24-well plates (Beyotime, China). After incubation at 37 °C for 24 h without shaking, the catheter fragments were washed for 5 min with PBS three times. The strains were then fixed for 20 min with 4% paraformaldehyde (Sigma), followed by a secondary immobilization with 2.5% glutaraldehyde (Sigma). After overnight incubation, samples were washed three times with phosphate buffers and dehydrated in 30%, 50%, 70%, 80%, 90%, 95%, or 100% ethanol (Sigma) for at least 10 min each. Eventually, the samples were dried, ion-sputter gilded, and then captured using an FEI Quanta FEG 250 SEM with a magnification of x5000 and x20000 times [34].
Adherence of arlR Delete Mutant and Complementation
The bacterial primary adhesion ability determination was used as previously described in the literature [35] after being modified to meet our study conditions. The overnight-grown strains were inoculated in TSB to the early stationary phase and were centrifuged at 4000 rpm. The bacterial suspensions were adjusted with PBS to an optical density at OD600 of 0.1, and 5 mL of each suspension was put on 6-well plates (Beyotime, China) and incubated at 37 °C for 30 min. Each plate was washed five times with PBS, and microscope images of adherent cells were captured (1000 folds magnification).
Biofilm Formation Assay
The overnight-grown bacterial cultures were adjusted 1:200 in TSB, transferred to 96-well polymer plates (Beyotime, China), and incubated for 24 h at 37 °C. The Petri dishes were gently washed thrice with PBS, fixed for 15 min with 200 µL of 99% methyl alcohol, and dried. The plates were stained with 2% crystal violet for 5 min before being measured at an OD570 wavelength.
Triton X-100-Induced Autolysis Assays
The strain’s autolysis assays were performed as previously described in the literature [28]. Briefly, overnight-grown bacterial cultures were diluted 1:100 in TSB containing 1 M of NaCl (Sigma) and cultivated at 37 °C with continuous shaking until OD580 reached approximately 0.7. The cells were centrifuged at 13,800 g at 4 °C for 10 min and washed twice with 50 mL ice-cold water. Then, the cells were resuspended in the same volume containing 50 mM Tris-HCl (PH 7.5) and 0.1% Triton X-100 (Sigma). Bacterial strains were incubated at 30 °C with continuous shaking for 3–5 h, while the OD580 was measured every 30 min.
Results
Construction of an arlR Delete Mutant Strain of S. aureus
A previous study showed the influence of the locus on autolysis, adhesion, and extracellular proteolytic activity in arlS mutants [19]. An arlR deleted mutant was created to investigate virulence gene regulation, in which the gene of arlR (NZ_CP015758) was deleted. The mutation (△arlR) leads to the deletion of 219 amino acids from the predicted protein (Fig. 1a). The experimental procedures were performed as follows (Fig. 1b).
PCR validation of PKOR1 allelic replacement. a The primers (attB1-up-F and attB2–down-R) are shown as short black arrows. arlR is indicated by an arrow box, while the PCR products used for deletion of arlR were shown as boxes with 1 kb size. b Schematic chart for the construction of the arlR deleted mutant
The Growth Curve of Strains
The arlR mutant strain was used to study the influence of arlR mutation on the growth rate, autolysis, and biofilm formation. The growth curve of the strains was cultured under the same condition and measured using a spectrophotometer at an optical density of 600 (OD600). Both the pLi50 and the arlR mutant adopted comparable growth rate patterns. Contrary, the growth rate behavior of the complementary strain was somehow similar to the wild-type strain; however, it was obvious that the wild-type strain’s growth rate was rapidly rising during the mid-exponential phase, as demonstrated in Fig. 2a. After six h of incubation, all four strains reached the stationary growth phase. Unlike the yellowish colonies of the wild-type and complementary strains, the deleted-mutant colonies were white-colored (Fig. 2b).
