Abstract
Generally, rapid detection of viral infection is necessary for preventing the virus from spreading among people in a society as a pandemic. Although there are many effective standard techniques used for virus identification, they are laborious, required skilled person to handle and time-consuming. Particularly, the detection of viral infection involved in the isolation and nucleic acid detection by collecting specimens (sample) from the appropriate sites. For instance, oral or nasal swab, nasopharyngeal or tracheal extract, lung tissue, blood, sputum and feces are collected in order to investigate the pandemic, COVID-19 for the effective and rapid diagnosis and eventually for the treatment. In this mini-review, it is summarized that the advanced testing methods which include RNA, immunologic and radiological based tests that could be used to detect COVID-19 and their cost, reliability and functionality are discussed in this review. This mini-review might help the researcher and health care sector to plan the diagnostic procedures as per the severity of the new infection, COVID-19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The COVID-19 is a contagious pandemic disease, rapidly spreading globally by a newly discovered coronavirus, SARS-CoV2. Coronaviruses are a broad group of viruses that can infect both animals and humans. The outbreak of pandemic originated in Wuhan city of China during 2019 and drew major attention and concern from the World Health Organization (WHO, 2020). Currently, COVID-19 has affected a large population in over 200 countries and territories around the world. Compared to other viral outbreaks such as SARS (2003) and MERS (2012), COVID-19 is being spread very rapidly, and entered a treacherous new phase and controlling the spread of COVID-19 is being a great hurdle. It is an emergency period for the world to show the scientific potential to find an appropriate and rapid diagnostic protocol to investigate the viral infection such as pandemic, COVID-19. Generally, before proceeding to diagnosis, a suspected individual consented with some questionnaire mentioning their travel history from an affected place or have contact with a person with the affected place and by observing the clinical presentation such as symptoms of COVID-19 [1]. Alternatively, chest X-ray revealed the bilateral infiltrates, but in early stage it is in normal condition. CT scan is more sensitive and specific because it showed infiltrates, segmental consolidation and peripheral or multifocal ground-glass opacities in both lungs of the affected person [2]. Invariably, a chest CT scan is used for the complement to RT-PCR to diagnose COVID-19. The schematic diagram represented showing the clinical sample collection and laboratory diagnosis of COVID-19 as per the Centre disease control and prevention (CDC) protocol (Fig. 1).
Surprisingly, about 1014 patients were tested with RT-PCR, and performance of CT scan was assessed, 580 patients showed positive results both RT-PCR and CT scan, 380 patients showed positive CT scan and negative RT-PCR results, 23 patients showed RT-PCR positive, and negative CT scan results and 105 patients showed negative results in both RT-PCR and CT scan on February 6, 2020, in Wuhan, China. Hence, CT scan has more sensitivity than RT-PCR [3]. CT scan is used for diagnosing COVID-19 patients when a person is more suspicious and has a negative RT-PCR result with repeated testing. RT-PCR is a reverse transcription-PCR used for RNA, which involved in the additional process of converting RNA to cDNA (complementary DNA). The RT-PCR is carried out in a thermocycler at various temperatures for denaturation, annealing, and multiplication of specific target the sequence of nucleic acid using specific primers. We are comprehensively reporting the testing methods for the rapid diagnosis on the basis of accuracy, reliability, and duration for the tests are summarized in Table 1 [4,5,6,7,8,9,10,11,12,13].
Sample Collection for COVID-19 Detection
Preferably, early morning patient’s sputum collected from the posterior oropharnyx for the investigation. Generally, blood, sputum, feces, urine specimens to be collected during illness and tested under different biochemical procedures. The physiological examinations such as hematology, radiology, microbiological examinations must be processed thereafter. Once the serum is drawn from the patients and immediately stored at -20 °C until all the testing procedures to be done. Furthermore, the severely affected patients must be immediately taken to the hospital for ventilation because of breathing illness and taken samples such as broncho alveolar lavage fluid and fibro bronchoscope brush biopsy to be collected. RT-PCR for the identification of RNA dependent-RNA polymerase gene region for determining the receptor binding spike protein and the nucleoprotein are examined. In addition, the electron microscopy is used to detect the live cells from the viral culture [14, 15].
