Abstract
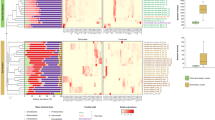
Termites are well recognized for their thriving on recalcitrant lignocellulosic diets through nutritional symbioses with gut-dwelling microbiota; however, the effects of diet changes on termite gut microbiota are poorly understood, especially for the lower termites. In this study, we employed high-throughput 454 pyrosequencing of 16S V1–V3 amplicons to compare gut microbiotas of Tsaitermes ampliceps fed with lignin-rich and lignin-poor cellulose diets after a 2-week-feeding period. As a result, the majority of bacterial taxa were shared across the treatments with different diets, but their relative abundances were modified. In particular, the relative abundance was reduced for Spirochaetes and it was increased for Proteobacteria and Bacteroides by feeding the lignin-poor diet. The evenness of gut microbiota exhibited a significant difference in response to the diet type (filter paper diets < corn stover diets < wood diets), while their richness was constant, which may be related to the lower recalcitrance of this biomass to degradation. These results have important implications for sampling and analysis strategies to probe the lignocellulose degradation features of termite gut microbiota and suggest that the dietary lignocellulose composition could cause shifting rapidly in the termite gut microbiota.





Similar content being viewed by others
References
Bugg, T. D., Ahmad, M., Hardiman, E. M., & Singh, R. (2011). The emerging role for bacteria in lignin degradation and bio-product formation. Current Opinion in Biotechnology, 22, 394–400.
McDonald, J. E., Rooks, D. J., & McCarthy, A. J. (2012). Methods for the isolation of cellulose-degrading microorganisms. Methods in Enzymology, 510, 349–374.
Brune, A. (2014). Symbiotic digestion of lignocellulose in termite guts. Nature Reviews Microbiology, 12, 168–180.
Scharf, M. E. (2015). Omic research in termites: an overview and a roadmap. Frontiers in Genetics, 6, 76.
Fraune, S., & Bosch, T. C. G. (2010). Why bacteria matter in animal development and evolution. BioEssays, 32, 571–580.
Scharf, M. E. (2015). Termites as targets and models for biotechnology. Annual Review of Entomology, 60, 77–102.
Warnecke, F., Luginbühl, P., Ivanova, N., Ghassemian, M., Richardson, T. H., Stege, J. T., Cayouette, M., McHardy, A. C., Djordjevic, G., Aboushadi, N., Sorek, R., Tringe, S. G., Podar, M., Martin, H. G., Kunin, V., Dalevi, D., Madejska, J., Kirton, E., Platt, D., Szeto, E., Salamov, A., Barry, K., Mikhailova, N., Kyrpides, N. C., Matson, E. G., Ottesen, E. A., Zhang, X., Hernández, M., Murillo, C., Acosta, L. G., Rigoutsos, I., Tamayo, G., Green, B. D., Chang, C., Rubin, E. M., Mathur, E. J., Robertson, D. E., Hugenholtz, P., & Leadbetter, J. R. (2007). Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature, 450, 560–565.
Werren, J. H., Baldo, L., & Clark, M. E. (2008). Wolbachia: master manipulators of invertebrate biology. Nature Reviews Microbiology, 6, 740–751.
Benjamino, J., & Graf, J. (2016). Characterization of the core and caste-specific microbiota in the termite, Reticulitermes flavipes. Frontiers in Microbiology, 7, 171.
Ohkuma, M., Sato, T., Noda, S., Ui, S., Kudo, T., & Hongoh, Y. (2007). The candidate phylum ‘termite group 1’ of bacteria: phylogenetic diversity, distribution, and endosymbiont members of various gut flagellated protists. FEMS Microbiology Ecology, 60, 467–476.
Brugerolle, G., & Radek, R. (2006). Symbiotic protozoa of termites. In H. König & A. Varma (Eds.), Intestinal microorganisms of soil invertebrates. Soil biology. Vol. 6 (pp. 243–269). Berlin: Springer.
Berlanga, M., Pasteur, B. J., Grandcolas, P., & Guerrero, R. (2011). Comparison of the gut microbiota from soldier and worker castes of the termite Reticulitermes grassei. International Microbiology, 14, 83–93.
