Abstract
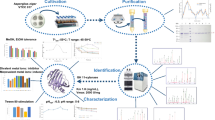
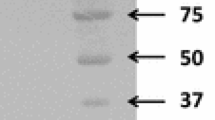
Streptomyces coelicolor A3(2) sco5931 gene was predicted to encode a putative xylanase A, a 477 amino acid protein belonging to glycoside hydrolase family 10. The entire sco5931 coding region was cloned and overexpressed in Streptomyces lividans TK24. Mature SCO5931 protein comprising 436 amino acids (47 kDa) was purified by single-step gel filtration chromatography from culture broth after ammonium sulfate precipitation, with 25.8-fold purification and yield of 30.6 %. The purified protein displayed a pronounced activity toward beechwood xylan as a substrate, but no activity was detected toward carboxymethylcellulose, Avicel, galactan, barley β-glucan, and xyloglucan, demonstrating that SCO5931 is a substrate-specific xylanase. Optimal xylanase activity was observed at 60 °C and pH 6.0. The addition of metal ions or EDTA did not affect the xylanase activity, while 4 mM MnCl2 severely inhibited the enzyme, reducing its activity by 87 %. Kinetic parameters of SCO5931 toward beechwood xylan were determined (K m = 0.24 mg/mL, V max = 6.86 μM/min). Thin layer chromatography and mass spectrometry analyses of the beechwood xylan SCO5931 hydrolysis products were conducted. Product masses corresponded to sodium adducts of xylobiose (m/z 305.24) and xylopentaose (m/z 701.59), indicating that SCO5931 specifically cleaves the β-1,4 linkage of xylan to yield xylobiose and xylopentaose.




Similar content being viewed by others
References
Bajpai, P. (1997). Microbial xylanolytic enzyme system: properties and applications. Advances in Applied Microbiology, 43, 141–194.
Dashtban, M., Schraft, H., & Qin, W. (2009). Fungal bioconversion of lignocellulosic residues; opportunities & perspectives. International Journal of Biological Sciences, 5, 578–595.
Collins, T., Gerday, C., & Feller, G. (2005). Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiology Reviews, 29, 3–23.
Beg, Q. K., Kapoor, M., Mahajan, L., & Hoondal, G. S. (2001). Microbial xylanases and their industrial applications: a review. Applied Microbiology and Biotechnology, 56, 326–338.
Chi, W. J., Youn, Y. S., Park, J. S., & Hong, S. K. (2015). Bacillus coreaensis sp. nov.: a xylan-hydrolyzing bacterium isolated from the soil of Jeju Island, Republic of Korea. The Journal of Microbiology, 53, 448–453.
Zhang, J., Matti, S. A., Terhi, P., Ming, T., Maija, T., & Liisa, V. (2011). Thermostable recombinant xylanases from Nonomuraea flexuosa and Thermoascus aurantiacus show distinct properties in the hydrolysis of xylans and pretreated wheat straw. Biotechnology for Biofuels, 4, 12–25.
Kaneko, S., Ichinose, H., Fujimoto, Z., Kuno, A., Yura, K., Go, M., Mizuno, H., Kusakabe, I., & Kobayashi, H. (2004). Structure and function of a family 10 beta-xylanase chimera of Streptomyces olivaceoviridis E-86 FXYN and Cellulomonas fimi Cex. Jornal of Biological Chemistry, 279, 26619–26626.
Kimdo, Y., Shin, D. H., Jung, S., Lee, J. S., Cho, H. Y., Bae, K. S., Sung, C. K., Rhee, Y. H., Son, K. H., & Park, H. Y. (2014). Biocatalytic properties and substrate-binding ability of a modular GH10 β-1,4-xylanase from an insect-symbiotic bacterium, Streptomyces mexicanus HY-14. Journal of Microbiology, 52, 863–870.
Morosoli, R., Bertrand, J. L., Mondou, F., Shareck, F., & Kluepfel, D. (1986). Purification and properties of a xylanase from Streptomyces lividans. Biochemical Journal, 239, 587–592.
Temuujin, U., Chi, W. J., Chang, Y. K., & Hong, S. K. (2012). Identification and biochemical characterization of Sco3487 from Streptomyces coelicolor A3(2), an exo- and endo-type β-agarase-producing neoagarobiose. Journal of Bacteriology, 194, 142–149.
Enkhbaatar, B., Temuujin, U., Lim, J. H., Chi, W. J., Chang, Y. K., & Hong, S. K. (2012). Identification and characterization of a xyloglucan-specific family 74 glycosyl hydrolase from Streptomyces coelicolor A3(2). Appllied and Environmental Microbiology, 78, 607–611.
Lim, J. H., Lee, C. R., Dhakshnamoorthy, V., Park, J. S., & Hong, S. K. (2015). Molecular characterization of Streptomyces coelicolor A(3) SCO6548 as a cellulose 1,4-β-cellobiosidase. FEMS Microbiology Letters, pii: fnv245.
