Abstract
Brain tissue is not only one of the most important but also the most complex and compliant tissue in the human body. While long underestimated, increasing evidence confirms that mechanics plays a critical role in modulating brain function and dysfunction. Computational simulations–based on the field equations of nonlinear continuum mechanics–can provide important insights into the underlying mechanisms of brain injury and disease that go beyond the possibilities of traditional diagnostic tools. Realistic numerical predictions, however, require mechanical models that are capable of capturing the complex and unique characteristics of this ultrasoft, heterogeneous, and active tissue. In recent years, contradictory experimental results have caused confusion and hindered rapid progress. In this review, we carefully assess the challenges associated with brain tissue testing and modeling, and work out the most important characteristics of brain tissue behavior on different length and time scales. Depending on the application of interest, we propose appropriate mechanical modeling approaches that are as complex as necessary but as simple as possible. This comprehensive review will, on the one hand, stimulate the design of new experiments and, on the other hand, guide the selection of appropriate constitutive models for specific applications. Mechanical models that capture the complex behavior of nervous tissues and are accurately calibrated with reliable and comprehensive experimental data are key to performing reliable predictive simulations. Ultimately, mathematical modeling and computational simulations of the brain are useful for both biomedical and clinical communities, and cover a wide range of applications ranging from predicting disease progression and estimating injury risk to planning surgical procedures.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Brain tissue is one of the most complex tissues in the human body. Neurological disorders, including stroke, encephalitis, dementias, and epilepsy, have been identified as one of the major public health concerns by the world health organization. An additional threat are the consequences of neurotrauma with over 2 million people affected by traumatic brain injury each year [111]. While over the past several decades, neuroscience has mostly been limited to electro-physiological, biochemical, molecular and genetic studies [162], more recent studies show that mechanics plays a critical role for neuronal function and dysfunction [9, 68]. External mechanical loads may cause damage of brain tissue during traumatic brain or spinal cord injury [101], but mechanical cues may also control developmental processes and progression of disease through mechanosensation of nerve cells [96, 162]. For instance, mechanics seem to not only drive cortical folding during mammalian brain development [21, 26, 141, 143], but also tumor cell migration and cell apoptosis [162]. Unphysiological changes in the mechanical environment induce cortical malformations [21, 27] associated with schizophrenia [76], autism [129], and epilepsy [14], or cell death in Alzheimer’s patients associated with dementia [15].
Computational modeling and personalized simulations can provide fundamental insights into the underlying mechanisms during injury and disease. Such predictive models not only reduce the necessity of experiments on humans and animals, but also allow for the development of novel treatment strategies or the detailed planning of surgical procedures [171]. However, realistic predictions of mechanobiological processes in the brain require sophisticated mechanical models that capture the complex and unique characteristics of this ultrasoft, highly adaptive and heterogeneous tissue. While great efforts have been made to mechanically model the behavior of brain tissue in health and disease [68], contradictory experimental results have constantly hindered progress [29] and caused confusion and delay [7]. This review aims to explain these discrepancies and provide a comprehensive overview on the mechanics of brain tissue.
After addressing the challenges associated with brain tissue testing and modeling in Sect. 2, we review experimental observations based on different testing techniques to work out the diverse characteristics of brain tissue behavior under different loading conditions in Sect. 3. Subsequently, we present mechanical modeling approaches that are capable of mathematically describing the characteristic tissue behavior in Sect. 4. Finally, we discuss application-specific features to propose appropriate modeling approaches together with the corresponding set of material parameters in Sect. 5. We will close this review with a short conclusion and open challenges in Sect. 6.
2 Challenges
2.1 Brain Tissue is Ultrasoft
With a shear modulus on the order of one kilopascal [22], brain tissue is ultrasoft—softer than any other tissue in the human body. For comparison, Fig. 1 highlights the stiffnesses of various organs in our body. Traditional mechanical test setups have originally been designed for much stiffer materials, even stiffer than bone. The extreme compliance of brain tissue pushes mechanical testing and modeling approaches to their limits. On the one hand, boundary conditions might be inappropriate. On the other hand, force sensors reach their sensitivity limit. As a result, early studies on the stiffness of brain tissue yielded significantly higher values than more recent studies [59]—the testing devices used at that time were merely not capable of recording the accurate tissue responses reliably.
To complicate matters, brain tissue deforms noticeably due to gravity—even just under its own weight. This could have important consequences on testing results, especially when the height of the specimen is relatively large [99]. It is thus difficult to control specimen geometry and local deformation states during biomechanical testing, which, in turn, will affect the forces recorded during testing [138]. This implies that the common assumption of homogeneous deformation during uniaxial tension or compression [22, 121, 137, 140] is debatable [147]. The only alternative to circumvent this concern are inverse studies, where finite element simulations are performed to capture local inhomogeneities in the deformation. However, it will always be difficult to exactly match the specimen geometry of experiment and simulation throughout the entire loading history. In general, ultrasoft tissues undergo large deformations–even under their own weight–and it is essential to formulate mechanical models with the framework of nonlinear continuum mechanics.
2.2 Brain Tissue is Highly Fragile
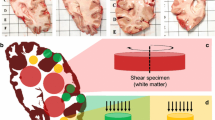
Due to the fragile nature of brain tissue, it is difficult to control tissue damage during biomechanical testing, especially when applying large deformations. In white matter brain tissue, axons, the long and slender nerve fibers, will tear when loaded above a certain threshold, as illustrated in Fig. 2 of [58]. Previously, this threshold has been determined at approximately 18% tensile strain. Measurements above, or even close to, this damage threshold are likely to render meaningless results.
