Abstract
Purpose of Review
To highlight pathways important for the development of autoimmune diabetes by investigating shared mechanisms of disease in polygenic and monogenic diabetes.
Recent Findings
Genome-wide association studies have identified 57 genetic risk loci for type 1 diabetes. Progress has been made in unravelling the mechanistic effects of some of these variants, providing key insights into the pathogenesis of type 1 diabetes. Seven monogenic disorders have also been described where diabetes features as part of an autoimmune syndrome. Studying these genes in relation to polygenic risk loci provides a unique opportunity to dissect pathways important for the development of immune-mediated diabetes.
Summary
Monogenic autoimmune diabetes can result from the dysregulation of multiple pathways suggesting that small effects on many immune processes are required to drive the autoimmune attack on pancreatic beta cells in polygenic type 1 diabetes. A breakdown in central and peripheral immune tolerance is a common theme in the genetic mechanisms of both monogenic and polygenic disease which highlights the importance of these checkpoints in the development and treatment of islet autoimmunity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Type 1 diabetes (T1D) is a complex disease that arises in the context of genetic risk and environmental triggers. Together, these alter immune pathways, resulting in the destruction of insulin-producing pancreatic beta cells. Despite the recognition of an autoimmune aetiology in T1D over 40 years ago [1], knowledge of the triggers and underlying mechanisms of T1D remain incomplete. Efforts to elucidate the cellular pathology of T1D in humans have been hampered by a limited ability to access human pancreatic tissue for direct examination and the immunologic and clinical heterogeneity of the disease.
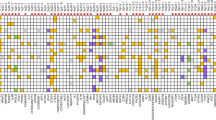
Genetic variation at the HLA region on chromosome 6p21 confers the greatest polygenic risk for the development of T1D. A further 56 non-HLA loci have also been identified but these confer a lower risk for development of the disease [2••, 3]. Many of these genetic loci are associated with additional autoimmune diseases (Table 1), including specific HLA alleles (Table 2). Seven monogenic conditions are also known to cause autoimmune diabetes that is clinically indistinguishable from T1D. In these patients, diabetes usually presents as part of a syndrome of multiple autoimmunity [9].
In this review, we will discuss how studying monogenic autoimmune disease has informed our understanding of mechanisms that contribute to polygenic disease. We will highlight pathways which are shared in the pathogenesis of T1D and other organ specific autoimmunity, focussing on those that have a role in both monogenic forms of autoimmune diabetes and polygenic T1D.
A Breach in Immune Tolerance Is Key to the Development of Autoimmunity
Loss of Central Tolerance
Central tolerance is the process of removing self-reactive T or B cells during their development thus preventing them from targeting normal tissues in the periphery [10]. This process occurs in the thymus for T cells and requires that the T cell receptor (TCR) of a developing T cell is able to bind human leukocyte antigen (HLA) molecules (those reacting to class I HLA go on to form CD8+ T cells, while class II form CD4+ T cells), while limiting the development of T cells with receptors that bind self-peptides in the context of HLA with high affinity. When central tolerance fails, autoimmunity can arise through the release and subsequent action of self-reactive T cells.
Loss of Central Tolerance in Monogenic Disease
The Autoimmune Regulator gene, AIRE, regulates the ectopic expression of self-peptides within the thymus in order to expose naïve T cells to these peptides during development [11]. Loss of function mutations in AIRE (either recessive or dominantly inherited) cause autoimmune polyendocrine syndrome type 1 (APS1, also known as autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy, APECED) by reducing or removing this function of AIRE in the thymus [12]. This allows high affinity autoreactive T cells to escape the thymus. Clinically, APS1 is highly variable but is usually characterised by chronic mucocutaneous candidiasis, adrenal insufficiency and autoimmune hypoparathyroidism. Approximately 13% of individuals develop autoimmune diabetes by 30 years of age [13].
T1D Genetic Risk Loci Involved in Central Tolerance
Variation in the insulin gene (INS) is linked to the development of T1D and is thought to result in a failure of central tolerance. The T1D-associated polymorphic variant is considered to be a variable number of tandem repeats (VNTR), located in the promoter of the INS gene to which AIRE binds, regulating INS RNA expression in the thymus [14,15,16,17,18,19]. VNTR variants of smaller size (class I VNTRs) are associated with increased T1D risk and lower INS mRNA expression in the thymus, allowing escape of insulin autoreactive CD4 T cells into the periphery during T cell development due to fewer insulin peptide-HLA class II interactions. Conversely, insulin autoreactive T cells are predicted to be deleted in individuals carrying the protective INS variants (Class III VNTRs) which drives higher levels of INS expression in the thymus [16]. In keeping with a failure in central tolerance, insulin autoreactive CD4 T cells are present at a higher frequency in the peripheral blood of T1D subjects carrying the INS susceptibility variants, whereas individuals with protective alleles have barely detectable levels of insulin autoreactive CD4 T cells [20].
A failure in central tolerance may also contribute to the association of HLA class II genes to T1D. Although the mechanism is not completely understood, evidence points to low affinity interactions between class II DQ8 molecules and islet peptides, which may result in failed deletion of islet autoreactive CD4 T cells [21, 22]. Another T1D-associated gene, PTPN22, has been linked with failures in both central and peripheral tolerance of T and B cells [23, 24]. A failure of B cell tolerance may be due in part to altered B cell receptor signalling in the presence of the risk variant p.R620W in PTPN22, allowing autoreactive B cells to escape central and peripheral tolerance checkpoints [25]. Although T1D is considered a T cell–mediated disease, B cell pancreatic infiltrate is present in many childhood onset T1D cases [26, 27] and anti-CD20 B cell depleting therapy temporarily slowed disease progression in established T1D [28], indicating a role for B cells in T1D pathogenesis.
Reduced Peripheral Tolerance
Central tolerance is an imperfect process, and as such peripheral tolerance exists to regulate self-reactive cells that escape thymic negative selection. Regulatory T cells (Tregs), a specialised subset of CD4+ T cells, are critical for peripheral tolerance [29, 30]. Tregs suppress conventional T cell (CD8+ and CD4+) activation, proliferation and cytokine production after an immune response to prevent collateral damage to tissues once a pathogen has been removed. There is also growing evidence supporting suppression of B and dendritic cells by Tregs [31, 32].