Scharf, M. E., Karl, Z. J., Sethi, A., & Boucias, D. G. (2011). Multiple levels of synergistic collaboration in termite lignocellulose digestion. PloS One, 6, e21709.
Scharf, M. E., Karl, Z. J., Sethi, A., Sen, R., Raychoudhury, R., & Boucias, D. G. (2011). Defining host-symbiont collaboration in termite lignocellulose digestion, “the view from the tip of the iceberg. Communicative & Integrative Biology, 4, 761–763.
Sinma, K., Khucharoenphaisan, K., Kitpreechavanich, V., & Tokuyama, S. (2011). Purification and characterization of a thermostable xylanase from Saccharopolyspora pathumthaniensis S582 isolated from the gut of a termite. Bioscience, Biotechnology, and Biochemistry, 75, 1957–1963.
Hongoh, Y. (2010). Diversity and genomes of uncultured microbial symbionts in the termite gut. Bioscience, Biotechnology, and Biochemistry, 74, 1145–1151.
Tai, V., James, E. R., Nalepa, C. A., Scheffrahn, R. H., Perlman, S. J., & Keeling, P. J. (2015). The role of host phylogeny varies in shaping microbial diversity in the hindguts of lower termites. Applied and Environmental Microbiology, 81, 1059–1070.
Abdul Rahman, N., Parks, D. H., Willner, D. L., Engelbrektson, A. L., Goffredi, S. K., Warnecke, F., Scheffrahn, R. H., & Hugenholtz, P. (2015). A molecular survey of Australian and north American termite genera indicates that vertical inheritance is the primary force shaping termite gut microbiomes. Microbiologica, 3, 5.
Karl, Z. J., & Scharf, M. E. (2015). Effects of five diverse lignocellulosic diets on digestive enzyme biochemistry in the termite Reticulitermes flavipes. Archives of Insect Biochemistry and Physiology, 90, 89–103.
Huang, X. F., Bakker, M. G., Judd, T. M., Reardon, K. F., & Vivanco, J. M. (2013). Variations in diversity and richness of gut bacterial communities of termites (Reticulitermes flavipes) fed with grassy and woody plant substrates. Microbial Ecology, 65, 531–536.
Boucias, D. G., Cai, Y., Sun, Y., Lietze, V. U., Sen, R., Raychoudhury, R., & Scharf, M. E. (2013). The hindgut lumen prokaryotic microbiota of the termite Reticulitermes flavipes and its responses to dietary lignocellulose composition. Molecular Ecology, 22, 1836–1853.
Raychoudhury, R., Sen, R., Cai, Y., Sun, Y., Lietze, V. U., Boucias, D. G., & Scharf, M. E. (2013). Comparative metatranscriptomic signatures of wood and paper feeding in the gut of the termite Reticulitermes flavipes (Isoptera: Rhinotermitidae). Insect Molecular Biology, 22, 155–171.
Brauman, A., Doré, J., Eggleton, P., Bignell, D., Breznak, J. A., & Kane, M. D. (2001). Molecular phylogenetic profiling of prokaryotic communities in guts of termites with different feeding habits. FEMS Microbiology Ecology, 35, 27–36.
Mikaelyan, A., Dietrich, C., Köhler, T., Poulsen, M., Sillam-Dussès, D., & Brune, A. (2015). Diet is the primary determinant of bacterial community structure in the guts of higher termites. Molecular Ecology, 24, 5284–5295.
Huang, F. S., Zhu, S. M., Ping, Z. M., He, X. S., & Li, G. X. (2000). Fauna Sinica Insecta, vol. 17 Isoptera (pp. 430–865). Beijing: Science Press.
Su, L. J., Liu, Y. Q., Liu, H., Wang, Y., Li, Y., Lin, H. M., Wang, F. Q., & Song, A. D. (2015). Linking lignocellulosic dietary patterns with gut microbial Enterotypes of Tsaitermes ampliceps and comparison with Mironasutitermes shangchengensis. Genetics and Molecular Research, 14, 13954–13967.
Liu, N., Yan, X., Zhang, M., Xie, L., Wang, Q., Huang, Y., Zhou, X., Wang, S., & Zhou, Z. (2011). Microbiome of fungus-growing termites: a new res ervoir for lignocellulase genes. Applied and Environmental Microbiology, 77, 48–56.