Damage evolution and fracture process in a (14 mm/9.5 mm initial height/edge) prismatic specimen of white matter harvested from the occipital lobe in the frontal direction. Adapted from [58]
In addition, the effect of adhesion, as illustrated in Fig. 3, enhances the risk of damaging specimens during preparation and handling [61]. This demonstrates that biomechanical tests need to be well-designed and performed with exceptional caution.
2.3 Brain Tissue is Biphasic
Unlike other soft tissues, the brain is not only ultrasoft, but also has an exceptionally high water content, 0.83 g/ml in gray matter and 0.71 g/ml in white matter [180]. From a total of about 80% water, approximately 20–40% is free flowing cerebrospinal fluid, while the rest resides inside the cells. Due to the biphasic nature of brain tissue, it is difficult to control drainage conditions during biomechanical testing [58]. As a result, depending on the testing setup, the incompressible fluid is trapped within the tissue or free to escape, which will largely affect the recorded response. This interaction between solid and fluid phases is highly sensitive to the time and length scales [29]. Depending on the type of loading and the testing setup, e.g., unconfined compression/extension versus nanoindentation experiments, different tissue components control the recorded mechanical stiffness. Nanoindentation experiments on intact brain tissue slices [20] closely mimic the nature of confined compression tests, where the fluid phase is mostly kept within the solid matrix and contributes to the tissue stiffness. In contrast, during unconfined compression and tension tests, the fluid is free to escape, and we primarily probe the elastic properties of the solid skeleton consisting of cells, intercellular connections, and extracellular matrix. Naturally, these tests introduce an additional time scale, the time for the fluid to escape the solid. Therefore, the results are sensitive to the type of preconditioning and the recovery time between two consecutive tests [23].
2.4 Brain Tissue is Highly Heterogeneous
Brain tissue displays a high degree of microstructural heterogeneity, as shown in Fig. 4. Its cellular and extracellular structure conforms with regionally varying functional demands, and so do its mechanical properties [173]. For most practical purposes, it seems too simplistic to consider brain tissue as a single homogeneous material.
Microstructure of different brain regions, the cortex (C), the basal ganglia (BG), the corona radiata (CR), and the corpus callosum (CC). Klüver-Barrera (K.B.) staining was used to color myelin wrapped around nerve fibers in blue, neuropil in pink, and nerve cells in purple. Images are shown at 20 magnification. Adapted from [22]
Cellular organization of brain tissue. In white matter (corona radiata), myelinated axons allow for rapid nerve impulse conduction; intermediate oligodendrocytes connect and form several myelin sheaths. Fibrous astrocytes ensure supply of nutrients and synaptic processing. In gray matter (cortex), neurons form synapses with each other and with protoplasmic astrocytes. In both white and gray matter, microglial cells contribute to clearance of debris and synapse remodeling. Reprinted from [27] under Creative Commons Attribution License (CC BY)
Figure 5 illustrates the major cell types in white and gray matter tissue. Gray matter regions contain mainly neuronal cell bodies, protoplasmic astrocytes providing neurons with nutrients, and microglia as active immune defense. White matter regions, on the contrary, contain axons, oligodendrocytes which wrap isolating myelin sheath around the axons, fibrous astrocytes, and microglia [27]. Notably, not only the cellular composition may be relevant for macroscopic tissue mechanics but also extracellular matrix components. The latter may show regional trends that even differ from those of brain cells.
Notably, the microstructure of brain tissue not only varies in space but also in time. When we learn a new task, for example juggling, the neurons responsible for fine motor skills will form further connections with one another and create new synapses. Brain plasticity, the temporal changes in the tissue’s microstructure, also affects the mechanical response. This emphasizes the complexity of brain tissue behavior and the high demands when aiming to develop mechanical models for brain tissue.
2.5 Brain Tissue is Not Easily Available
One major issue considering the mechanical testing of brain tissue is that it is exceptionally challenging to obtain adequate material. While it would be desirable to test the mechanical properties of brain tissue in vivo—in its natural environment—there are major issues with in vivo testing [97]. Firstly, traditional mechanical testing setups, initially developed and used for non-organic materials, require to prepare specimens of a certain geometry—optimized to allow for homogeneous deformation states during testing [63]. This is obviously not possible without removing the material from its natural environment and destroying its biological functionality. Secondly, the calibration of mechanical models that are useful to study injury and disease require a versatile set of different loading conditions, including deformations that go beyond physiological conditions and might induce tissue damage. Again, those tests can not be performed in vivo on a healthy individual.
On the other hand, the deformations imposed by common ex vivo testing techniques such as nanoindentation, uniaxial compression and extension, or simple shear will undoubtedly differ from those experienced in vivo. Recently, a newly established testing technique, magnetic resonance elastography, has enabled to study the mechanical properties of human brain tissue in vivo [112]. While magnetic resonance elastography is a powerful tool to perform non-invasive longitudinal studies to monitor the stiffness evolution under diseased conditions, for example in multiple sclerosis [182], it can not yet be immediately used to derive mechanical models for brain tissue, and fully replace ex vivo experiments [12]. Magnetic resonance elastography is based on the assumption of linear elasticity or viscoelasticity [146], and does not offer enough information to establish nonlinear, finite strain constitutive models for realistic computational simulations. Besides, the mechanical properties obtained from magnetic resonance elastography are sensitive to positioning, to the method to extract elastic parameters, and to the excitation frequency [177]. It is not at all trivial to accurately quantify regional mechanical properties, especially considering thin structures such as the cortex and the corpus callosum [131]. In summary, the better we understand the different “shades” of brain tissue behavior, the easier it will be to establish new appropriate testing techniques in the future.