Reduced Peripheral Tolerance in Monogenic Disease
Reduced function or number of Tregs has been implicated in the disease mechanism of several monogenic autoimmune disorders. Immunodysregulation, polyendocrinopathy, enteropathy, X-linked (IPEX) syndrome highlights the requirement of Tregs to restrain autoimmunity. IPEX syndrome, which often proves fatal in early life, results from hemizygous mutations in the FOXP3 gene, a key regulator of Treg development [33]. The syndrome typically presents in the neonatal period, with > 90% of affected boys having severe protein losing enteropathy and ~ 80% developing autoimmune diabetes. Additional autoimmune diseases can develop including severe atopic dermatitis (70%) and autoimmune hypothyroidism (35%) [34].
Individuals with recessively inherited IL2RA mutations develop immunodeficiency 31C syndrome [35], which is similar to IPEX syndrome (enteropathy, hypothyroidism and severe eczema) and can include neonatal-onset autoimmune diabetes [36]. IL2RA encodes CD25, a subunit of the IL-2 receptor which is constitutively and highly expressed on regulatory T cells facilitating their recruitment and suppressive ability. IL-2 is a key signalling molecule involved in regulating the immune system and induces FOXP3 expression; therefore, loss of the IL-2 receptor on T cells reduces FOXP3 expression and Treg development [37]. As in IPEX, mutations that lead to a loss of CD25 expression result in a reduced Treg compartment, promoting autoimmunity through the failure of peripheral tolerance [38].
Heterozygous mutations in CTLA4 cause autoimmune lymphoproliferative syndrome which can include autoimmune diabetes [39,40,41]. CTLA4 is constitutively expressed by Tregs and can also be expressed by CD4+ and CD8+ effector T cells where it functions as a potent suppressive receptor molecule, by preventing co-activation of T effector cells via CD28 [42]. It mediates inhibition in effector T cells by competing with CD28 for binding to CD80/CD86 on antigen presenting cells (APC) but may also inhibit T cell receptor signalling [31, 43, 44]. Tregs from individuals with dominantly inherited CTLA4 mutations show reduced expression of CTLA4, FOXP3 and IL2RA [39].
Recessively inherited mutations in LRBA cause common variable immunodeficiency-8 (CVID8) with autoimmunity [45]. This includes extremely young onset of haematological autoimmune disorders (80%), enteropathy (70%) and autoimmune diabetes (30%) [46]. LRBA plays an essential role in the post-translational regulation and trafficking of CTLA4 (see above), whereby it prevents lysosomal degradation of CTLA4 containing vesicles [47•]. Interestingly, studies have identified six individuals with LRBA mutations who had a reduced number of Tregs [48] and an individual with normal cell-surface CTLA4 expression [49]. In the latter patient, there was increased Th17 cell activity (measured by IL-17 production) suggesting that the disease could also be mediated through effector cells.
Gain of function (GOF) mutations in STAT3, which links extracellular cytokine signals to gene expression, cause infancy-onset multiple autoimmune disease [50]. These mutations cause haematological autoimmune disorders (70%), enteropathies (50%) and autoimmune diabetes in ~ 30% of individuals which often presents in the neonatal period. Some patients present with similar features to autoimmune lymphoproliferative syndrome (ALPS) [51, 52]. STAT3 is involved in multiple signalling pathways that influence the fate of CD4 T cells, enhancing development of Th17 and T follicular helper cells, while blocking the development and survival of regulatory T cells. Tregs are numerically and functionally reduced in most individuals with GOF STAT3 mutations, while Th17 cells may be normal, reduced or increased [53].
T1D Genetic Risk Loci Involved in Peripheral Tolerance
Genetic variants in genes that function in the IL-2 pathway are associated with T1D, including IL2RA (described above) and PTPN2 which encodes a non-receptor tyrosine phosphatase that regulates IL-2 signalling [2••, 54, 55]. The most highly associated IL2RA single nucleotide polymorphism (SNP), rs61839660, is non-coding and located in an enhancer region in intron 1 of the gene. The enhancer binds multiple transcription factors and interacts with other regulatory elements in the locus in primary CD4 T cells, but only in response to T cell stimulation [56••]. The presence of the T1D risk allele at the enhancer resulted in enhanced CD25 (IL2RA) upregulation in CD4 conventional T cells but not Tregs in a knock-in mouse model [56••], indicating that this SNP primarily impacts CD4 conventional T cells.
Alternatively, several other IL2RA SNPs have been associated with decreased CD25 expression on CD4 conventional T cells and Tregs, and increased levels of soluble CD25 with the risk alleles, revealing the complexity of the IL2RA locus [57,58,59]. Functionally, this correlates with reduced IL-2 signalling and diminished Treg fitness and suppressive function [58, 59]. The T1D-associated SNPs in the PTPN2 gene are also non-coding and have been correlated with decreased PTPN2 RNA levels and reduced IL-2 signalling in genotyped healthy control subjects and longstanding T1D patients [60, 61]. The PTPN2 T1D risk allele was also associated with decreased FOXP3 expression in activated CD4 T cells [60]. The effects of these IL2RA and PTPN2 T1D risk alleles on IL-2 signalling are independent but additive, both potentially contributing to reduced peripheral tolerance through effects on Tregs [58]. An additional T1D-associated SNP is located in the IL2-IL21 intergenic region, although the impact of this SNP has not been evaluated yet [2••].