RNA Based Diagnostic Techniques
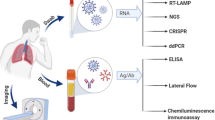
General detection of viral infections involved in the isolation and nucleic acid detection by collecting specimens from the oral swab, nasal swab, nasopharyngeal or tracheal extract, lung tissue, blood, sputum and feces [16]. Some of the alternative testing methods that were used to detect COVID-19 (Fig. 2) based on the genetic material of the virus in general and COVID-19 in particular.
Reverse Transcription-PCR (RT - PCR)
RT-PCR is a standard tool for detection since human coronavirus is an RNA virus. After the PCR process the, PCR product is analyzed by electrophoresis analysis, restriction enzyme analysis, high-throughput sequencing for confirming the presence of Viral RNA [17,18,19]. The broncho-alveolar lavage fluid was collected from the patients are infected with COVID-19. The RNA of CoV2 is extracted then to RT-PCR and unbiased; high-throughput sequencing was used to identify the presence of viral RNA using RdRp region of CoV2 as a target [5]. Although this method is more sensitive to collect viral particle directly from the affected alveolar the region, it is not used routinely because it needs special equipment, preparation and skilled person and not convenient for patients as well.
There also other alternative method but it is less sensitive when compared to the previous method, Corman et al., (2020) obtained the clinical specimens (sputum) from nose and throat swap, then the viral RNA is isolated using MagNA Pure 96 system. Then, the RT-PCR was done with a reaction mixture of 25 μL having RNA(5 μL), 2 × reaction buffer (12.5 μL, Taq mixture (1 μL), 50 mM magnesium sulfate solution (0.4 μL) and nonacetylated bovine serum albumin (1 μg), then the thermal cycling are carried out, for RT at 55 °C (10 min) followed at 95 °C (3 min) then 45 cycles at 95 °C (15 s), and 58 °C (30 s) [4]. Moreover, Real Time PCR (RT qPCR) is a simple qualitative assay used for detecting most of the human coronaviruses with more specificity and sensitivity than other conventional PCR, and it is used for early diagnosis. RT qPCR is prone to contamination and requires more time for post-analysis [18]. Meanwhile, many false-negative results have been reported due to several factors, previous investigations with a total of 957 individuals suspected or confirmed with COVID-19, the false-negative rate ranged from 2% to 29% [19]. This is because Corona virus has the mutating nature, so results from RT qPCR using primers in the gene are affected by the variation of viral RNA sequences. A COVID-19 patient’s RT qPCR test showed a false-negative result for the first three times. The RT-PCR test for SARS-CoV-2 virus, on the other hand, has several flaws that demand changes in how the technology is employed. The RT-PCR test, like immunodiagnostic assays, can have difficulty discriminating between real positive and true negative results. Individuals infected with COVID-19 [20]. The timing of sampling after the beginning of symptoms is one of the key reasons for such a high false-negative rate in RT-PCR results. Because it has been demonstrated that the false-negative rate increases with sampling time, the time of sample is critical. The test’s difficulty level changes with time [21]. To summarize, RT-PCR has a substantial false-negative rate that varies between specimens and time periods. When immunodiagnostic tests and clinical symptoms are examined with the RT-PCR test result, the false negative rate can be reduced.. Otherwise, the negative PCR test findings will no longer provide us with sufficient assurance that the suspected case is disease-free.
Microarray Technique
COVID-19 is used to diagnose with rapid and high-throughput method. The cDNA labeled with the specific probe was produced from the RNA of the SARS-CoV2 by the process of reverse transcription. These labeled cDNA are loaded in each well-containing solid-phase oligonucleotide and get hybridized, and the microarray plates are washed to remove free DNA. Eventually, the presence of coronavirus RNA is found by the detection of a specific probe [7]. A 60mer oligonucleotide was designed that corresponds to the TOR2 sequence of coronavirus, and then it’s hybridized with cDNA sequence of the RNA extracted from the nasal swab and gargling fluid [22]. In addition, microarray approaches that use a scanner to display the hybridization between the probe and the target are a quick, sensitive, specific, and accurate way of detecting hybridization which is successfully used for detecting 24 single nucleotide polymorphisms mutation in the gene that encodes spike protein of human coronavirus, this technique showed 100% accuracy [23]. They can also do simultaneous analyses of multiple microbial genes. Although it can detect many samples, this approach does not allow for the detection of a few viral genes in small samples.