3 Experimental Observations
Mechanical testing techniques and their range of application. AFM atomic force microscopy, IND indentation, MMT multiaxial mechanical testing, OST oscillatory shear testing, MRE magnetic resonance elastography, NDG neurodegeneration, NRG neuroregeneration, DAI diffuse axonal injury, TG tumor growth, NPH normal pressure hydrocephalus, NDV neurodevelopment, NS neurosurgery, TBI traumatic brain injury
From all these challenges, it becomes clear why experimental studies on the mechanical properties of brain tissue have been equivocal and why stiffness values in the literature have varied by an order of magnitude or more [119]. Depending on the testing techniques different aspects of the complex mechanics of brain tissue control the recorded response as illustrated in Fig. 6. In this section, we summarize the experimentally observed characteristics of brain tissue, discuss how different mechanical testing techniques may have resulted in contradictory findings, and critically evaluate facts that are still under debate. Ultimately, this will allow us to work out the key characteristics to establish appropriate constitutive models for brain tissue in Sect. 4.
3.1 Brain Tissue Stiffness Increases with Increasing Strain
Figure 7 illustrates the loading-mode-specific elastic behavior of brain tissue—the response we expect for very slow processes such as tissue growth or progress of neurodegenerative diseases—here tested during unconfined compression, tension, and simple shear experiments for four different brain regions, the cortex, the basal ganglia, the corona radiata, and the corpus callosum as introduced in Fig. 4 [22]. Independent of the loading mode, we observe a strain-stiffening behavior—the stiffness increases with increasing strain—which highlights the strong nonlinearity of the tissue response.
Average elastic response of brain tissue with standard deviations during multiple loading modes, simple shear, compression, and tension, for different brain regions: cortex, basal ganglia, corona radiata, and corpus callosum. Adapted from [22]
3.2 Brain Tissue is Stiffer in Compression Than in Tension
Figure 7 reveals that stresses are significantly higher in compression than in tension. This important characteristic of brain tissue has been consistently observed by several research groups [22, 58, 85, 120, 121]. A convenient test to capture the loading-mode-specificity of the mechanical response of brain tissue is to perform tests under combined loading conditions, as illustrated in Fig. 8 [22, 133]: For combined compression/tension-shear experiments, the shear stresses increase noticeably with increasing compression, but only marginally with increasing tension [22]. We may attribute this behavior to the biphasic nature of brain tissue discussed in Sect. 2.3: Cerebrospinal fluid trapped inside the solid network of cells and extracellular matrix offers noticeably resistance in compression but only marginal resistance to tension.
Combined compression/tension-shear loading: average elastic shear stress versus amount of shear for sinusoidal simple shear superimposed on axial stretch \(\lambda =1.0\), 0.95, 0.9, 0.85, 0.8, 0.75, 1.05, 1.1, 1.15, 1.2, and 1.25. Shear stresses increase with increasing compressive strain but not with increasing tensile strain. Adapted from [22]
3.3 Brain Tissue is Stiffer During Loading Than During Unloading
Figure 9 demonstrates that even for a relatively slow loading speed of 2mm/min corresponding to strain rates of approximately 0.0067 1/s, brain tissue exhibits a highly hysteretic response [22]. We expect this hysteresis to vanish for much slower rates, which might occur in vivo during brain growth or disease, but are difficult to reproduce during ex vivo testing.
Average unconditioned response of brain tissue with standard deviations during multiple loading modes, simple shear, compression, and tension, for different brain regions: cortex, basal ganglia, corona radiata, and corpus callosum. Adapted from [22]
Figure 10 shows stress relaxation experiments for all four brain regions [22], which confirm the extreme time-dependence of brain tissue with a stress relaxation of up to 80% within only 300 s. Interestingly, white matter tissue, with a stress relaxation of more than 70% after 300 s, is more viscous and responds more slowly than gray matter, with a stress relaxation of approximately 65%. Within white matter, specimens from the corpus callosum relax faster than specimens from the corona radiata. Within gray matter, the basal ganglia and the cortex exhibit a similar relaxation behavior. Stress relaxation percentages are slightly higher in shear than in compression, but both loading modes show similar regional dependencies.
Stress relaxation behavior of brain tissue with standard deviations and stress relaxation percentages after 5 minutes for simple shear and compression loadings in different brain regions: cortex, basal ganglia, corona radiata, and corpus callosum. Adapted from [22]
3.4 Brain Stiffness Increases with Increasing Strain Rate
Figures 11 and 12 illustrate that brain tissue not only stiffens with increasing strain but also with increasing strain rate. Rate-dependence of brain stiffness has consistently been reported in the literature using different testing techniques: shear testing or oscillatory shear testing [44, 127, 151], uniaxial compression or tension [57, 83, 85, 132, 137, 140], indentation [20, 107, 136, 163], and magnetic resonance elastography [36, 145]. Figure 11 illustrates uniaxial compression, tension, and simple shear experiments performed at strain rates of 0.33 and 0.0067 1/s, respectively. Figure 12 shows nanoindentation experiments over a loading rate spectrum from 1 to \(160\,\mu \hbox {m/s}\). Within the analyzed loading rate regime both gray and white matter double their maximum forces and corresponding moduli when increasing the loading rate by two orders of magnitude [20]. This effect becomes particularly important for applications such as blunt or traumatic brain injury, where even higher strain rates occur [137, 158].