A non-coding SNP rs3087243 located 3′ of the CTLA4 gene has been associated with T1D, as well as other autoimmune diseases [2••] (Table 1). How the CTLA4 rs3087243 SNP affects CTLA4 function is not completely understood. Initial studies indicated that the rs3087243 variant affected CTLA4 alternative splicing, resulting in lower levels of a soluble CTLA4 isoform in CD4 T cells carrying the T1D susceptibility allele [62]. However, this finding was not replicated in a subsequent study [63]. More recently the rs3087243 SNP was shown to be in high linkage disequilibrium with an (AT)n dinucleotide repeat in the 3′ untranslated region of CTLA4, with the T1D susceptibility allele associated with longer (AT)n repeat length compared with the non-risk allele [64]. Human islet autoreactive T cell lines with longer (AT)n repeats expressed lower levels of CTLA4 RNA and protein relative to T cell lines with shorter repeats, and longer (AT)n repeats destabilised a GFP reporter expressed in Jurkat T cells [64]. Confirmation of these findings in rs3087243 genotyped peripheral blood CD4 T cells and elucidation of corresponding functional phenotypes will clarify the mechanism of the T1D association with CTLA4.
Defects in Interferon Signalling Pathways
Interferons (IFNs) are cytokines released by mononuclear cells in response to the presence of pathogens and tumours that modulate the immune system. They have a role in the regulation of immune responses, activating natural killer cells and macrophages and upregulating antigen presentation by HLA molecules [65]. Type 1 IFNs (IFN-1) are central to the anti-viral response, yet the presence of an IFN signature is well described in multiple autoimmune diseases including T1D [66, 67]. Perturbed IFN signalling is associated with systemic lupus erythematosus, a systemic autoimmune condition directed against ubiquitous proteins such as those within the cell nucleus [68]. Indeed, an IFN signature consisting of expression levels of type 1 IFN-responsive genes correlates with the severity of disease in SLE [69]. The mechanistic role of IFN signalling in the induction of organ specific autoimmunity such as T1D remains unclear although evidence is mounting that they may induce increased autoantigen presentation by islet cells and thus increase recognition and activation of effector T cells [70]. Studies on deceased T1D patients’ pancreata have shown increased expression of IFN alpha [71, 72]; however, whether this was secondary to, for example, viral infection of the islets, remains unknown. The role of IFNs in organ specific autoimmunity is further supported by the triggering of autoimmune disease, including diabetes, in patients undergoing interferon treatment for malignancies [73, 74].
Defects in IFN Signalling in Monogenic Disease
Patients with dominant GOF mutations in STAT1 present with chronic mucocutaneous candidiasis and lower respiratory tract infections with a subset of patients developing severe organ-specific autoimmunity including T1D (4% of patients) [75]. STAT1 responds to multiple cytokines to translate extracellular signals to gene transcription. IFN alpha induces the transcription of numerous targets via the JAK-STAT pathway increasing the proliferation of immune cells, as well as augmenting other immune processes such as antibody production. Increased activity of STAT1, as a result of impaired nuclear de-phosphorylation, is proposed to lead to autoimmunity by increasing IFN alpha signalling.
T1D Genetic Risk Loci Involved in IFN Signalling
Two coding variants in genes within the interferons pathway, IFIH1 A946T and TYK2 P1104A, support a role of IFN-1 in the development of T1D [2••, 76].
IFIH1 (interferon-induced helicase C-domain-containing protein 1, also known as MDA5) is a component of the innate response to RNA viruses. Upon recognition of double-stranded RNA, IFIH1 undergoes a conformational change that result in a series of signalling events leading to transcription of genes encoding IFN-1 and interferon-stimulated genes (ISGs). Rare GOF missense mutations in IFIH1 are described which result in interferonopathies [77]. A rare loss of function mutation in the IFIH1 gene is associated with protection from T1D [78]. The common variant rs1990760, a non-synonymous coding variant in IFIH1(A946T), is associated with risk for T1D [76] as well as other autoimmune diseases including SLE [79]. This variant results in enhanced basal expression of IFN-1 and improved response to viral challenge indicating that the IFIH1 risk variant 946T is a gain-of-function variant that is triggered by RNA self-ligands as well as viral infection [80]. This variant has likely been selected in the population as it provides an advantage in the setting of viral infection, despite the fact that it also promotes the risk of autoimmunity.
TYK2 encodes a JAK family kinase that functions to mediate proximal IFN-1-, IL-12- and IL-23-dependent signals [81, 82]. TYK2 deficient patients exhibit susceptibility to viral infections and impairment in cellular responses to IFN-1, as well as mycobacterial infections consistent with an impaired response to IL-12 and IL-23. A missense change within the kinase domain of TYK2, with substitution of alanine for a conserved proline (TYK2-P1104A) is associated with a lack of catalytic activity and protection for T1D [2••, 83]. The protective TYK2 1104A allele results in impaired cellular responses to IFN-1, IL-12 and IL-23 and notably leads to a decrease in the induction of experimental autoimmune encephalomyelitis in murine models and a striking decrease in IL-17/IFN-γ positive CD4 T cells [83, 84]. In humans carrying the TYK2-1104A allele, IFN-1 receptor signalling is decreased, and alterations in memory B cell populations and a decrease in Tfh cells are seen in the peripheral blood [84, 85]. Similar to TYK2 deficient patients, individuals homozygous for the TYK2 1104A variant have an increased frequency of tuberculosis [83]. The incidence of the TYK2 1104A allele has decreased over the past 4000 years in Europeans, consistent with negative selection of this allele by tuberculosis infection [83]. Thus, the T1D risk allele TYK2 P1104 is increasing in frequency in European populations. Taken together, these studies suggest that alterations in IFN signalling in T1D may play a role in disease, but in the case of the TYK2 variant, this is in the broader context of other cytokine responses.
Conclusions
The finding that monogenic autoimmune disease can result from the dysregulation of multiple immune pathways suggests that small effects on multiple processes may be required to drive the autoimmune attack on pancreatic beta cells in polygenic T1D. A common theme in the genetic pathways identified in diabetes and other autoimmunity is the breakdown of immune tolerance; however, the part of the pathway perturbed can be different. Monogenic disorders include those which disrupt central tolerance (AIRE), or impact the development (FOXP3, STAT3) or function (IL2RA, LRBA, CTLA4) of regulatory T cells required for the maintenance of peripheral tolerance. The risk alleles identified by genome-wide association studies mirror this, with effects on the ectopic expression of insulin, reducing central tolerance (e.g. INS VNTR), and reduced expression of genes essential for the maintenance of peripheral tolerance also identified (e.g. IL2RA, PTPN2, CTLA4).