Schloss, P. D., Gevers, D., & Westcott, S. L. (2011). Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PloS One, 6, e27310.
Schloss, P. D., Westcott, S. L., Ryabin, T., et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Applied and Environmental Microbiology, 75, 7537–7541.
Kunin, V., Engelbrektson, A., Ochman, H., & Hugenholtz, P. (2010). Wrinkles in the rare biosphere: pyrosequencing errors can lead to artificial inflation of diversity estimates. Environmental Microbiology, 12, 118–123.
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., & Knight, R. U. (2011). CHIME improves sensitivity and speed of chimera detection. Bioinformatics, 27, 2194–2200.
Kuhnigk, T., & König, H. (1997). Degradation of dimeric lignin model compounds by aerobic bacteria isolated from the hindgut of xylophagous termites. Journal of Basic Microbiology, 37, 205–211.
Aickin, M., & Gensler, H. (1996). Adjusting for multiple testing when reporting research results: the Bonferroni vs Holm methods. American Journal of Public Health, 86, 726–728.
Berchtold, M., Chatzinotas, A., Schönhuber, W., Brune, A., Amann, R., Hahn, D., & König, H. (1999). Differential enumeration and in situ localization of microorganisms in the hindgut of the lower termite Mastotermes darwiniensis by hybridization with rRNA-targeted probes. Archives of Microbiology, 172, 407–416.
Ni, J., & Tokuda, G. (2013). Lignocellulose-degrading enzymes from termites and their symbiotic microbiota. Biotechnology Advances, 31, 838–850.
Brune, A. (2006). Symbiotic associations between termites and prokaryotes. In M. Dworkin, S. Falkow, E. Rosenberg, K.-H. Schleifer, & E. Stackebrandt (Eds.), The prokaryotes, vol. 1. Symbiotic associations, biotechnology, applied microbiology (pp. 439–474). New York: Springer.
He, S., Ivanova, N., Kirton, E., Allgaier, M., Bergin, C., Scheffrahn, R. H., Kyrpides, N. C., Warnecke, F., Tringe, S. G., & Hugenholtz, P. ((2013)). Comparative metagenomic and metatranscriptomic analysis of hindgut paunch microbiota in wood- and dung-feeding higher termites. PloS One, 8, e61126.
Dietrich, C., Köhler, T., & Brune, A. (2014). The cockroach origin of the termite gut microbiota: patterns in bacterial community structure reflect major evolutionary events. Applied and Environmental Microbiology, 80, 2261–2269.
Dröge, S., Fröhlich, J., Radek, R., & König, H. (2006). Spirochaeta coccoides sp. nov., a novel coccoid spirochete from the hindgut of the termite Neotermes castaneus. Applied and Environmental Microbiology, 72, 392–397.
Hongoh, Y., Sharma, V. K., Prakash, T., Noda, S., Toh, H., Taylor, T. D., Kudo, T., Sakaki, Y., Toyoda, A., Hattori, M., & Ohkuma, M. (2008). Genome of an endosymbiont coupling N2 fixation to cellulolysis within protist cells in termite gut. Science, 322, 1108–1109.
Yang, Y. J., Zhang, N., Ji, S. Q., Lan, X., Zhang, K. D., Shen, Y. L., Li, F. L., & Ni, J. F. (2014). Dysgonomonas macrotermitis sp. nov., isolated from the hindgut of a fungus-growing termite. International Journal of Systematic and Evolutionary Microbiology, 64, 2956–2961.
Pramono, A. K., Sakamoto, M., Iino, T., Hongoh, Y., & Ohkuma, M. (2015). Dysgonomonas termitidis sp. nov., isolated from the gut of the subterranean termite Reticulitermes speratus. International Journal of Systematic and Evolutionary Microbiology, 65, 681–685.
Pinheiro, G. L., Correa, R. F., Cunha, R. S., Cardoso, A. M., Chaia, C., Clementino, M. M., Garcia, E. S., de Souza, W., & Frasés, S. (2015). Isolation of aerobic cultivable cellulolytic bacteria from different regions of the gastrointestinal tract of giant land snail Achatina fulica. Frontiers in Microbiology, 6, 860.