Average initial loading segment with standard deviations for simple shear as well as uniaxial compression experiments loaded at different speeds of \(v=100\) mm/min and \(v=2\) mm/min corresponding to strain rates of approximately 0.33 and 0.0067 1/s, respectively, in four different brain regions: cortex, basal ganglia, corona radiata, and corpus callosum
Our results in Fig. 11 suggest that the effect of strain rate is more pronounced in white matter regions than in gray matter regions. This observation agrees well with the stress relaxation experiments in Fig. 10, which show that white matter relaxes faster than gray matter. A possible explanation for this behavior could be the difference in the permeability of gray and white matter. While for slower loadings, fluid has more time to escape, for faster loadings, it offers resistance, which leads to higher stresses. As white matter specimens seem to loose a higher amount of fluid during unconfined experiments than gray matter specimens [22, 23, 25], their strain rate effect is more pronounced.
Loading rate sensitivity of gray and white matter. Sensitivity of indentation force versus indentation depth for varying loading rates reveals the rate-dependent nature of brain tissue. Indentation force and modulus increase with increasing loading rate. Adapted from [20]
3.5 Brain Tissue Softens upon Preconditioning
Average conditioned response of brain tissue with standard deviations during multiple loading modes, simple shear, compression, and tension, for different brain regions: cortex, basal ganglia, corona radiata, and corpus callosum. Adapted from [22]
Figure 13 shows the conditioned tissue response during the third cycle of cyclic loading in simple shear, uniaxial compression, and uniaxial tension for four different brain regions, the cortex, the basal ganglia, the corona radiata, and the corpus callosum. Notably, during the third loading cycle, the stresses are significantly lower than during the first loading cycle, the unconditioned tissue response in Fig. 9. We conclude that brain tissue softens upon preconditioning. This observation agrees well with reported results, where the equilibrium shear modulus during the fifth cycle was consistently about 20% lower than during the first cycle [62].
Representative stress versus amount of shear/stretch behavior in simple shear, compression, and tension for stepwise increasing shear/stretch—three cycles per shear/stretch level. The tissue exhibits substantial pre-conditioning during the first cycle of each level, and successively softens with increasing maximum shear/stretch
Figure 14 illustrates that a characteristic softening between the first loading cycle and all subsequent cycles occurs whenever the tissue is loaded past the previous maximum strain. We once more attribute this characteristic conditioning to fluid leaking out of the sample: As the maximum strain increases, more fluid escapes the sample and no longer offers resistance to loading. In turn, the offset at zero strain upon unloading increases.
For simple shear and tension, we observe jagged peaks during the first loading cycle of each strain level. We suggest two possible explanations for this phenomenon according to the challenges introduced in Sects. 2.1–2.4. Firstly, it is possible that part of the water molecules of the cerebrospinal fluid are non-covalently bonded to the highly charged macromolecules in the extracellular matrix; during initial loading, those weak interactions break and the water is free to escape. Secondly, these peaks could result from tissue that initially adheres to the upper or lower specimen holder, without being glued, but comes loose during loading.
3.6 Brain Tissue Recovers from Preconditioning
Representative nominal stress versus stretch behavior of a specimen from the corona radiata loaded twice with a set of three cycles up to 10% compressive strain separated by a 60 min recovery period. The tissue exhibited a similar pre-conditioning pattern for both compression tests. Adapted from [22]
Figure 15 demonstrates that the characteristic conditioning behavior discussed in Sect. 3.5 is fully recoverable. When we load the same specimen after a recovery period of one hour, the cyclic response of brain tissue up to 10% compression follows almost the identical path of the initial loading—again showing the evident softening after the first loading cycle. A similar behavior has been observed during nanoindentation experiments: The tissue continuously softens throughout multiple indents, but fully recovers within a few hours of rest [20]. We conclude that the softening due to conditioning is not associated with tissue damage but with visco- or poro-elastic effects that recede over time. This is an important finding towards developing appropriate mechanical models for brain tissue behavior in Sect. 4. Notably, the tissue only recovers when it is kept hydrated. If the tissue was left in air, it would dry out and the response would stiffen over time [44].
3.7 Brain Tissue is not Notably Anisotropic
From a mechanics point of view, a fundamental question that needs to be answered is whether brain tissue is an isotropic or anisotropic material. Due to controversies regarding previous results in the literature, where some suggested that brain tissue was isotropic [22, 22, 91, 120, 137], but others showed that there were significant directional trends [52, 53, 85, 134, 165], we combined biomechanical testing of the same specimen in three orthogonal loading directions with antecedent diffusion tensor imaging to carefully analyze to which extent the fibrous microstructure of axonal networks in white matter results in an anisotropic macroscopic mechanical response [22]. This allows us to minimize falsifying effects of non-uniform fiber distributions or inter-specimen variations that might have affected previous studies. We determined nerve fiber distributions prior to specimen extraction and tested the same microstructurally anisotropic sample in multiple loading directions—the three possible modes for simple shear (FT, TT, and TF) and the two possible modes for compression and tension (FF and TT), as denoted in Fig. 16a.