Defects in interferon signalling are increasingly recognised as being involved in multiple autoimmune diseases. While diabetes is a rare feature of monogenic interferonopathies, two loci within the IFN pathway (IFIH-1 and TYK2) have been robustly associated with polygenic T1D. Furthermore, the presence of an IFN signature in some patients with T1D, increased expression of IFN in pancreata from deceased T1D patients and mounting evidence of viral involvement in some T1D cases highlight that defects in IFN signalling may be an important contributor to the pathophysiology of T1D. Further molecular characterisation of the mechanisms by which common risk loci that impart risk for T1D and of genes causing monogenic autoimmunity will be the key to detecting the pathways that underlie this shared aetiology.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Bottazzo GF, Florin-Christensen A, Doniach D. Islet-cell antibodies in diabetes mellitus with autoimmune polyendocrine deficiencies. Lancet (London, England) [Internet]. 1974 Nov 30 [cited 2019 Jan 4];2(7892):1279–83. Available from: http://www.ncbi.nlm.nih.gov/pubmed/4139522
•• Onengut-Gumuscu S, Chen W-M, Burren O, Cooper NJ, Quinlan AR, Mychaleckyj JC, et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat Genet [Internet]. 2015 Apr 9 [cited 2019 Jan 4];47(4):381–6. Available from: http://www.nature.com/articles/ng.3245 . Fine-mapped susceptibility loci in T1D and showed that T1D was more genetically similar to antibody-driven autoimmune diseases than non-antibody-associated.
Barrett JC, Clayton DG, Concannon P, Akolkar B, Cooper JD, Erlich HA, et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet [Internet]. 2009 Jun 10 [cited 2019 Jan 4];41(6):703–7. Available from: http://www.nature.com/articles/ng.381
Andlauer TFM, Buck D, Antony G, Bayas A, Bechmann L, Berthele A, et al. Novel multiple sclerosis susceptibility loci implicated in epigenetic regulation. Sci Adv [Internet]. 2016 Jun [cited 2019 Jan 30];2(6):e1501678. Available from: https://doi.org/10.1126/sciadv.1501678
Erlich H, Valdes AM, Noble J, Carlson JA, Varney M, Concannon P, et al. HLA DR-DQ haplotypes and genotypes and type 1 diabetes risk: analysis of the type 1 diabetes genetics consortium families. Diabetes [Internet]. 2008 Apr 1 [cited 2019 Jan 8];57(4):1084–92. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18252895
Liu E, Lee H-S, Aronsson CA, Hagopian WA, Koletzko S, Rewers MJ, et al. Risk of pediatric celiac disease according to HLA haplotype and country. N Engl J Med [Internet]. 2014 Jul 3 [cited 2019 Jan 8];371(1):42–9. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24988556
Zamani M, Spaepen M, Bex M, Bouillon R, Cassiman JJ. Primary role of the HLA class II DRB1*0301 allele in Graves disease. Am J Med Genet [Internet]. 2000 Dec 18 [cited 2019 Jan 8];95(5):432–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/11146462
Stahl EA, Raychaudhuri S, Remmers EF, Xie G, Eyre S, Thomson BP, et al. Genome-wide association study meta-analysis identifies seven new rheumatoid arthritis risk loci. Nat Genet [Internet]. 2010 Jun [cited 2019 Jan 30];42(6):508–14. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20453842
Johnson MB, Hattersley AT, Flanagan SE. Monogenic autoimmune diseases of the endocrine system 2016 Oct [cited 2019 Jan 4];4(10):862–72. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27474216
Hogquist KA, Baldwin TA, Jameson SC. Central tolerance: learning self-control in the thymus. Nat Rev Immunol [Internet]. 2005 Oct 1 [cited 2019 Jan 4];5(10):772–82. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16200080
Zhao B, Chang L, Fu H, Sun G, Yang W. The role of autoimmune regulator (AIRE) in peripheral tolerance. J Immunol Res [Internet]. 2018 Sep 4 [cited 2019 Jan 4];2018:1–6. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30255105
Aaltonen J, Björses P, Perheentupa J, Horelli-Kuitunen N, Palotie A, Peltonen L, et al. An autoimmune disease, APECED, caused by mutations in a novel gene featuring two PHD-type zinc-finger domains. Nat Genet [Internet]. 1997 Dec [cited 2019 Jan 4];17(4):399–403. Available from: http://www.ncbi.nlm.nih.gov/pubmed/9398840
Husebye ES, Perheentupa J, Rautemaa R, Kämpe O. Clinical manifestations and management of patients with autoimmune polyendocrine syndrome type i. J Intern Med [Internet]. 2009 May [cited 2019 Jan 4];265(5):514–29. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19382991
Lucassen AM, Julier C, Beressi JP, Boitard C, Froguel P, Lathrop M, et al. Susceptibility to insulin dependent diabetes mellitus maps to a 4.1 kb segment of DNA spanning the insulin gene and associated VNTR. Nat Genet [Internet]. 1993 Jul [cited 2019 Jan 4];4(3):305–10. Available from: http://www.nature.com/articles/ng0793-305
Owerbach D, Gabbay KH. Localization of a type I diabetes susceptibility locus to the variable tandem repeat region flanking the insulin gene. Diabetes [Internet]. 1993 Dec [cited 2019 Jan 4];42(12):1708–14. Available from: http://www.ncbi.nlm.nih.gov/pubmed/8243816
Pugliese A, Zeller M, Fernandez A, Zalcberg LJ, Bartlett RJ, Ricordi C, et al. The insulin gene is transcribed in the human thymus and transcription levels correlated with allelic variation at the INS VNTR-IDDM2 susceptibility locus for type 1 diabetes. Nat Genet [Internet]. 1997 Mar [cited 2019 Jan 4];15(3):293–7. Available from: http://www.nature.com/articles/ng0397-293
Vafiadis P, Bennett ST, Todd JA, Nadeau J, Grabs R, Goodyer CG, et al. Insulin expression in human thymus is modulated by INS VNTR alleles at the IDDM2 locus. Nat Genet [Internet]. 1997 Mar [cited 2019 Jan 4];15(3):289–92. Available from: http://www.nature.com/articles/ng0397-289
Cai CQ, Zhang T, Breslin MB, Giraud M, Lan MS. Both polymorphic variable number of tandem repeats and autoimmune regulator modulate differential expression of insulin in human thymic epithelial cells. Diabetes [Internet]. 2011 Jan 1 [cited 2019 Jan 4];60(1):336–44. Available from: https://doi.org/10.2337/db10-0255
Sparks AE, Chen C, Breslin MB, Lan MS. Functional domains of Autoimmune Regulator (AIRE) modulate INS-VNTR transcription in human thymic epithelial cells. J Biol Chem [Internet]. 2016 May 20 [cited 2019 Jan 4];291(21):11313–22. Available from: https://doi.org/10.1074/jbc. M116.722488.