RT Loop-Mediated Isothermal Amplification (RT-LAMP)
RT-LAMP is used to amplify the specific gene by a set 4 or 6 unique primers in general. The amplification detection is done by either precipitation of magnesium pyrophosphate or by fluorescence under UV-radiation and also by monitoring the pyrophosphate turbidity level. LAMP assay is also used to detect other RNA virus such as respiratory CoV2. RT-LAMP uses a single tube under the isothermal condition of a single-stranded temperature of (63 °C) detect the MERS. This procedure requires only 1 h to perform and requires only a single standard temperature for amplification. It is done by primer or probe that targets the conserved nucleocapsid protein region (upE and ORF1a gene) [8]. During the SARS epidemic, 49 samples were collected from the patients and performed RT-LAMP assay and compared to RT-PCR, concluded that sensitivity is 100% and specificity is 87% for RT-LAMP with regard to RT-PCR respectively [24, 25]. The fundamental disadvantage of the LAMP-based approach is the difficulty of designing primers which is required to obtain the specificity of a PCR test maintaining the benefit of running the entire experiment at a constant temperature [26]. Another possible drawback of such methods is that they can provide false-positive results as a result of carry-over from earliest tests particularly when upgraded into an automated platform. RT-LAMP detection using quenched fluorescent primers, for example, can circumvent these constraints. Several other improvements to the LAMP method are being optimized and validated in various laboratories around the world to meet the needs of the COVID-19 pandemics [27, 28].
Multiplex Test
It is used to detect and distinguishing coronavirus and other viruses, multiplexed coronavirus. The mass spectrometry method detects the pathogenic human coronavirus and provides phylogenetic evidence about unknown or novel coronavirus. This method is performed in a Mass ARRAY using MALDI-TOF data, and it is highly sensitive and simple to perform [9]. Novel arch-shaped multiple-target sensor involves indirect amplification using primers that readily amplify the multiple targets since it has high sensitivity, accuracy, and specificity. This technique also helps in distinguishing the coronavirus and another respiratory virus in clinical specimens within 20 min. More research is being done now to solve some of the major flaws in LFIA test systems, particularly in terms of generating quantitative data and documenting them. They can be digitized with scanners or cameras and specific software that allows them to be recorded and transmitted across long distances. However, technological advancements will necessitate more complex technology, increasing the cost and duration of analysis in the long run.
Paper-Based Colorimetric Test
The paper-based colorimetric is based on the aforementioned principle, and this is a simple, rapid approach, where the aggregation of pyrrolidinyl peptide nucleic acid-induced silver nanoparticles is used for detection of CoV2 oligonucleotide [29]. The acpcPNA probe is used as an attractive alternative to RNA and DNA probes. They are chemically and biologically stable and effectively hybridized with cDNA sequence. The color change indicates the presence of target oligonucleotide by the electrostatic repulsion, where anionic DNA-acpcPNA duplex results in the dispersion of silver nanoparticles. Moreover, this method has some disadvantages concerning sensitivity and selectivity because it depends on pH, concentration of nanoparticles and DNA probe and mismatched DNA strand.
Immunodiagnostic Techniques
Immunodiagnostic of Antigen
This diagnostic test involved in identifying the availability of CoV2 antigen and considered as the best test for detection of acute or early infection typically within 30 min [13]. The specific antibodies for the viral antigen are adsorbed on the paper strip and enclosed by a plastic casing. The visually detectable signal is observed if the target antigens bind with specific antibodies. WHO does not recommend the antigen-detecting rapid diagnostic tests for patient care due to limited available data [12]. The sensitivity of the rapid diagnostic test is expected to be varied in the range (34–80%). The sensitivity of this method depends on time from onset of infection, virus concentration and quality of the sample collected from sputum or throat swab, and precise formulating of reagents in test kits [11].
Immunodiagnostic of Antibody
Alternatively, a rapid test for COVID-19, used to identify the specific antibodies in the blood against CoV2 antigen. Generally, antibodies are generated over some days after the infection, and its strength depends on age, nutritional health status, disease severity and some medications or infection that suppress the immune system [13].