Bentley, S. D., Chater, K. F., Cerdeño-Tárraga, A. M., Challis, G. L., Thomson, N. R., James, K. D., Harris, D. E., Quail, M. A., Kieser, H., Harper, D., Bateman, A., Brown, S., Chandra, G., Chen, C. W., Collins, M., Cronin, A., Fraser, A., Goble, A., Hidalgo, J., Hornsby, T., Howarth, S., Huang, C. H., Kieser, T., Larke, L., Murphy, L., Oliver, K., O’Neil, S., Rabbinowitsch, E., Rajandream, M. A., Rutherford, K., Rutter, S., Seeger, K., Saunders, D., Sharp, S., Squares, R., Squares, S., Taylor, K., Warren, T., Wietzorrek, A., Woodward, J., Barrell, B. G., Parkhill, J., & Hopwood, D. A. (2002). Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature, 417, 141–147.
Kieser, T., Bibb, M. J., Buttner, M. J., Chater, K. F., & Hopwood, D. A. (2000). Practical Streptomyces Genetics. John Innes Foundation, Norwich Research Park, Colney, Norwich NR4 7UH, England
Doumith, M., Weingarten, P., Wehmeier, U. F., Salah-Bey, K., Benhamou, B., Capdevila, C., Michel, J. M., Piepersberg, W., & Raynal, M. C. (2000). Analysis of genes involved in 6-deoxyhexose biosynthesis and transfer in Saccharopolyspora erythraea. Molecular Genetics and Genomics, 264, 477–485.
Green, M. R., & Sambrook, J. (2012). Molecular cloning. A laboratory manual (4th ed.). Cold Spring Harbor: Cold Spring Harbor Laboratory Press.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72, 248–254.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Percival Zhang, Y.-H., & Lynd, L. R. (2005). Determination of the number-average degree of polymerization of cellodextrins and cellulose with application to enzymatic hydrolysis. Biomacromolecules, 6, 1510–1515.
Lineweaver, H., & Burk, D. (1934). The determination of enzyme dissociation constants. Journal of the American Chemical Society, 56, 658–666.
Petersen, T. N., Brunak, S., von Heijne, G., & Nielsen, H. (2011). SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature Methods, 8, 785–786.
Hazes, B. (1996). The (QxW)3 domain: a flexible lectin scaffold. Protein Science, 5, 1490–1501.
Rutenber, E., Ready, M., & Robertus, J. D. (1987). Structure and evolution of ricin B chain. Nature, 326, 624–626.
Shareck, F., Roy, C., Yaguchi, M., Morosoli, R., & Kluepfel, D. (1991). Sequences of three genes specifying xylanases in Streptomyces lividans. Gene, 107, 75–82.
Fujimoto, Z., Mizuno, H., Kuno, A., Yoshida, S., Kobayashi, H., & Kusakabe, I. (1997). Crystallization and preliminary X-ray crystallographic study of Streptomyces olivaceoviridis E-86 beta-xylanase. Journal of Biochemistry, 121, 826–828.
Lisov, A. V., Belova, O. V., Andreeva-Kovalevskaya, Z. I., Budarina, Z. I., Solonin, A. A., Vinokurova, N. G., & Leontievsky, A. A. (2014). Recombinant xylanase from Streptomyces coelicolor Ac-738: characterization and the effect on xylan-containing products. World Journal of Microbiology and Biotechnology, 30, 801–808.
Polizeli, M. L., Rizzatti, A. C., Monti, R., Terenzi, H. F., Jorge, J. A., & Amorim, D. S. (2005). Xylanases from fungi: properties and industrial applications. Applied Microbiology and Biotechnology, 67, 577–591.
Yang, J., Summanen, P. H., Henning, S. M., Hsu, M., Lam, H., Huang, J., Tseng, C. H., Dowd, S. E., Finegold, S. M., Heber, D., & Li, Z. (2015). Xylooligosaccharide supplementation alters gut bacteria in both healthy and prediabetic adults: a pilot study. Frontiers in Physiology, 6, 216.
Hansen, C. H., Frøkiær, H., Christensen, A. G., Bergström, A., Licht, T. R., Hansen, A. K., & Metzdorff, S. B. (2013). Dietary xylooligosaccharide downregulates IFN-γ and the low-grade inflammatory cytokine IL-1β systemically in mice. Journal of Nutrition, 143, 533–540.
Acknowledgements
This work was supported by the Advanced Biomass R&D Center (ABC) of Global Frontier Project funded by the Ministry of Science, ICT, and Future Planning (NRF-2015M3A6A2065700).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Enkhbaatar, B., Lee, CR., Hong, YS. et al. Molecular Characterization of Xylobiose- and Xylopentaose-Producing β-1,4-Endoxylanase SCO5931 from Streptomyces coelicolor A3(2). Appl Biochem Biotechnol 180, 349–360 (2016). https://doi.org/10.1007/s12010-016-2103-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2103-y