Durinovic-Belló I, Wu RP, Gersuk VH, Sanda S, Shilling HG, Nepom GT. Insulin gene VNTR genotype associates with frequency and phenotype of the autoimmune response to proinsulin. Genes Immun [Internet]. 2010 Mar 7 [cited 2019 Jan 4];11(2):188–93. Available from: http://www.nature.com/articles/gene2009108
Lee KH, Wucherpfennig KW, Wiley DC. Structure of a human insulin peptide-HLA-DQ8 complex and susceptibility to type 1 diabetes. Nat Immunol [Internet]. 2001 Jun 1 [cited 2019 Jan 4];2(6):501–7. Available from: https://doi.org/10.1038/88694
Yang J, Chow ITIT, Sosinowski T, Torres-Chinn N, Greenbaum CJ, James EA, et al. Autoreactive T cells specific for insulin B:11-23 recognize a low-affinity peptide register in human subjects with autoimmune diabetes. Proc Natl Acad Sci USA [Internet]. 2014 Oct 14 [cited 2019 Jan 4];111(41):14840–5. Available from: https://doi.org/10.1073/pnas.1416864111
Rawlings DJ, Dai X, Buckner JH. The role of PTPN22 risk variant in the development of autoimmunity: finding common ground between mouse and human. J Immunol [Internet]. 2015 Apr 1 [cited 2019 Jan 4];194(7). Available from: https://doi.org/10.4049/jimmunol.1403034
Menard L, Saadoun D, Isnardi I, Ng Y-S, Meyers G, Massad C, et al. The PTPN22 allele encoding an R620W variant interferes with the removal of developing autoreactive B cells in humans. J Clin Invest [Internet]. 2011 Sep 1 [cited 2019 Jan 4];121(9). Available from: http://www.jci.org/articles/view/45790
Habib T, Funk A, Rieck M, Brahmandam A, Dai X, Panigrahi AK, et al. Altered B cell homeostasis is associated with type I diabetes and carriers of the PTPN22 allelic variant. J Immunol [Internet]. 2012 Jan 1 [cited 2019 Jan 4];188(1). Available from: https://doi.org/10.4049/jimmunol.1102176
Leete P, Willcox A, Krogvold L, Dahl-Jørgensen K, Foulis AK, Richardson SJ, et al. Differential insulitic profiles determine the extent of β-cell destruction and the age at onset of type 1 diabetes. Diabetes [Internet]. 2016 May [cited 2019 Jan 4];65(5). Available from: https://doi.org/10.2337/db15-1615
Arif S, Leete P, Nguyen V, Marks K, Nor NM, Estorninho M, et al. Blood and islet phenotypes indicate immunological heterogeneity in type 1 diabetes. Diabetes [Internet]. 2014 Nov [cited 2019 Jan 4];63(11). Available from: https://doi.org/10.2337/db14-0365
Pescovitz MD, Greenbaum CJ, Krause-Steinrauf H, Becker DJ, Gitelman SE, Goland R, et al. Rituximab, B-lymphocyte depletion, and preservation of beta-cell function. N Engl J Med [Internet]. 2009 Nov 26 [cited 2019 Jan 4];361(22):2143–52. Available from: https://doi.org/10.1056/NEJMoa0904452
Schmetterer KG, Neunkirchner A, Pickl WF. Naturally occurring regulatory T cells: markers, mechanisms, and manipulation. FASEB J [Internet]. 2012 Jun 23 [cited 2019 Jan 4];26(6):2253–76. Available from: https://doi.org/10.1096/fj.11-193672
Sakaguchi S, Miyara M, Costantino CM, Hafler DA. FOXP3+ regulatory T cells in the human immune system. Nat Rev Immunol [Internet]. 2010 Jul 18 [cited 2019 Jan 4];10(7):490–500. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20559327
Chen J, Ganguly A, Mucsi AD, Meng J, Yan J, Detampel P, et al. Strong adhesion by regulatory T cells induces dendritic cell cytoskeletal polarization and contact-dependent lethargy. J Exp Med [Internet]. 2017 Feb [cited 2019 Jan 4];214(2):327–38. Available from: https://doi.org/10.1084/jem.20160620
Xu A, Liu Y, Chen W, Wang J, Xue Y, Huang F, et al. TGF-β–induced regulatory T cells directly suppress B cell responses through a noncytotoxic mechanism. J Immunol [Internet]. 2016 May 1 [cited 2019 Jan 4];196(9):3631–41. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27001954
Chatila TA, Blaeser F, Ho N, Lederman HM, Voulgaropoulos C, Helms C, et al. JM2, encoding a fork head-related protein, is mutated in X-linked autoimmunity-allergic disregulation syndrome. J Clin Invest [Internet]. 2000 Dec 15 [cited 2019 Jan 4];106(12):R75–81. Available from: http://www.jci.org/articles/view/11679
d’Hennezel E, Dhuban K Bin, Torgerson T, Piccirillo CA, Bin Dhuban K, Torgerson T, et al. The immunogenetics of immune dysregulation, polyendocrinopathy, enteropathy, X linked (IPEX) syndrome. J Med Genet [Internet]. 2012 May 11 [cited 2019 Jan 4];49(5):291–302. Available from: https://doi.org/10.1136/jmedgenet-2012-100759
Sharfe N, Dadi HK, Shahar M, Roifman CM. Human immune disorder arising from mutation of the alpha chain of the interleukin-2 receptor. Proc Natl Acad Sci U S A [Internet]. 1997 Apr 1 [cited 2019 Jan 4];94(7):3168–71. Available from: http://www.ncbi.nlm.nih.gov/pubmed/9096364
Caudy AA, Reddy ST, Chatila T, Atkinson JP, Verbsky JW. CD25 deficiency causes an immune dysregulation, polyendocrinopathy, enteropathy, X-linked-like syndrome, and defective IL-10 expression from CD4 lymphocytes. J Allergy Clin Immunol [Internet]. 2007 Feb [cited 2019 Jan 4];119(2):482–7. Available from: http://linkinghub.elsevier.com/retrieve/pii/S0091674906021191
Zorn E, Nelson EA, Mohseni M, Porcheray F, Kim H, Litsa D, et al. IL-2 regulates FOXP3 expression in human CD4+CD25+ regulatory T cells through a STAT-dependent mechanism and induces the expansion of these cells in vivo. Blood [Internet]. 2006 May 9 [cited 2019 Jan 4];108(5):1571–9. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16645171
Bezrodnik L, Caldirola MS, Seminario AG, Moreira I, Gaillard MI. Follicular bronchiolitis as phenotype associated with CD25 deficiency. Clin Exp Immunol [Internet]. 2014 Feb [cited 2019 Jan 4];175(2):227–34. Available from: https://doi.org/10.1111/cei.12214
Schwab C, Gabrysch A, Olbrich P, Patiño V, Warnatz K, Wolff D, et al. Phenotype, penetrance, and treatment of 133 cytotoxic T-lymphocyte antigen 4–insufficient subjects. J Allergy Clin Immunol. 2018;1–15.