Colloidal Gold-Based Immunochromatographic Assay (CG-ICG)
CG-ICG is also known as IgM/IgG antibody test involved in the collection of blood samples containing serum, plasma samples for the detection of CoV2. This technique contains two strips for IgM and IgG. Briefly, 1–2 volume of serum or whole blood samples is added into the strip, after about 15 min of incubation, the strip containing the positive sample will appear line called T and C; if the sample is present only in C line then it is negative; It will be considered as invalid test if there is no C line is showed up. The sensitivity of CG-ICG assay in nuclear acid confirmed cases were 11.1, 92.9 and 96.8% at early, intermediate and late stages, respectively [13].
Radiation Mediated Diagnosis
Chest X-Ray
Chest X-ray images revealed about the rapid progression of pneumonia and also showed differences between the left and right lung by the observation of bilateral or unilateral opacities due to infections of COVID-19 using X-ray images [6].
Ultrasonography
Lung ultrasonographic imaging has similar results with respect to chest CT with some additional advantage reported elsewhere (24). The characteristics a feature of pneumonia due to COVID-19 includes, pleural line was irregularly thickened, B lines were observed in various identical patterns. The merging of these identical patterns, non-translobar, translobar and multifocal small, with bronchograms, lines have appeared during the remedial phase, and uncommon pleural effusions [30].
Other Qualitative Assay
Cobas SARS-CoV2
The cobas® 6800/8800 system is a qualitative test used for the detection of SARS-CoV2 RNA. Copan or BD™ universal transport systems are used for collecting the sample from the oropharyngeal and nasopharyngeal swab. The entire sample preparation and PCR is regulated by RNA internal control, some external controls such as low titer positive control and negative control are regulated by this system. Positive results of SARS-CoV2 RNA detection do not confirm the presence of the transmissible virus and negative results do preclude SARS-CoV2 infection, it must be considered along with symptoms, patient’s travel host and epidemiological information [12]. On the whole, there are plenty of in silico studies are reported to study the anti-COVID property of essential oils, microalgal peptides and so on [31].
Conclusion
To summarize, many COVID-19 cases were asymptomatic; hence, it is a great challenge for healthcare workers to diagnosis appropriately. There are many standard methods available reported in this review for the diagnosis of COVID-19 accordingly, if any suspected person has positive results by following diagnosis methods is considered to be a confirmed case.
References
Jin, Y., Yang, H., Ji, W., Wu, W., Chen, S., Zhang, W., & Duan, G. (2020). Virology, epidemiology, pathogenesis, and control of COVID-19. Viruses, 12(4), 372. https://doi.org/10.3390/v12040372
Singhal, T. (2020). A review of coronavirus disease-2019 (COVID-19). The Indian Journal of Pediatrics, 87(4), 281–286. https://doi.org/10.1007/s12098-020-03263-6
Ai, T., Yang, Z., Hou, H., Zhan, C., Chen, C., Lv, W., ... & Xia, L. (2020). Correlation of chest CT and RT-PCR testing for coronavirus disease 2019 (COVID-19) in China: A report of 1014 cases. Radiology, 296(2), E32-E40. https://doi.org/10.1148/radiol.2020200642.
Corman, V. M., Landt, O., Kaiser, M., Molenkamp, R., Meijer, A., Chu, D. K., ... & Drosten, C. (2020). Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance, 25(3), 2000045. https://doi.org/10.2807/15607917.ES.2020.25.3.2000045.
Zhu, N., Zhang, D., Wang, W., Li, X., Yang, B., Song, J., ... & Tan, W. (2020). A novel coronavirus from patients with pneumonia in China, 2019. New England Journal of Medicine. https://doi.org/10.1056/NEJMoa2001017.
Xu, Z., Shi, L., Wang, Y., Zhang, J., Huang, L., Zhang, C., ... & Wang, F. S. (2020). Pathological findings of COVID-19 associated with acute respiratory distress syndrome. The Lancet Respiratory Medicine, 8(4), 420–422. https://doi.org/10.1016/S2213-2600(20)30076-X.