Effects of arlR knockout on S. aureus growth. a Growth curves of S. aureus strains. Overnight cultures were diluted to OD600 = 0.15 as the start point by TSB and incubated at 37 °C with shaking. The value of OD600 was measured every 30 min for about 8 h. The experiment was repeated thrice, and the representative data was shown. b Bacterial morphology. Overnight cultures were diluted to 1:10000 with fresh TSB after incubating on TSB plates at 37 °C for 24 h
Real-Time qPCR
As previously mentioned by Yurong Wen et al., arlR can bind to the promoter region between icaR and icaADBC, then regulate the gene expression of icaADBC [32]. The impact of arlR on icaADBC and icaR in S. aureus biofilm formation and adhesive ability was first executed via the real-time qPCR. Transcript levels of icaA and icaR were examined at the stationary phase of the wild-type mutant and complement strains. The transcript expression of icaA was reduced almost ten-fold in the complementary strains compared to the wild-type strains (Fig. 3a). Although the transcriptional level of icaR did not differ significantly, a minor decrease was seen in the arlR mutation but neither in the wild-type nor complementary strains (Fig. 3b). The housekeeping gene 16 S rRNA was used as an endogenous factor to standardize all data. These results demonstrated a reduction in the icaA expression of the arlR mutation, which was correlated with the adherence and the agglutination. Nevertheless, the icaR expression was not affected. The results are consistent with the EMSA results in our previous study, which was activated ArlR recognizes a 20-bp AT-rich inverted repeat [32].
Adherence and Biofilm Formation Assay
A previous study showed intercellular adherence and accumulation of poly-laminate cell clumps that lead to biofilm formation, which could be measured by crystal violet staining [36]. The attached bacterial cells of wild-type strains showed that the number of adherent cells was significantly higher than the corresponding mutants, with a minor variation between the wild-type and the complementary strains (Fig. 4a). Following incubation of the strains in polystyrene plates, the arlR delete mutant appearance (middle panel) had a distinct pattern with the wild-type (upper panel) and complementary strains (lower panel). In contrast to the wild-type strain, the arlR delete mutant strain exhibited a negligible affinity for the polystyrene surface, resulting in a minimal adhesion of cell density as measured by 2% crystal violet staining once the biofilm formed on the bacterial surface. The cell density of complementary strains was similar to that of the wild-type (Fig. 4b). Since cell agglutination is an indicator of the strain’s primary attachment and the accumulation for the multilayered cell clusters; therefore, the strains were observed using a spectrophotometer at OD570, the wild-type and complementary strains produced several aggregates, and intercellular adhesion significantly decreased in the deleted mutant (Fig. 4c). The arlR can reinforce the PIA production, and the PIA is necessary for biofilm and aggregate formation [29, 37]. To find out how arlR deleted mutant influences the icaADBC, which is essential for PIA biosynthesis, we further study the gene expression levels of these factors.
Adherence and biofilm formation capacities of S. aureus. a Overnight cultures were diluted to 1:100 in 50 mL TSB and grown to the early stationary phase. The adherent cell images were captured using the microscope, with a magnification power of 50× in the upper line and 250× in the lower line. The black boxes in the upper line reflect the regions magnified in the lower line. b Overnight cells were diluted by 1:200 with fresh TSB, added into 96-well polystyrene plates, and incubated for 24 h at 37 °C. The biofilms were fixed in 99% methyl alcohol and stained with crystal violet for observation. c The bacterial cells were stained with crystal violet, and the quantitative assay of the biofilm formation capacity determined the absorbance at 570 nm. The results of a representative experiment are shown as (n = 30; p < 0.001), and the error bars represent the standard deviation (SD)
Triton X-100-Induced Autolysis in arlR Mutant
Autolysis plays a vital role in the bacterial cell wall. S. aureus has multiple known or putative autolysin genes, including lytM, lytN, and atl; the expression of autolysins is tightly controlled by several negative autolysis regulators, such as arlRS [38]. Aiming to determine whether the mutation leads to an increase in autolysis, we examined the contribution of arlR on the cell autolytic activities using Triton X-100. The deleted mutant enhanced the cell lysis level compared to the wild-type strain, while the complementary strain’s lysis behaved similarly to the wild-type one (Fig. 5).