Kuehn HS, Ouyang W, Lo B, Deenick EK, Niemela JE, Avery DT, et al. Immune dysregulation in human subjects with heterozygous germline mutations in CTLA4. Science (80- ). 2014;345(6204):1623–7.
Schubert DD, Bode C, Kenefeck R, Hou TZ, Wing JB, Kennedy A, et al. Autosomal dominant immune dysregulation syndrome in humans with CTLA4 mutations. Nat Med [Internet]. 2014 Dec 20 [cited 2019 Jan 4];20(12):1410–6. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25329329
Sansom DM, Walker LSK The role of CD28 and cytotoxic T-lymphocyte antigen-4 (CTLA-4) in regulatory T-cell biology. Immunol Rev [Internet]. 2006 Aug [cited 2019 Jan 4];212(1):131–48. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16903911
Collins A V, Brodie DW, Gilbert RJC, Iaboni A, Manso-Sancho R, Walse B, et al. The interaction properties of costimulatory molecules revisited. Immunity [Internet]. 2002 Aug [cited 2019 Jan 4];17(2):201–10. Available from: http://www.ncbi.nlm.nih.gov/pubmed/12196291
Walunas TL, Lenschow DJ, Bakker CY, Linsley PS, Freeman GJ, Green JM, et al. CTLA-4 can function as a negative regulator of T cell activation. Immunity [Internet]. 1994 Aug [cited 2019 Jan 4];1(5):405–13. Available from: http://www.ncbi.nlm.nih.gov/pubmed/7882171
Lopez-Herrera G, Tampella G, Pan-Hammarström Q, Herholz P, Trujillo-Vargas CMM, Phadwal K, et al. Deleterious mutations in LRBA are associated with a syndrome of immune deficiency and autoimmunity. Am J Hum Genet [Internet]. 2012 Jun 8 [cited 2019 Jan 4];90(6):986–1001. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22608502
Johnson MB, De Franco E, Lango Allen H, Al Senani A, Elbarbary N, Siklar Z, et al. Recessively inherited LRBA mutations cause autoimmunity presenting as neonatal diabetes. Diabetes [Internet]. 2017 Aug [cited 2019 Jan 4];66(8):2316–22. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28473463
• Lo B, Zhang K, Lu W, Zheng L, Zhang Q, Kanellopoulou C, et al. Patients with LRBA deficiency show CTLA4 loss and immune dysregulation responsive to abatacept therapy. Science (80). 2015;349(6246):436–40 An elegant study that defined the role of LRBA in maintaining tolerance by repurposing a known therapeutic agent, abatacept, simultaneously providing a personalised therapy for patients.
Charbonnier A-MM, Janssen E, Chou J, Ohsumi TK, Keles S, Hsu JT, et al. Regulatory T-cell deficiency and immune dysregulation, polyendocrinopathy, enteropathy, X-linked-like disorder caused by loss-of-function mutations in LRBA. J Allergy Clin Immunol [Internet]. 2015 Jan [cited 2019 Jan 4];135(1):217–27. Available from: https://doi.org/10.1016/j.jaci.2014.10.019
De Bruyne M, Bogaert DJ, Venken K, Van den Bossche L, Bonroy C, Roels L, et al. A novel LPS-responsive beige-like anchor protein (LRBA) mutation presents with normal cytotoxic T lymphocyte-associated protein 4 (CTLA-4) and overactive TH17 immunity. J Allergy Clin Immunol [Internet]. 2018 Dec [cited 2019 Jan 4];142(6):1968–71. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30193839
Flanagan SE, Haapaniemi E, Russell MA, Caswell R, Allen HL, De Franco E, et al. Activating germline mutations in STAT3 cause early-onset multi-organ autoimmune disease. Nat Genet [Internet]. 2014 Aug 20 [cited 2019 Jan 4];46(8):812–4. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25038750
Milner JD, Vogel TP, Forbes L, Ma CA, Stray-Pedersen A, Niemela JE, et al. Early-onset lymphoproliferation and autoimmunity caused by germline STAT3 gain-of-function mutations. Blood [Internet]. 2015 Jan 22 [cited 2019 Jan 4];125(4):591–9. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25359994
Nabhani S, Schipp C, Miskin H, Levin C, Postovsky S, Dujovny T, et al. STAT3 gain-of-function mutations associated with autoimmune lymphoproliferative syndrome like disease deregulate lymphocyte apoptosis and can be targeted by BH3 mimetic compounds. Clin Immunol [Internet]. 2017 Aug [cited 2019 Jan 4];181:32–42. Available from: https://linkinghub.elsevier.com/retrieve/pii/S1521661616304478
Olbrich P, Freeman AF. STAT1 and STAT3 mutations: important lessons for clinical immunologists. Expert Rev Clin Immunol [Internet]. 2018 Dec 2 [cited 2019 Jan 4];14(12):1029–41. Available from: https://doi.org/10.1080/1744666X.2018.1531704
Todd JA, Walker NM, Cooper JD, Smyth DJ, Downes K, Plagnol V, et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat Genet [Internet]. 2007 Jul 6 [cited 2019 Jan 4];39(7):857–64. Available from: http://www.nature.com/articles/ng2068
Farh KK-H, Marson A, Zhu J, Kleinewietfeld M, Housley WJ, Beik S, et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature [Internet]. 2015 Feb 19 [cited 2019 Jan 4];518(7539):337–43. Available from: http://www.nature.com/articles/nature13835
•• Simeonov DR, Gowen BG, Boontanrart M, Roth TL, Gagnon JD, Mumbach MR, et al. Discovery of stimulation-responsive immune enhancers with CRISPR activation. Nature [Internet]. 2017 Aug 7 [cited 2019 Jan 4];549(7670):111–5. Available from: https://doi.org/10.1038/nature23875. Provides a framework for identifying stimulus-responsive enhancers for genes, independent of exposure to the exposure to the stimulus. This will be key to elucidating the functional effects of many susceptibility loci in autoimmune and other diseases.