While diffusion tensor images showed highly anisotropic diffusion properties, mechanical testing revealed no significant directional dependencies, as illustrated in Fig. 16. According to these results, while structurally anisotropic, brain tissue can be considered as an isotropic material from a mechanical point of view. This finding seems reasonable considering the fact that nerve fibers are primarily functional elements of our brain that only marginally contribute to its mechanical strength. We note, however, that mean stresses along fibers appeared slightly higher than perpendicular to the fibers under tensile loading and slightly lower under compressive loading. This could imply that—although statistically insignificant—nerve fibers contribute to tissue strength in tension but not in compression.
a Two possible compression and tension modes, along (FF) and transverse (TT) to the uniaxial nerve fiber direction \(\varvec{f}_0\) in the corpus callosum (CC); b three simple shear modes IJ, for I,J \(\in\) {T,F}, where I denotes the normal vector of the face that is shifted by simple shear and J denotes the direction of shear, along (F) or transverse (T) to the fiber direction \(\varvec{f}_0\). c Direction-dependent behavior of brain tissue. Column plots of shear moduli (mean ± SD) averaged over three possible modes in simple shear (FT, TT, and TF), and two possible modes in compression and tension (FF and TT) for all specimens from the corpus callosum. The data indicate a marginally softer response along nerve fibers in compression and marginally stiffer response along fibers in tension but the differences appeared to be statistically not significant (n.s.). Reprinted with permission from [22]
Our results agree well with an early study on the rheological shear response of human brain tissue [151], which showed higher directional variation in gray matter than in white matter, but neither of the differences appeared significant. Contradictory to our findings, studies on porcine brain tissue found a significantly stiffer shear response orthogonal to nerve fibers than along fibers in the corpus callosum [134]. In the corona radiata, however, the trend was opposite. The authors of this study sheared each specimen in two orthogonal directions similar to our experiments, but only estimated fiber orientations from anatomical knowledge and used rectangular specimen dimensions of \(10\times 5\times 1\) mm\(^3\). Notably, the measured shear stresses were consistently higher in the direction of the longer axis corresponding to the direction orthogonal to fibers in the corpus callosum, and to the fiber direction in the corona radiata. This could indicate that directional dependencies are an artifact of the non-cuboidal specimen dimensions rather than a result of the anisotropic distribution of nerve fibers, which could explain the contradictory results. Interestingly, yet other studies on the porcine corpus callosum found opposite trends with a significantly stiffer response in the fiber direction than perpendicular to it in dynamic shear [53] and tensile tests [165]. In both studies, specimens were relatively large with dimensions of up to \(16\times 16\times 3\,\hbox {mm}^3\) and \(5\times 5\times 60\,\hbox {mm}^3\). Our diffusion tensor images showed that even in the much larger human brain, it would be challenging to extract specimens of that size that exhibit a sufficiently uniform microstructure. We therefore interpret the corresponding results with a degree of caution.
Overall, compared to other soft biological tissues including muscle, tendon, or arteries with pronounced directional stiffness differences of an order of magnitude or more, the directional stiffness of brain tissue only varies by a few percent and can reasonably well be assumed as isotropic. Nonetheless, while isotropy seems to be a valid assumption for the elasticity of brain tissue, diffusion or permeability properties might still be anisotropic. Furthermore, the role of axons could be decisive for white matter brain damage, where rupture of axons can lead to loss of brain function [37, 66].
3.8 Brain Tissue Stiffness is Region-Dependent
Due to its high microstructural heterogeneity, brain tissue can hardly be considered as a single material with unified material properties. While early studies on the mechanical properties of the brain focused on brain tissue as a whole, more recent experimental studies have distinguished between different regions, i.e., white and gray matter [20, 57, 114, 163], or, even more refined, cortex, basal ganglia, corona radiata and corpus callosum, as depicted in Fig. 4 [22, 85]. Others tested the individual properties of the cerebrum, cerebellum, pons, and medulla [109].
Regional variation of gray and white matter moduli. Measurements at three different slices and three different locations reveal that the specimen moduli vary markedly across the brain. Gray matter, left, is softer than white matter, right, and displays smaller regional variations. Black horizontal lines indicate the mean; gray zones indicate the standard deviation. Adapted from [20]
Astonishingly, experimental results towards regional trends in brain tissue properties have been contradictory. While indentation experiments including those shown in Fig. 17 yield higher stiffnesses in white matter from the corona radiata than in the cortex [20, 92, 163], simple shear, unconfined compression and tension tests in Figs. 7, 8, 9, 10 and 11 show the opposite trend [22]. The origin of these, at first sight, contradictory results lies in the complexity of brain tissue behavior, which highly depends on the length and time scales of the applied loading. Furthermore, not only the tissue’s elasticity shows local variations but also other properties such as viscosity and porosity. To disclose regional differences in brain tissue properties, it is therefore indispensable to thoroughly understand the different “shades” of brain tissue mechanics and their loading-mode specific effects.
3.8.1 Regional Trends Depend on the Loading Rate
Firstly, the rheological difference discussed in Sect. 3.4 leads to a rate-dependency of regional trends: White matter stiffens relative to gray matter with increasing loading rate. As a result, shear, compression or tension experiments in the fast loading regime associated with phenomena on the order of seconds or milliseconds such as traumatic brain injury have reported a higher stiffness for white matter from the corona radiata than for cortical gray matter [85, 114], while experiments in the slow loading regime, as those shown in Fig. 11, indicate the reversed relationship. The strain-rate dependence of regional trends is also supported by a recent study showing that inter-regional mechanical properties become increasingly heterogeneous with increasing strain rate [108].