SEM Images of Biofilms
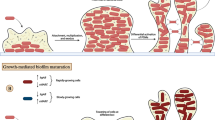
S. aureus has been reported to bind with human matrix proteins and form stable clumps in the agglutination process [20]. To further evaluate the clumps-difference between the wild-type, deleted mutant, and complementary strains, the strains were incubated for 24 h at 37 °C to ensure biofilm formation. Scanning electron microscopy (SEM) images indicated a microstructural level of cell morphologies in which the wild-type strains developed a compact, thick biofilm. In contrast, the mutant formed just a few bacterial cell clusters and had clumping defects (Fig. 6a and b). With a 20 K magnification, we observed the cross-links between the wild-type bacterial cells (Fig. 6d and e), which led to an abnormal clump formation. pLi50-arlR developed a biofilm similar to wild-type strains (Fig. 6d and f).
Discussion
As concerns increase regarding antibiotics bacterial resistance and broad-spectrum antibiotics destroying the beneficial microflora, directly targeting the signaling pathways that regulate pathogen virulence emerges as a promising alternative strategy. Biofilm formation has been recognized as a virulence factor attributing the pathogenesis and infections [1, 39]. Despite its clinical significance, little is known about the expression and regulation of S. aureus biofilms. Several S. aureus genes, such as ica, arlRS, and sarA, have been identified as crucial for biofilm formation. Ica ADBC-encoded proteins biosynthesize the PIA, which is essential for S. aureus and Staphylococcus epidermidis biofilm formation [40, 41]. The icaADBC operon plays a vital role in biofilm formation, which is considered a major invasive factor associated with agglutination and pathogenesis [37]. IcaR, located adjacent to the ica operon, encodes a transcriptional repressor associated with the ica operon expression in S. aureus [42].
The response regulator ArlR can bind to the promoter region of the ica operon and regulate the expression of ica operon and PIA production [14, 43]. Our previous study elucidated the ArlR-binding site located 11 bp upstream of the − 10 sequence, indicating that they are regulated by ArlR directly [32]. This study revealed that the S. aureus response-regulatory ArlR influences biofilm formation, agglutination, and autolysis. The result of the biofilm formation assay demonstrated that the deletion of arlR has reduced biofilm formation and decreased PIA production (Fig. 7). In S. epidermidis, arlR could bind with the icaA promoter region and the upstream of icaR, negatively regulating biofilm formation in an ica-dependent manner [44].
The qRT-PCR outcomes revealed that the icaR transcriptional level has slightly changed compared to the wild-type strains. Meanwhile, the gene expression of icaA has significantly decreased in arlR mutants. Triton X-100-induced autolysis showed an increased cell lysis rate in the arlR mutant compared to the wild-type and complementary strains. We hypothesize that arlR mutations mediate the adherence to polystyrene surfaces and reduce cell clusters by inducing autolysins expression. The SEM images of clumps demonstrated that arlR regulated the agglutination on the plastic surfaces. It has been reported that several regulatory factors, including MgrA, Rbf, SigB, TcaR, SrrAB, and Spx, affect ica expression, and its regulation is extremely complicated [16, 45].
This study confirmed ArlR’s involvement in the physiological processes of biofilm formation, agglutination, and autolysis in S. aureus. However, it did not extensively investigate the molecular-level regulatory mechanism of ArlR, including its response to external signals and the specific genes involved in biofilm formation, agglutination, and autolysis. Although previous studies and this research have demonstrated the direct binding and regulation of icaABC by ArlR, future investigations will focus on elucidating the signaling mechanism of the ArlRS two-component regulatory system and uncovering additional downstream regulatory genes.