Dendrou CA, Plagnol V, Fung E, Yang JHM, Downes K, Cooper JD, et al. Cell-specific protein phenotypes for the autoimmune locus IL2RA using a genotype-selectable human bioresource. Nat Genet [Internet]. 2009 Sep 23 [cited 2019 Jan 4];41(9):1011–5. Available from: http://www.nature.com/articles/ng.434
Cerosaletti K, Schneider A, Schwedhelm K, Frank I, Tatum M, Wei S, et al. Multiple autoimmune-associated variants confer decreased IL-2R signaling in CD4+ CD25(hi) T cells of type 1 diabetic and multiple sclerosis patients. Jacobson S, editor. PLoS One [Internet]. 2013 Dec 23 [cited 2019 Jan 4];8(12):e83811. Available from: https://doi.org/10.1371/journal.pone.0083811
Garg G, Tyler JR, Yang JHM, Cutler AJ, Downes K, Pekalski M, et al. Type 1 diabetes-associated IL2RA variation lowers IL-2 signaling and contributes to diminished CD4+CD25+ regulatory T cell function. J Immunol [Internet]. 2012 May 1 [cited 2019 Jan 4];188(9). Available from: https://doi.org/10.4049/jimmunol.1100272
Long SA, Cerosaletti K, Wan JY, Ho J-C, Tatum M, Wei S, et al. An autoimmune-associated variant in PTPN2 reveals an impairment of IL-2R signaling in CD4(+) T cells. Genes Immun [Internet]. 2011 Mar 23 [cited 2019 Jan 4];12(2). Available from: http://www.nature.com/articles/gene201054
Yang JHM, Cutler AJ, Ferreira RC, Reading JL, Cooper NJ, Wallace C, et al. Natural Variation in Interleukin-2 sensitivity influences regulatory T-Cell frequency and function in individuals with long-standing type 1 diabetes. Diabetes [Internet]. 2015 Nov [cited 2019 Jan 4];64(11):3891–902. Available from: https://doi.org/10.2337/db15-0516
Ueda H, Howson JMM, Esposito L, Heward J, Snook H, Chamberlain G, et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease Nature [Internet]. 2003 May 29 [cited 2019 Jan 4];423(6939):506–11. Available from: http://www.nature.com/articles/nature01621
Mayans S, Lackovic K, Nyholm C, Lindgren P, Ruikka K, Eliasson M, et al. CT60 genotype does not affect CTLA-4 isoform expression despite association to T1D and AITD in northern Sweden. BMC Med Genet [Internet]. 2007 Feb 6 [cited 2019 Jan 4];8(1):3. Available from: https://doi.org/10.1186/1471-2350-8-3
de Jong VM, Zaldumbide A, van der Slik AR, Laban S, Koeleman BPC, Roep BO. Variation in the CTLA4 3’UTR has phenotypic consequences for autoreactive T cells and associates with genetic risk for type 1 diabetes. Genes Immun [Internet]. 2016 Jan 10 [cited 2019 Jan 4];17(1):75–8. Available from: http://www.nature.com/articles/gene201551
de Weerd NA, Samarajiwa SA, Hertzog PJ. Type I interferon receptors: biochemistry and biological functions. J Biol Chem [Internet]. 2007 Jul 13 [cited 2019 Jan 7];282(28):20053–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17502368
Theofilopoulos AN, Baccala R, Beutler B, Kono DH. Type I interferons (alpha/beta) in immunity and autoimmunity. Annu Rev Immunol [Internet]. 2005 Apr [cited 2019 Jan 7];23(1):307–36. Available from: https://doi.org/10.1146/annurev.immunol.23.021704.115843
Ferreira RC, Guo H, Coulson RMR, Smyth DJ, Pekalski ML, Burren OS, et al. A Type I interferon transcriptional signature precedes autoimmunity in children genetically at risk for type 1 diabetes. Diabetes [Internet]. 2014 Jul 1 [cited 2019 Jan 7];63(7):2538–50. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24561305
Kaul A, Gordon C, Crow MK, Touma Z, Urowitz MB, van Vollenhoven R, et al. Systemic lupus erythematosus. Nat Rev Dis Prim [Internet]. 2016 Jun 16 [cited 2019 Jan 7];2:16039. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27306639
Bezalel S, Guri KM, Elbirt D, Asher I, Sthoeger ZM. Type I interferon signature in systemic lupus erythematosus. Isr Med Assoc J [Internet]. 2014 Apr [cited 2019 Jan 7];16(4):246–9. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24834763
Lombardi A, Tsomos E, Hammerstad SS, Tomer Y. Interferon alpha: The key trigger of type 1 diabetes. J Autoimmun [Internet]. 2018 Nov [cited 2019 Jan 7];94:7–15. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30115527
Foulis AK, Liddle CN, Farquharson MA, Richmond JA, Weir RS. Diabetologia in the United Kingdom. Diabetologia. 1986;1:267–74.
