Abstract
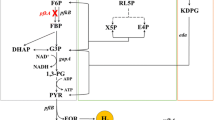
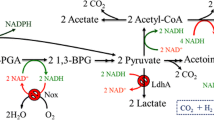
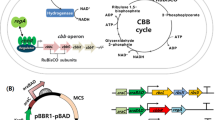
Genetic modifications are considered as one of the most important technologies for improving fermentative hydrogen yield. Herein, we overexpress fhlA and pncB genes from Klebsiella HQ-3 independently to enhance hydrogen molar yield. HQ-3-fhlA/pncB strain is developed by manipulation of pET28-Pkan/fhlA Kanr and pBBR1-MCS5/pncB Gmr as expression vectors to examine the synchronous effects of fhlA and pncB. Optimization of anaerobic batch fermentations is achieved and the maximum yield of biohydrogen (1.42 mol H2/mol of glucose) is produced in the range of pH 6.5–7.0 at 33–37 °C. Whole cell H2 yield is increased up to 40 % from HQ-3-fhlA/pncB, as compared with HQ-3-fhlA 20 % and HQ-3-pncB 12 % keeping HQ-3-C as a control. Mechanism of improved H2 yield is studied in combination with metabolic flux analysis by measuring glucose consumption and other metabolites including formate, succinate, 2,3 butanediol, lactate, acetate, ethanol, and hydrogen. The results suggest that under transient conditions, the increase in the total level of NAD by NAPRTase can enhance the rate of NADH-dependent pathways, and therefore, final distribution of metabolites is changed. Combined overexpression of fhlA and pncB eventually modifies the energy and carbon balance leading to enhanced H2 production from FHL as well as by NADH pathway.






Similar content being viewed by others
References
Mathews, J., & Wang, G. (2009). Metabolic pathway engineering for enhanced biohydrogen production. Int J Hydrog Energy, 34, 7404–16.
Lin, C. Y., & Lay, C. H. (2004). Effects of carbonate and phosphate concentrations on hydrogen production using anaerobic sewage sludge microflora. Int J Hydrog Energy, 29, 275–81.
Oh, S. E., Van Ginkel, S., & Logan, B. E. (2003). The relative effectiveness of pH control and heat treatment for enhancing biohydrogen gas production. Environ Sci Technol, 37, 5186–90.
Kheshgi, H. S., & Prince, R. C. (2005). Sequestration of fermentation CO2 from ethanol production. Energy, 30, 1865–71.
Levin, D. B., Pitt, L., & Love, M. (2004). Biohydrogen production: prospects and limitations to practical application. Int J Hydrog Energy, 29, 173–85.
Brentner, L. B., Peccia, J., & Zimmerman, J. B. (2010). Challenges in developing biohydrogen as a sustainable energy source: implications for a research agenda. Environ Sci Technol, 44, 2243–54.
Koku, H., Eroğlu Gündüz, U., Yücel, M., & Türker, L. (2002). Aspects of the metabolism of hydrogen production by Rhodobacter sphaeroides. Int J Hydrog Energy, 27, 1315–29.
Dubini, A., & Ghirardi, M. L. (2015). Engineering photosynthetic organisms for the production of biohydrogen. Photosynth Res, 123, 241–53.
Maeda, T., Sanchez‐Torres, V., & Wood, T. K. (2012). Hydrogen production by recombinant Escherichia coli strains. J Microbial Biotechnol, 5, 214–25.
Sinha, P., & Pandey, A. (2011). An evaluative report and challenges for fermentative biohydrogen production. Int J Hydrog Energy, 36, 7460–78.
Seol, E., Jang, Y., Kim, S., Oh, Y. K., & Park, S. (2012). Engineering of formate-hydrogen lyase gene cluster for improved hydrogen production in Escherichia coli. Int J Hydrog Energy, 37, 15045–51.
Wang, S., Wang, J., Xu, L., Pi, J., Zhang, H., & Yan, Y. (2013). Enhanced biohydrogen production by homologous over-expression of fnr, pncB, fdhF in Klebsiella sp. HQ-3. Sheng Wu Gong Cheng Xue Bao, 29, 1278–89.
Leonhartsberger, S., Ehrenreich, A., & Böck, A. (2000). Analysis of the domain structure and the DNA binding site of the transcriptional activator FhlA. Eur J Biochem, 267, 3672–84.
Schlensog, V., & Böck, A. (1990). Identification and sequence analysis of the gene encoding the transcriptional activator of the formate hydrogenlyase system of Escherichia coli. Mol Microbiol, 4, 1319–27.
Sanchez-Torres, V., Maeda, T., & Wood, T. K. (2009). Protein engineering of the transcriptional activator FhlA to enhance hydrogen production in Escherichia coli. Appl Environ Microbiol, 75, 5639–46.
Leonhartsberger, S., Korsa, I., & Bock, A. (2002). The molecular biology of formate metabolism in Enterobacteria. J Mol Microbiol Biotechnol, 4, 269–76.
Sauter, M., Bohm, R., & Bock, A. (1992). Mutational analysis of the operon (hyc) determining hydrogenase 3 formation in Escherichia coli. Mol Microbiol, 6, 1523–32.
Liang, L., Liu, R., Wang, G., Gou, D., Ma, J., Chen, K., Jiang, M., Wei, P., & Ouyang, P. (2012). Regulation of NAD(H) pool and NADH/NAD+ ratio by overexpression of nicotinic acid phosphoribosyltransferase for succinic acid production in Escherichia coli NZN111. Enzyme Microb Technol, 51, 286–93.