Conclusion
To conclude, the current study discussed the role of the response regulator ArlR in S. aureus biofilm formation, agglutination, and autolysis. Biofilm formation is a crucial factor contributing to the pathogeneses and infections caused by S. aureus. The expression and regulation of S. aureus biofilms are not yet fully understood, but the ica, arlRS, and sarA genes have been identified as key players in biofilm formation. The icaADBC operon, responsible for polysaccharide intercellular adhesin (PIA) synthesis, plays a vital role in S. aureus and S. epidermidis biofilm formation. The transcriptional repressor IcaR, located adjacent to the ica operon, is associated with the expression of the ica operon in S. aureus. The response regulator ArlR has been found to bind to the promoter region of the ica operon, directly regulating its expression and PIA production.
The findings of this study indicate that the deletion of arlR resulted in reduced biofilm formation and decreased PIA production. Additionally, the arlR mutant showed increased autolysis compared to the wild-type and complementary strains. It is hypothesized that arlR mutations mediate adherence to surfaces and reduce cell clusters by inducing the expression of autolysins. Furthermore, arlR was found to regulate agglutination on plastic surfaces. Understanding the mechanisms and regulatory factors involved in biofilm formation is crucial for developing alternative strategies to combat antibiotic resistance and target pathogen virulence directly. Further research is warranted to unravel the complex regulation of biofilm formation and explore the potential of ArlR as a therapeutic target in combating S. aureus infections.
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- S. aureus :
-
Staphylococcus aureus
- ESKAPE:
-
Enterococcus faecium, Staphylococcus aureus, Klebsiella. pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species
- MRSA:
-
Methicillin-resistant Staphylococcus aureus
- MIC:
-
Minimum inhibitory concentration
- TCS:
-
Two-component regulatory system
- MgrA:
-
Multiple antibiotic resistance regulator A
- Ica:
-
Intracellular adhesion
- PIA:
-
Polysaccharide intercellular adhesin
- USA300:
-
A widely spread community-associated MRSA strain
- Rbf:
-
Repressor of biofilm formation
- SarX:
-
Sar transcriptional regulatory protein X
- SigB:
-
Sigma factor B
- SrrAB:
-
Sensor kinase/response regulator system SrrAB
- Spx:
-
Global transcriptional regulator Spx
- TcaR:
-
Transcriptional regulator of the tca operon
References
Venkateswaran, P., Vasudevan, S., David, H., Shaktivel, A., Shanmugam, K., Neelakantan, P., & Solomon, A. P. (2023). Revisiting ESKAPE pathogens: Virulence, resistance, and combating strategies focusing on quorum sensing. Frontiers in Cellular and Infection Microbiology, 13, 1159798. https://doi.org/10.3389/fcimb.2023.1159798
Zhou, K., et al. (2018). A review on nanosystems as an effective approach against infections of Staphylococcus aureus. International Journal of Nanomedicine, 13, 7333–7347. https://doi.org/10.2147/IJN.S169935
Mottola, C., et al. (2016). Susceptibility patterns of Staphylococcus aureus biofilms in diabetic foot infections. BMC Microbiology, 16(1), 119. https://doi.org/10.1186/s12866-016-0737-0
Parente, D. M., Cunha, C. B., Mylonakis, E., & Timbrook, T. T. (Jun. 2018). The clinical utility of Methicillin-Resistant Staphylococcus aureus (MRSA) nasal screening to rule out MRSA Pneumonia: A diagnostic Meta-analysis with antimicrobial stewardship implications. Clinical Infectious Diseases, 67(1), 1–7. https://doi.org/10.1093/cid/ciy024
Tong, S. Y. C., Davis, J. S., Eichenberger, E., Holland, T. L., & Fowler, V. G. (2015). Staphylococcus aureus infections: Epidemiology, pathophysiology, clinical manifestations, and management. Clinical Microbiology Reviews, 28(3), 603–661. https://doi.org/10.1128/CMR.00134-14
Ellis, M. W. (2014). Prevalence of nasal colonization and strain concordance in patients with community-associated Staphylococcus aureus skin and soft-tissue infections, Infection Control & Hospital Epidemiology, 35, 1251–1256.