3.8.2 Regional Trends Depend on Drainage Conditions
To explain differences between nanoindentation and unconfined macroscopic experiments—both performed at relatively low rates—we refer to the biphasic nature of brain tissue discussed in Sect. 2.3. Nanoindentation of intact tissue slices can be regarded as a virtually confined test, where the free flowing fluid is mostly kept within the solid matrix. During simple shear, unconfined compression and tension tests, in contrast, the fluid is free to escape. As a result, unconfined tests reveal a markedly stiffer response for the unconditioned response, see Fig. 9, than for conditioned response, see Fig. 13, which emphasizes that the fluid plays a significant role in the tissue’s mechanical strength: Brain tissue softens when part of the fluid escapes. In turn, when tissue samples have time to saturate themselves again after loading during a recovery period of one hour, in Fig. 15, they regain their initial stiffness.
When comparing the porous nature of different brain regions, our experiments indicate that the largest amount of fluid escapes from white matter specimens of the corpus callosum, closely followed by the corona radiata, while gray matter specimens from the cortex loose the least amount of fluid [23]. This difference in the permeability of gray and white matter is also reflected in larger hysteresis areas in white matter than in gray matter during cyclic loading [22]. We can rationalize these observations with the underlying tissue microstructure: the corpus callosum consists of a sparsely cross-linked network of unidirectional fibers, whereas the cortex consists of a densely connected network of dendrites that traps the fluid phase inside the tissue.
With regard to these observations, we suppose that during unconfined experiments, we primarily probe the “elastic” properties of the solid skeleton, the cells and the extracellular matrix. Those are stiffer in gray matter from the cortex than in white matter from the corona radiata, see Fig. 9 [22]. We observe a different behavior for confined compression experiments during nanoindentation with relatively large indenters, as shown in Fig. 18: The fluid cannot escape freely and we probe both solid and fluid phases. Due to the differences in the porous nature of gray and white matter, we record larger stiffnesses for white matter from the corona radiata than for cortical gray matter, see Fig. 17.
Experimental nanoindentation setup. A 5 mm-thick coronal slice of freshly harvested brain tissue is placed in a 100mm-diameter petri dish and mounted underneath the force transducer of the TriboIndenter\(^{\mathrm{TM}}\). A 12 mm-diameter washer marks the indentation region and stabilizes the sample. A circular flat punch of 1.5 mm diameter ensures a homogenized specimen response. Reprinted with permission from [20]
In summary, because of its ultrasoft nature, brain tissue stiffness recordings are highly sensitive to the fluid content of the sample. Undrained samples are stiffer than drained samples, and drainage rates depend critically on the tissue microstructure. These effects are less pronounced in other types of tissues with a lower fluid volume fraction. This explains why the reported stiffness values of brain tissue vary hugely. Without an explicit mention of loading rates, drainage conditions, and sample size and geometry, it is virtually impossible to compare stiffness values recorded under different test conditions. The concept of a single one gray or white matter stiffness value simply does not exist for brain tissue, and it is critical for computational simulations to understand exactly which situation applies to select the appropriate model and parameter values.
3.8.3 Regional Trends Depend on the Length Scale
Figure 6 summarizes various testing techniques to probe the mechanical behavior of brain tissue at different spatial and temporal scales. A prominent method to probe brain tissue at small spatial scales is atomic force microscopy [35]. Unlike nanoindentation on the meso-scale with relatively large indenter tips shown in Fig. 18, atomic force microscopy has resulted in yet higher stiffness in gray than in white matter [94]. In atomic force microscopy, the size of the indenter tip is on the order of the dimensions of individual cells. The indenter seems to be small enough not to trap the fluid beneath the tip which suggests that these tests probe the solid component of brain tissue similar to unconfined experiments at slow loading rates.
A prominent method to probe brain tissue at large spatial scales, embedded in the skull, is magnetic resonance elastography [97]. Its major advantage is that it allows us to probe the brain tissue in vivo in its natural environment. Here, we most likely measure additional effects due to the interaction of brain tissue with surrounding structures including the meninges and the skull. Because of its limited spatial resolution, it remains questionable whether in vivo magnetic resonance elastography can accurately quantify regional mechanical properties, especially in thin structures such as the cortex and the corpus callosum [131]. Furthermore, the stiffnesses from magnetic resonance elastography are highly sensitive to the actuation frequency [177]. Not surprisingly, some studies found cortical gray matter to be stiffer than white matter [71], while others reported the opposite trend [114], or no significant differences [10].
3.8.4 Conclusion Towards Regional Trends
In summary, while it seems important to take the significant local variations in tissue properties into account when mechanically modeling brain tissue, we have to pay caution with regard to the specific application we have in mind. There are no right or wrong testing results; depending on the length and time scales during testing, as well as the boundary conditions, we observe different trends. For instance, depending on the application of interest, confined or unconfined testing data could be relevant. The former represent the behavior of brain tissue at intermediate and short time scales during surgery [171] or impact loading [38], while the latter might rather represent the behavior for slow processes during brain development [26, 143, 157] or tumor growth [90].