Chen, Q., Li, J., Deng, Z., Xiong, W., Wang, Q., & Hu, Y. Q. (2010). Comprehensive detection and identification of seven animal coronaviruses and human respiratory coronavirus 229E with a microarray hybridization assay. Intervirology, 53(2), 95–104. https://doi.org/10.1159/000264199
Shirato, K., Yano, T., Senba, S., Akachi, S., Kobayashi, T., Nishinaka, T., ... & Matsuyama, S. (2014). Detection of Middle East respiratory syndrome coronavirus using reverse transcription loop-mediated isothermal amplification (RT-LAMP). Virology Journal, 11(1), 1–11. https://doi.org/10.1186/1743-422X-11-139.
Xiu, L., Zhang, C., Wu, Z., & Peng, J. (2017). Establishment and application of a universal coronavirus screening method using MALDI-TOF mass spectrometry. Frontiers in Microbiology, 8, 1510. https://doi.org/10.3389/fmicb.2017.01510
Bhadra, S., Jiang, Y. S., Kumar, M. R., Johnson, R. F., Hensley, L. E., & Ellington, A. D. (2015). Real-time sequence-validated loop-mediated isothermal amplification assays for detection of Middle East respiratory syndrome coronavirus (MERS-CoV). PLoS One, 10(4), e0123126.
Bruning, A. H., Leeflang, M. M., Vos, J. M., Spijker, R., de Jong, M. D., Wolthers, K. C., & Pajkrt, D. (2017). Rapid tests for influenza, respiratory syncytial virus, and other respiratory viruses: A systematic review and meta-analysis. Clinical Infectious Diseases, 65(6), 1026–1032. https://doi.org/10.1093/cid/cix461
WHO lists two COV2ID-19 tests for emergency use. (2020). https://www.who.int/news-room/detail/07-04-2020-who-lists-two-Covid-19-tests-for-emergency-use
Pan, Y., Li, X., Yang, G., Fan, J., Tang, Y., Zhao, J., ... & Li, Y. (2020). Serological immunochromatographic approach in diagnosis with SARS-CoV-2 infected COVID-19 patients. Journal of Infection, 81(1), e28-e32. https://doi.org/10.1101/2020.03.13.20035428.
Wang, W., Xu, Y., Gao, R., Lu, R., Han, K., Wu, G., & Tan, W. (2020). Detection of SARS-CoV-2 in different types of clinical specimens. Jama, 323(18), 1843–1844. https://doi.org/10.1001/jama.2020.3786
To, K. K. W., Tsang, O. T. Y., Leung, W. S., Tam, A. R., Wu, T. C., Lung, D. C., ... & Yuen, K. Y. (2020). Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study. The Lancet Infectious Diseases, 20(5), 565–574. https://doi.org/10.1016/S1473-3099(20)30196-1.
Yu, F., Du, L., Ojcius, D. M., Pan, C., & Jiang, S. (2020). Measures for diagnosing and treating infections by a novel coronavirus responsible for a pneumonia outbreak originating in Wuhan, China. Microbes and Infection, 22(2), 74–79. https://doi.org/10.1016/j.micinf.2020.01.003
Balboni, A., Gallina, L., Palladini, A., Prosperi, S., & Battilani, M. (2012). A real-time PCR assay for bat SARS-like coronavirus detection and its application to Italian greater horseshoe bat faecal sample surveys. The Scientific World Journal, 2012. https://doi.org/10.1100/2012/989514
Shen, M., Zhou, Y., Ye, J., Al-Maskri, A. A. A., Kang, Y., Zeng, S., & Cai, S. (2020). Recent advances and perspectives of nucleic acid detection for coronavirus. Journal of Pharmaceutical Analysis, 10(2), 97–101. https://doi.org/10.1016/j.jpha.2020.02.010
Arevalo-Rodriguez, I., Buitrago-Garcia, D., Simancas-Racines, D., Zambrano-Achig, P., Del Campo, R., Ciapponi, A., ... & Zamora, J. (2020). False-negative results of initial RT-PCR assays for COVID-19: A systematic review. PLoS One, 15(12), e0242958. https://doi.org/10.1371/journal.pone.0242958.