Deleo, F. R., & Chambers, H. F. (2009). Reemergence of antibiotic-resistant. JCI The Journal of Clinical Investigation, 119(9), 2464–2474. https://doi.org/10.1172/JCI38226.2464
Hassoun, A., Linden, P. K., & Friedman, B. (2017). Incidence, prevalence, and management of MRSA bacteremia across patient populations-A review of recent developments in MRSA management and treatment. Critical Care, 21(1), 1–10.
Yadav, R., Kumar, A., Singh, V. K., Jayshree, & Yadav, S. K. (2018). Prevalence and antibiotyping of Staphylococcus aureus and methicillin-resistant S. Aureus (MRSA) in domestic animals in India. Journal of Global Antimicrobial Resistance, 15, 222–225. https://doi.org/10.1016/j.jgar.2018.08.001
Bai, J., et al. (2019). The role of ArlRS in regulating oxacillin susceptibility in methicillin-resistant Staphylococcus aureus indicates it is a potential target for antimicrobial resistance breakers. Emerging Microbes & Infections, 8(1), 503–515. https://doi.org/10.1080/22221751.2019.1595984
Rybak, M. J., & Akins, R. L. (2001). Emergence of methicillin-resistant Staphylococcus aureus with intermediate glycopeptide resistance. Drugs, 61, 1–7. https://doi.org/10.2165/00003495-200161010-00001
Hsu, L. Y., Wijaya, L., & Koh, T. H. (2010). Community-associated meticillin-resistant Staphylococcus aureus. The Lancet, 376, 767. https://doi.org/10.1016/S0140-6736(10)61369-4
Guo, Y., Song, G., Sun, M., Wang, J., & Wang, Y. (2020). Prevalence and therapies of antibiotic-resistance in Staphylococcus aureus. Frontiers in Cellular and Infection Microbiology, 10, 107.
Burgui, S., Gil, C., Solano, C., Lasa, I., & Valle, J. (2018). A systematic evaluation of the two-component systems network reveals that ArlRS is a key regulator of catheter colonization by Staphylococcus aureus. Frontiers in Microbiology, 9, 1–11. https://doi.org/10.3389/fmicb.2018.00342
Capra, E. J., & Laub, M. T. (2012). The evolution of two-component. Annual Review of Microbiology, 66, 325–347. https://doi.org/10.1146/annurev-micro-092611-150039.The
Schilcher, K., & Horswill, A. R. (2020). Staphylococcal biofilm development: Structure, regulation, and treatment strategies. Microbiology and Molecular Biology Reviews, 84(3), 10–1128.
Qiu, Y., et al. (2021). Five major two components systems of Staphylococcus aureus for adaptation in diverse hostile environment. Microbial Pathogenesis, 159, 105119.
Haag, A. F., & Bagnoli, F. (2017). The role of two-component signal transduction systems in Staphylococcus aureus virulence regulation, Staphylococcus aureus: Microbiology, Pathology, Immunology, Therapy and Prophylaxis (pp. 145–198).