In addition to the different regions within the cerebrum, we can distinguish between the cerebrum, cerebellum, corpus callosum, thalamus, and brain stem, amongst others [176]. Indentation experiments, for instance, show that the mouse cerebellum is softer than the mouse cerebrum [108]. This agrees with results on human brain tissue using magnetic resonance elastography [115, 123, 176]. Finally, it is also important to note that, even within the brain regions we have introduced in Fig. 4, tissue properties may vary significantly. Indentation experiments, for instance, revealed noticeable inter-regional variations within the corona radiata of porcine brain tissue [31] and studies suggest that stiffness variations in white matter tissue are linearly correlated to the local myelin content [173]. Understanding the effects of the tissue microstructure on the macroscopic mechanical response is critical for the interpretation of the constitutive behavior of the human brain for computational simulations.
3.9 Open Questions
There is a general agreement that the ultrasoft nature, the high fluid content, and the biochemical composition make brain tissue very different from all other soft tissues in our body. This implies that factors that have traditionally been considered irrelevant in soft tissue mechanics could play an important role when characterizing the material properties of the brain. In this section, we highlight several open questions that could point towards new studies with a view to create a more holistic picture of the behavior of our brain under healthy and diseased conditions.
3.9.1 Is Brain Stiffness Species-Dependent?
Due to limited availability and ethical considerations, only a few studies have actually tested human brain tissue [30, 44, 50, 51, 58, 59, 85, 134, 151]. Alternatively, researchers consulted porcine [120, 121, 134, 137, 139, 140, 160, 165] or bovine brain tissue [13, 20, 40, 163] due to their structural similarities with the human brain. Others tested the properties of rat [35, 49, 54] or mouse brains [133]. Since the primary goal of developing and calibrating mechanical models for the brain is to assist diagnosis and treatment of human patients, it is important to understand species-dependent peculiarities. Early compression stress relaxation experiments suggested that monkey brain tissue was stiffer than human brain tissue [59]. More recent indentation experiments show that mouse brain tissue is stiffer than rat brain tissue, which is again stiffer than porcine brain tissue [109]. Interestingly, these observations imply a negative correlation between stiffness and total brain volume: the smaller the brain, the stiffer the response. This hypothesis, however, contradicts shear relaxation experiments in which human brain tissue was stiffer than porcine brain tissue [134]. However, in this study, the specimen thickness of only 1mm might have affected tissue integrity and the obtained results. To date, there is no general agreement on the species-dependence of brain tissue properties and it remains unclear whether the observed differences are an artifact of the testing method or the result of a true size effect that we can observe in microstructural engineering materials.
3.9.2 Is Brain Stiffness Correlated with Cell Density?
One approach to understand and predict regional variations in brain tissue properties discussed in Sect. 3.8 is to disclose the correlation between macroscopic mechanics and the locally varying microstructure. First steps towards this direction have only been taken recently. Due to the functionality of nervous tissue, most attention has been paid to cellular components, while extracellular matrix components were given less consideration. The composition of different cell types such as neurons, astrocytes, oligodendrocytes, or microglia, and even their local morphological changes in response to functional demands is illustrated in Fig. 5. Independent of those differences, different brain regions, and cell decomposition, however, we find a negative correlation between the shear modulus and the total number of cell nuclei, as illustrated in Fig. 19.
Correlation between shear moduli and the number of cell nuclei in different brain regions: cortex (C), basal ganglia (BG), corona radiata (CR), and corpus callosum (CC), exemplary shown for the unconditioned response. Adapted from [24]
These preliminary results agree well with a recent study on live mouse brain tissue [5] and indicate that the cells might actually be the softest component of brain tissue. Accordingly, extracellular matrix components could play an important role. In this context, it is interesting to note that the cells in the central nervous system have been shown to be very soft compared to cells from other tissues [106]—as is the overall tissue response. This conjecture is supported by another recent study showing that the stiffness of brain tissue can not be solely determined by the stiffness of the cells that constitute the tissue [81]. But, it contradicts a previous study on spinal cord tissue, where the stiffness positively correlated with the relative tissue area covered by cell nuclei [94, 95]. The latter finding could be attributed to the fact that those measurements were performed using atomic force microscopy indentation on a smaller length scale than the experiments which are the basis for Fig. 19 [22]. On the resolution of individual cells, it was further shown that glial cells, including astrocytes, oligodendrocytes, and microglia, are even softer than neurons [106]. Importantly, however, the stiffness measured for individual cortical cells changed depending on the extracellular matrix material used for coating the dish during experimentation [81]. This emphasizes that when aiming to understand tissue mechanics on the continuum scale, it is insufficient to test the stiffness of each component individually. Rather, the overall tissue response and its correlation with the underlying microstructure needs to be characterized, when cells are embedded in their natural environment.
3.9.3 Is Brain Stiffness Correlated with Myelin Content?
Neurons in the central nervous system are surrounded and cross-linked by myelin, a fatty white substance that wraps around axons to create an electrically insulating layer. Fig. 5 illustrates the microstructural implications of myelin in white matter tissue. While the electrical function of myelin is widely recognized, its mechanical importance remains underestimated.
Stiffness-myelin relation in different regions of cerebral white matter. Across n=11 samples, the stiffness increased with increasing myelin content with a Pearson correlation coefficient of 0.92. Dots indicate mean, ellipses indicate standard deviations in stiffness and myelin content. Reprinted with permission from [172]
Figure 20 suggests that white matter stiffness is positively correlated with the local myelin content. These results were obtained by combining nanoindentation testing and histological staining in immature, pre-natal brains [172] and mature, post-natal brains [173] and agree well with uniaxial tension experiments on chick embryo spinal cord tissue, which suggest that myelin and cellular coupling of axons via the glial matrix in large part dictate the tensile response of the tissue [150]. The positive correlation between myelination and stiffness is also confirmed by magnetic resonance elastography studies showing that demyelination reduces the stiffness in a murine model of multiple sclerosis [149]. Interestingly, those processes were shown to be reversible after remyelination. We may conclude that myelin is not only important to ensure smooth electrical signal propagation in neurons, but also to protect neurons against physical forces and provide a strong microstructural network that stiffens the white matter tissue as a whole. The strong correlation between the white matter stiffness and the local myelin content points towards the potential of tissue stiffness as a biomarker for multiple sclerosis and other forms of demyelinating disorders.