Huang X, Yuang J, Goddard A, Foulis A, James RF, Lernmark A, et al. Interferon expression in the pancreases of patients with type I diabetes. Diabetes [Internet]. 1995 Jun [cited 2019 Jan 7];44(6):658–64. Available from: http://www.ncbi.nlm.nih.gov/pubmed/7540571
Nakamura K, Kawasaki E, Imagawa A, Awata T, Ikegami H, Uchigata Y, et al. Type 1 diabetes and interferon therapy: a nationwide survey in Japan. Diabetes Care [Internet]. 2011 Sep 1 [cited 2019 Jan 7];34(9):2084–9. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21775762
Zornitzki T, Malnick S, Lysyy L, Knobler H. Interferon therapy in hepatitis C leading to chronic type 1 diabetes. World J Gastroenterol [Internet]. 2015 Jan 7 [cited 2019 Jan 7];21(1):233. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25574096
Toubiana J, Okada S, Hiller J, Oleastro M, Lagos Gomez M, Aldave Becerra JC, et al. Heterozygous STAT1 gain-of-function mutations underlie an unexpectedly broad clinical phenotype. Blood [Internet]. 2016 Jun 23 [cited 2019 Jan 7];127(25):3154–64. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27114460
Smyth DJ, Cooper JD, Bailey R, Field S, Burren O, Smink LJ, et al. A genome-wide association study of nonsynonymous SNPs identifies a type 1 diabetes locus in the interferon-induced helicase (IFIH1) region. Nat Genet [Internet]. 2006 Jun 14 [cited 2019 Jan 7];38(6):617–9. Available from: http://www.nature.com/articles/ng1800
Rice GI, Del Toro Duany Y, Jenkinson EM, Forte GM, Anderson BH, Ariaudo G, et al. Gain-of-function mutations in IFIH1 cause a spectrum of human disease phenotypes associated with upregulated type I interferon signaling. Nat Genet [Internet]. 2014 May 30 [cited 2019 Jan 7];46(5):503–9. Available from: http://www.nature.com/articles/ng.2933
Nejentsev S, Walker N, Riches D, Egholm M, Todd JA. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science [Internet]. 2009 Apr 17 [cited 2019 Jan 7];324(5925):387–9. Available from: https://doi.org/10.1126/science.1167728
Cunninghame Graham DS, Morris DL, Bhangale TR, Criswell LA, Syvänen A-C, Rönnblom L, et al. Association of NCF2, IKZF1, IRF8, IFIH1, and TYK2 with systemic lupus erythematosus. McCarthy MI, editor. PLoS Genet [Internet]. 2011 Oct 27 [cited 2019 Jan 7];7(10):e1002341. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22046141
Gorman JA, Hundhausen C, Errett JS, Stone AE, Allenspach EJ, Ge Y, et al. The A946T variant of the RNA sensor IFIH1 mediates an interferon program that limits viral infection but increases the risk for autoimmunity Nat Immunol [Internet]. 2017 Jul 29 [cited 2019 Jan 7];18(7):744–52. Available from: https://doi.org/10.1038/ni.3766
Shaw MH, Freeman GJ, Scott MF, Fox BA, Bzik DJ, Belkaid Y, et al. Tyk2 negatively regulates adaptive Th1 immunity by mediating IL-10 signaling and promoting IFN-gamma-dependent IL-10 reactivation. J Immunol [Internet]. 2006 Jun 15 [cited 2019 Jan 7];176(12):7263–71. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16751369
Shimoda K, Kato K, Aoki K, Matsuda T, Miyamoto A, Shibamori M, et al. Tyk2 plays a restricted role in IFN alpha signaling, although it is required for IL-12-mediated T cell function. Immunity [Internet]. 2000 Oct [cited 2019 Jan 7];13(4):561–71. Available from: http://www.ncbi.nlm.nih.gov/pubmed/11070174
Boisson-Dupuis S, Ramirez-Alejo N, Li Z, Patin E, Rao G, Kerner G, et al. Tuberculosis and impaired IL-23-dependent IFN-γ immunity in humans homozygous for a common TYK2 missense variant. Sci Immunol [Internet]. 2018 Dec 21 [cited 2019 Jan 14];3(30):eaau8714. Available from: https://doi.org/10.1126/sciimmunol.aau8714
Dendrou CA, Cortes A, Shipman L, Evans HG, Attfield KE, Jostins L, et al. Resolving TYK2 locus genotype-to-phenotype differences in autoimmunity. Sci Transl Med [Internet]. 2016 Nov 2 [cited 2019 Jan 7];8(363):363ra149. Available from: https://doi.org/10.1126/scitranslmed.aag1974
Gorman JA, Hundhausen C, Kinsman M, Arkatkar T, Allenspach EJ, Clough C, et al. The TYK2-P1104A autoimmune protective variant limits coordinate signals required to generate specialized T cell subsets. Front Immunol. 2019;in press.
Funding
Jane H. Buckner and Karen Cerosaletti are supported by NIH grant 1DP3DK111802. Sarah E. Flanagan has a Sir Henry Dale Fellowship jointly funded by the Wellcome Trust and the Royal Society (grant no. 105636/Z/14/Z).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Pathogenesis of Type 1 Diabetes
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Johnson, M.B., Cerosaletti, K., Flanagan, S.E. et al. Genetic Mechanisms Highlight Shared Pathways for the Pathogenesis of Polygenic Type 1 Diabetes and Monogenic Autoimmune Diabetes. Curr Diab Rep 19, 20 (2019). https://doi.org/10.1007/s11892-019-1141-6
Published:
DOI: https://doi.org/10.1007/s11892-019-1141-6