Hongoh, Y., Sato, T., Dolan, M. F., Noda, S., Ui, S., Kudo, T., & Ohkuma, M. (2007). The motility symbiont of the termite gut flagellate Caduceia versatilis is a member of the “Synergistes” group. Applied and Environmental Microbiology, 73, 6270–6276.
Wasi, S., Tabrez, S., & Ahmad, M. (2013). Use of Pseudomonas spp. for the bioremediation of environmental pollutants: a review. Environmental Monitoring and Assessment, 185, 8147–8155.
Jiménez, D. J., Korenblum, E., & van Elsas, J. D. (2014). Novel multispecies microbial consortia involved in lignocellulose and 5-hydroxymethylfurfural bioconversion. Applied Microbiology and Biotechnology, 98, 2789–2803.
Kannisto, M. S., Mangayil, R. K., Shrivastava-Bhattacharya, A., Pletschke, B. I., Karp, M. T., & Santala, V. P. (2015). Metabolic engineering of Acinetobacter Baylyi ADP1 for removal of clostridium butyricum growth inhibitors produced from lignocellulosic hydrolysates. Biotechnology for Biofuels, 8, 198.
Oosterkamp, M. J., Méndez-García, C., Kim, C.-H., Bauer, S., Ibáñez, A. B., Zimmerman, S., Hong, P.-Y., Cann, I. K., & Mackie, R. I. (2016). Lignocellulose-derived thin stillage composition and efficient biological treatment with a high-rate hybrid anaerobic bioreactor system. Biotechnology for Biofuels, 9, 120.
He, Y., Ding, Y., & Long, Y. (1991). Two cellulolytic clostridium species: Clostridium cellulosi sp. nov. and Clostridium cellulofermentans sp. nov. International Journal of Systematic Bacteriology, 41, 306–309.
Huang, X. F., Santhanam, N., Badri, D. V., Hunter, W. J., Manter, D. K., Decker, S. R., Vivanco, J. M., & Reardon, K. F. (2013). Isolation and characterization of lignin-degrading bacteria from rainforest soils. Biotechnology and Bioengineering, 110, 1616–1626.
Geib, S. M., Filley, T. R., Hatcher, P. G., et al. (2008). Lignin degradation in wood-feeding insects. Proceedings of the National Academy of Sciences of the United States of America, 105, 12932–12937.
Chaffron, S., & von Mering, C. (2007). Termites in the woodwork. Genome Biology, 8, 229.
Wei, H., Xu, Q., Taylor, L. E., Baker, J. O., Tucker, M. P., & Ding, S. Y. (2009). Natural paradigms of plant cell wall degradation. Current Opinion in Biotechnology, 20, 330–338.
Vikman, M., Karjomaa, S., Kapanen, A., Wallenius, K., & Itavaara, M. (2002). The influence of lignin content and temperature on the biodegradation of lignocellulose in composting conditions. Applied Microbiology and Biotechnology, 59, 591–598.
Reichling, J. (2010). Plant–microbe interactions and secondary metabolites with antibacterial, antifungal and antiviral properties. In M. Wink (Ed.), Annual plant reviews volume 39: functions and biotechnology of plant secondary metabolites (pp. 214–347). Oxford: Wiley.
Acknowledgments
This research was supported by National Science Foundation of China Grant 31170350.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
We declare that we have no financial and personal relationships with other people or organizations that can inappropriately influence our work. There is no professional or other personal interest of any nature or kind in any product, service, and/or company that could be construed as influencing the position presented in the manuscript entitled, “Variation in the gut microbiota and sensitivity to dietary changes in termite hosts”.
Additional information
Lijuan Su and Lele Yang contributed equally to this work.
Electronic supplementary material
Supplementary Fig. S1
(DOCX 156 kb)
Supplementary Table S1
(DOCX 29 kb)
Supplementary Table S2
(DOCX 68 kb)
Rights and permissions
About this article
Cite this article
Su, L., Yang, L., Huang, S. et al. Variation in the Gut Microbiota of Termites (Tsaitermes ampliceps) Against Different Diets. Appl Biochem Biotechnol 181, 32–47 (2017). https://doi.org/10.1007/s12010-016-2197-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2197-2