Párraga Solórzano, P. K., Yao, J., Rock, C. O., & Kehl-Fiea, T. E. (2019). Disruption of glycolysis by nutritional immunity activates a two-component system that coordinates a metabolic and antihost response by staphylococcus aureus. MBio, 10(4), 1–15. https://doi.org/10.1128/mBio.01321-19
Walker, J. N., et al. (2013). The Staphylococcus aureus ArlRS two-component system is a novel regulator of agglutination and pathogenesis. Plos Pathogens, 9, 1–17. https://doi.org/10.1371/journal.ppat.1003819
Fournier, B., & Hooper, D. C. (2000). A new two-component regulatory system involved in adhesion, autolysis, and extracellular proteolytic activity of Staphylococcus aureus. Journal of Bacteriology, 182, 3955–3964. https://doi.org/10.1128/JB.182.14.3955-3964.2000
Crosby, H. A., et al. (2020). The Staphylococcus aureus ArlRS two-component system regulates virulence factor expression through MgrA. Molecular Microbiology, 113(1), 103–122. https://doi.org/10.1111/mmi.14404
Crosby, H. A., Schlievert, P. M., Merriman, J. A., King, J. M., Salgado-Pabón, W., & Horswill, A. R. (2016). The Staphylococcus aureus global regulator MgrA modulates clumping and virulence by controlling surface protein expression. Plos Pathogens, 12(5), 1–31. https://doi.org/10.1371/journal.ppat.1005604
Kwiecinski, J. M. (2019). Staphylococcus aureus adhesion in endovascular infections is controlled by the arlRS–MgrA signaling cascade, vol. 15, no. 5. https://doi.org/10.1371/journal.ppat.1007800
Villet, R. A., Truong-Bolduc, Q. C., Wang, Y., Estabrooks, Z., Medeiros, H., & Hooper, D. C. (2014). Regulation of expression of abcA and its response to environmental conditions. Journal of Bacteriology, 196(8), 1532–1539. https://doi.org/10.1128/JB.01406-13
Cue, D., Lei, M. G., & Lee, C. Y. (2013). Activation of sarX by Rbf is required for biofilm formation and icaADBC expression in Staphylococcus aureus. Journal of Bacteriology, 195(7), 1515–1524. https://doi.org/10.1128/JB.00012-13
Jenul, C., & Horswill, A. R. (2019). Regulation of Staphylococcus aureus virulence. Gram-Positive Pathogens, 6(1), 669–686. https://doi.org/10.1128/9781683670131.ch41
Wu, Y., et al. (2014). Role of the two-component regulatory system arlRS in ica operon and aap positive but non-biofilm-forming Staphylococcus epidermidis isolates from hospitalized patients. Microbial Pathogenesis, 76, 89–98. https://doi.org/10.1016/j.micpath.2014.09.013
Jin, Z., Jiang, Q., Fang, B., & Sun, B. (2019). The ArlR-MgrA regulatory cascade regulates PIA-dependent and protein-mediated biofilm formation in rbf-dependent and rbf-independent pathways. International Journal of Medical Microbiology, 309(2), 85–96. https://doi.org/10.1016/j.ijmm.2018.12.006
Büttner, H., Mack, D., & Rohde, H. (2015). Structural basis of Staphylococcus epidermidis biofilm formation: Mechanisms and molecular interactions. Frontiers in Cellular and Infection Microbiology, 5(FEB), 1–15. https://doi.org/10.3389/fcimb.2015.00014
Liang, X., Zheng, L., Landwehr, C., Lunsford, D., Holmes, D., & Ji, Y. (2005). Global regulation of gene expression by ArlRS, a two-component signal transduction regulatory system of Staphylococcus aureus. Journal of Bacteriology, 187(15), 5486–5492. https://doi.org/10.1128/JB.187.15.5486-5492.2005
Ouyang, Z., et al. (2019). Deciphering the activation and recognition mechanisms of Staphylococcus aureus response regulator ArlR. Nucleic Acids Research, 47, 11418–11429. https://doi.org/10.1093/nar/gkz891
Bae, T., & Schneewind, O. (2006). Allelic replacement in Staphylococcus aureus with inducible counter-selection. Plasmid, 55(1), 58–63. https://doi.org/10.1016/j.plasmid.2005.05.005