Wu, H., Li, Z., Zhou, L., & Ye, Q. (2007). Improved succinic acid production in the anaerobic culture of an Escherichia coli pflB ldhA double mutant as a result of enhanced anaplerotic activities in the preceding aerobic culture. Appl Environ Microbiol, 73, 7837–43.
San, K. Y., Bennett, G. N., Berrios-Rivera, S. J., Vadali, R. V., Yang, Y. T., Horton, E., Rudolph, F. B., Sariyar, B., & Blackwood, K. (2002). Metabolic engineering through cofactor manipulation and its effects on metabolic flux redistribution in Escherichia coli. Metab Eng, 4, 182–92.
Berrı, X., Os-Rivera, S. J., San, K. Y., & Bennett, G. N. (2002). The effect of NAPRTase overexpression on the total levels of NAD, the NADH/NAD+ ratio, and the distribution of metabolites in Escherichia coli. Metab Eng, 4, 238–47.
Wang, J., Yu, W., Xu, L., Wang, S., & Yan, Y. (2013). Effects of increasing the NAD(H) pool on hydrogen production and metabolic flux distribution in Enterobacter aerogenes mutants. Int J Hydrog Energy, 38, 13204–15.
Heuser, F., Schroer, K., Lütz, S., Bringer-Meyer, S., & Sahm, H. (2007). Enhancement of the NAD(P)(H) pool in Escherichia coli for biotransformation. Eng Life Sci, 7, 343–53.
Ma, Z., Rao, Z., Zhuge, B., Fang, H., Liao, X., & Zhuge, J. (2010). Construction of a novel expression system in Klebsiella pneumoniae and its application for 1,3-propanediol production. Appl Biochem Biotechnol, 162, 399–407.
Kovach, M. E., Elzer, P. H., Hill, D. S., Robertson, G. T., Farris, M. A., Roop, R. M., & Peterson, K. M. (1995). Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene, 166, 175–6.
Ito, T., Nakashimada, Y., Kakizono, T., & Nishio, N. (2004). High-yield production of hydrogen by Enterobacter aerogenes mutants with decreased alpha-acetolactate synthase activity. J Biosci Bioeng, 97, 227–32.
Maupin, J. A., & Shanmugam, K. T. (1990). Genetic regulation of formate hydrogenlyase of Escherichia coli: role of the fhlA gene product as a transcriptional activator for a new regulatory gene, fhlB. J Bacteriol, 172, 4798–806.
Sawers, R. G. (2005). Formate and its role in hydrogen production in Escherichia coli. Biochem Soc Trans, 33, 42–6.
Si, B., Liu, Z., Zhang, Y., Li, J., Xing, X.-H., Li, B., Duan, N., & Lu, H. (2015). Effect of reaction mode on biohydrogen production and its microbial diversity. Int J Hydrog Energy, 40, 3191–200.
Lee, K. S., Lin, P. J., & Chang, J. S. (2006). Temperature effects on biohydrogen production in a granular sludge bed induced by activated carbon carriers. Int J Hydrog Energy, 31, 465–72.
Wu, K. J., Saratale, G. D., Lo, Y. C., Chen, W. M., Tseng, Z. J., Chang, M. C., Tsai, B. C., Su, A., & Chang, J. S. (2008). Simultaneous production of 2,3-butanediol, ethanol and hydrogen with a Klebsiella sp. strain isolated from sewage sludge. Bioresour Technol, 99, 7966–70.
Sánchez, A. M., Bennett, G. N., & San, K.-Y. (2005). Effect of different levels of NADH availability on metabolic fluxes of Escherichia coli chemostat cultures in defined medium. J Biotechnol, 117, 395–405.
Yu, M., Huang, M., Song, Q., Shao, J., & Ying, X. (2015). Characterization of a (2R, 3R)-2,3-butanediol dehydrogenase from Rhodococcus erythropolis WZ010. Molecules, 20, 7156–73.
Plapp, B. V., Leidal, K. G., Murch, B., Green, P., & David, W. (2015). Contribution of liver alcohol dehydrogenase to metabolism of alcohols in rats. Chem Biol Interact, 234, 85–95.
Yoshida, A., Nishimura, T., Kawaguchi, H., Inui, M., & Yukawa, H. (2006). Enhanced hydrogen production from glucose using ldh- and frd-inactivated Escherichia coli strains. Appl Microbiol Biotechnol, 73, 67–72.
Lu, Y., Zhao, H., Zhang, C., Lai, Q., & Xing, X. (2009). Perturbation of formate pathway for hydrogen production by expressions of formate hydrogen lyase and its transcriptional activator in wild Enterobacter aerogenes and its mutants. Int J Hydrog Energy, 34, 5072–9.
Acknowledgments
This work was supported by the Fundamental Research Funds for the Central Universities (HUST: 2010MS029) and the National Natural Science Foundation of P. R. China (NSFC) (Nos. 31070089 and 31170078). The authors are deeply indebted to the Analytical and Testing Center of Huazhong University of Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Jawed, M., Pi, J., Xu, L. et al. Enhanced H2 Production and Redirected Metabolic Flux via Overexpression of fhlA and pncB in Klebsiella HQ-3 Strain. Appl Biochem Biotechnol 178, 1113–1128 (2016). https://doi.org/10.1007/s12010-015-1932-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1932-4