Abbasi-Oshaghi, E., Mirzaei, F., Farahani, F., Khodadadi, I., & Tayebinia, H. (2020). Diagnosis and treatment of coronavirus disease 2019 (COVID-19): Laboratory, PCR, and chest CT imaging findings. International Journal of Surgery, 79, 143–153. https://doi.org/10.1016/j.ijsu.2020.05.018
Wang, Y., Kang, H., Liu, X., & Tong, Z. (2020). Combination of RT-qPCR testing and clinical features for diagnosis of COVID-19 facilitates management of SARS-CoV-2 outbreak. Journal of Medical Virology. https://doi.org/10.1002/jmv.25721
Shi, R., Ma, W., Wu, Q., Zhang, B., Song, Y., Guo, Q., ... & Zheng, W. (2003). Design and application of 60mer oligonucleotide microarray in SARS coronavirus detection. Chinese Science Bulletin, 48(12), 1165–1169. https://doi.org/10.1007/BF03183928.
Guo, X., Geng, P., Wang, Q., Cao, B., & Liu, B. (2014). Development of a single nucleotide polymorphism DNA microarray for the detection and genotyping of the SARS coronavirus. Journal of Microbiology and Biotechnology, 24(10), 1445–1454. https://doi.org/10.4014/jmb.1404.04024
Behzadi, P., Ranjbar, R., & Alavian, S. M. (2015). Nucleic acid-based approaches for detection of viral hepatitis. Jundishapur Journal of Microbiology, 8(1). https://doi.org/10.5812/jjm.17449
Thai, H. T. C., Le, M. Q., Vuong, C. D., Parida, M., Minekawa, H., Notomi, T., ... & Morita, K. (2004). Development and evaluation of a novel loop-mediated isothermal amplification method for rapid detection of severe acute respiratory syndrome coronavirus. Journal of Clinical Microbiology, 42(5), 1956–1961. https://jcm.asm.org/content/42/5/1956.short
Jia, X., Zhang, P., Tian, Y., Wang, J., Zeng, H., Wang, J., ... & Liu, Y. (2020). Clinical significance of IgM and IgG test for diagnosis of highly suspected COVID-19 infection. MedRxiv. https://doi.org/10.1101/2020.02.28.20029025.
Xu, M., Wang, D., Wang, H., Zhang, X., Liang, T., Dai, J., ... & Yu, X. (2020). COVID-19 diagnostic testing: Technology perspective. Clinical and Translational Medicine, 10(4), e158. https://doi.org/10.1002/ctm2.158.
Harding, H., Broom, A., & Broom, J. (2020). Aerosol-generating procedures and infective risk to healthcare workers from SARS-CoV-2: The limits of the evidence. Journal of Hospital Infection, 105(4), 717–725. https://doi.org/10.1016/j.jhin.2020.05.037
Teengam, P., Siangproh, W., Tuantranont, A., Vilaivan, T., Chailapakul, O., & Henry, C. S. (2017). Multiplex paper-based colorimetric DNA sensor using pyrrolidinyl peptide nucleic acid-induced AgNPs aggregation for detecting MERS-CoV, MTB, and HPV oligonucleotides. Analytical Chemistry, 89(10), 5428–5435. https://doi.org/10.1021/acs.analchem.7b00255
Peng, Q. Y., Wang, X. T., & Zhang, L. N. (2020). Findings of lung ultrasonography of novel corona virus pneumonia during the 2019–2020 epidemic. Intensive Care Medicine, 46(5), 849–850. https://doi.org/10.1007/s00134-020-05996-6
MubarakAli, D., MohamedSaalis, J., Sathya, R., Irfan, N., & Kim, J.-W. (2021). An evidence of microalgal peptide to target spike protein of COVID19: In silico approach. Microbial Pathogenesis, 160, 105189.
Acknowledgements
This study was supported by AUN/SEED-Net, JICA through Special Program for Research against COVID19 (SPRAC 2020) program for the financial support (SN042/MI.KU/2020). The author is greatly acknowledges to Crescent Innovation and Incubation Council (CIIC), BSACIST.
Availability of Data and Materials
Not applicable.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
RD: data acquisition, RS: data acquisition; DM: review, guidance and proofreading.
Corresponding author
Ethics declarations
Conflict of Interest
There is no conflict of interest to this work.
Ethical Approval
Not applicable.
Consent to Participate
Not applicable.
Consent to Publish
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
MubarakAli, D. Comprehensive Review on Rapid Diagnosis of New Infection COVID-19. Appl Biochem Biotechnol 194, 1390–1400 (2022). https://doi.org/10.1007/s12010-021-03728-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-021-03728-0