Wen, Y. (2017). Crystal structure of master biofilm regulator CsgD regulatory domain reveals an atypical receiver domain. Protein Science, 26(10), 2073–2082. https://doi.org/10.1002/pro.3245
Wu, Y., et al. (2012). The two-component signal transduction system ArlRS regulates Staphylococcus epidermidis biofilm formation in an ica-dependent manner. PLoS One, 7(7). https://doi.org/10.1371/journal.pone.0040041
Vuong, C., et al. (2004). Polysaccharide intercellular adhesin (PIA) protects Staphylococcus epidermidis against major components of the human innate immune system. Cellular Microbiology, 6(3), 269–275. https://doi.org/10.1046/j.1462-5822.2004.00367.x
Moormeier, D. E., & Bayles, K. W. (2017). Staphylococcus aureus biofilm: A complex developmental organism graphical abstract HHS public access. Molecular Microbiology, 104(3), 365–376. https://doi.org/10.1111/mmi.13634.Staphylococcus
Chunhua, M., et al. (2012). The expression of LytM is down-regulated by RNAIII in Staphylococcus aureus. Journal of Basic Microbiology, 52, 636–641. https://doi.org/10.1002/jobm.201100426
Beltrame, C. O., et al. (2015). Inactivation of the autolysis-related genes lrgB and yycI in staphylococcus aureus increases cell lysis-dependent eDNA release and enhances biofilm development in vitro and in vivo. PLoS One, 10(9), 1–20. https://doi.org/10.1371/journal.pone.0138924
Wu, S., Zhang, J., Peng, Q., Liu, Y., Lei, L., & Zhang, H. (2021). The role of Staphylococcus aureus YycFG in gene regulation, biofilm organization and drug resistance. Antibiotics, 10(12), 1555.
Hoang, T. M., et al. (2019). Transcriptional regulation of icaADBC by both IcaR and TcaR in Staphylococcus epidermidis. Journal of Bacteriology, 201(6), 10–1128.
Arciola, C. R., Campoccia, D., Ravaioli, S., & Montanaro, L. (2015). Polysaccharide intercellular adhesin in biofilm: Structural and regulatory aspects. Frontiers in Cellular and Infection Microbiology, 5(FEB), 1–10. https://doi.org/10.3389/fcimb.2015.00007
Trotonda, M. P., Tamber, S., Memmi, G., & Cheung, A. L. (2008). MgrA represses biofilm formation in Staphylococcus aureus. Infection and Immunity, 76(12), 5645–5654. https://doi.org/10.1128/IAI.00735-08
Yan, H., Wang, Q., Teng, M., & Li, X. (2019). The DNA-binding mechanism of the TCS response regulator ArlR from Staphylococcus aureus. Journal of Structural Biology, 208(3), 107388. https://doi.org/10.1016/j.jsb.2019.09.005
Patel, H., & Rawat, S. (2023). A genetic regulatory see-saw of biofilm and virulence in MRSA pathogenesis. Frontiers in Microbiology, 14, 1204428.
Funding
FZ is partially supported by the National Natural Science Foundation of China (No. 32170187), the Natural Science Foundation of Shaanxi Provence (No. 2021JM-007), and the Science Foundation of the Chinese Academy of Medical Sciences (2021-JKCS-008).
Author information
Authors and Affiliations
Contributions
Data curation, J Zhou and J Zhang; formal analysis, J Zhou; funding acquisition, F Zhang; investigation, Z Ouyang, Y Guo, J Zhang, M Jiao, W He, X He, and M Rabie; methodology, J Zhou, Z Ouyang, Y Guo, M Jiao, and W He; software, W He; supervision, F Zheng; writing—original draft, J Zhou, Z Ouyang, and F Zheng; writing—review and editing, M Refat, Z Ouyang, and F Zheng.
Corresponding authors
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Zhenlin Ouyang and Fang Zheng and share senior authorship of this manuscript.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, J., Refat, M., Guo, Y. et al. The Functional Study of Response Regulator ArlR Mutants in Staphylococcus Aureus. Appl Biochem Biotechnol (2024). https://doi.org/10.1007/s12010-024-04919-1
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-024-04919-1