3.9.4 Is Brain Stiffness Correlated with DTI Properties?
Microstructural parameters that require histological staining, e.g. the number of cell nuclei and the myelin content, can only be reliably recorded ex vivo. However, it is also desirable to find correlations between mechanical tissue properties and structural data that can be obtained in vivo. One such parameter is the fractional anisotropy (FA) obtained from magnetic resonance imaging and diffusion tensor imaging (DTI). In white matter, fractional anisotropy illustrates the alignment of nerve fibre bundles. In cortical gray matter, it may be interpreted in terms of the orientation and distribution of axonal, dendritic, and glial cell processes [17]. Interestingly, low fractional anisotropy values have recently been attributed to high neuropil complexity [169], which strengthens the hypothesis that brain tissue stiffness closely correlates with interconnections and capillary density [22].
Correlation between the shear modulus \(\mu\) under compression and the fractional anisotropy (FA) from diffusion tensor magnetic resonance imaging. There is a significant decrease of the shear modulus with increasing fractional anisotropy, \(\mu = 1.84-2.17\) FA, \(r= -0.65\) and \(p < 0.001\). Reprinted with permission from [22]
Figure 21 seems to suggest that the value of fractional anisotropy negatively correlates with the shear modulus, \(\mu = 1.84-2.17\) FA, \(r = -0.65\), \(p < 0.001\), during compression loading [22]. Similar correlations can be observed for shear, \(\mu = 1.18-1.34\) FA, \(r = -0.65\), \(p < 0.001\), and tension, \(\mu = 1.3\)-1.55 FA \(r = -0.69\), \(p < 0.001\), and for all loading modes combined, \(\mu = 1.57-1.96\) FA, \(r = -0.66\), \(p < 0.001\)) [22]. Importantly, the observed correlation between tissue stiffness and fractional anisotropy could point towards new methods to access regional variations in tissue properties in vivo using magnetic resonance diffusion tensor imaging.
3.9.5 Does Our Brain Stiffen During Development?
Closely linked to the observations in the previous subsections, experimental studies have consistently shown that brain tissue stiffens during development. Our indentation experiments revealed that both bovine gray and white matter tissue stiffened significantly upon maturation: the gray matter stiffness doubled from \(0.31\pm 0.20\) kPa pre-natally to \(0.68\pm 0.20\hbox { kPa}\) post-natally; the white matter stiffness tripled from \(0.45\pm 0.18\hbox { kPa}\) pre-natally to \(1.33\pm 0.64\hbox {kPa}\) post-natally [172]. This is in perfect agreement with a significant increase in the indentation moduli of rat [49, 152] and mouse [109] brain tissue beginning at 10-12 days after birth and continuing to 180 days. Interestingly, in exactly this period, myelin basic protein as a measure of the progress of myelination increases, which confirms the close correlation between developmental brain stiffening and myelination discussed in the Sect. 3.9.3. Similarly, dynamic shear experiments on 2-3 day old pig brains yielded significantly lower shear moduli than experiments on adult pig brain samples [160]. Interestingly, also the strain-stiffenig character of the tissue samples increased with maturation [160]. According to magnetic resonance elastography measurements, the adult human brain appears to be three to four times stiffer than the brain of young children [30]. Even adolescent brains still show a softer response than adult brains in certain brain regions including the cerebellum as well as the parietal and temporal lobes [115]. Only one group reported the opposite trend with a decrease in tissue stiffness with age based on indentation and shear experiments on rat brains [62, 134].
3.9.6 Does Our Brain Soften with Age?
While it seems well established that brain tissue stiffens during development, the natural next question is whether brain tissue starts to degrade and soften again after it has passed a zenith. Interestingly, neither oscillatory shear tests [51] nor macroscale unconfined experiments [22, 85] showed strong age-dependent trends of brain tissue stiffness, as illustrated in Fig. 22. The graphs summarize ex vivo data from ten human brains and indicate that regional trends, as discussed in Sect. 3.8, are markedly stronger than age- or inter-subject-dependent effects—specimens from a specific region yielded moduli in a similar range independent of age or subject [22]. In contrast to these findings, in vivo measurements using magnetic resonance elastography on human subjects indeed yielded a linear decline in whole-brain elasticity within an age range from 18 to 72 years [77, 146]. We attribute this observation to changes in the fluid balance of the human brain and a decrease in total brain volume, which would affect magnetic resonance elastography but not ex vivo measurements on small tissue samples that primarily probe the elastic properties of the solid phase. This hypothesis is supported by the fact that the relative viscous-to-elastic behavior during magnetic resonance elastography did not differ between age groups, suggesting a preservation of the organization of the tissue’s microstructure, which is responsible for elastic tissue properties [77].
Age-dependent shear modulus in different brain regions: cortex (C), basal ganglia (BG), corpus callosum (CC), and corona radiata (CR), and for different loading conditions, simple shear, compression, and tension. Interestingly, independent of age or subject, specimens extracted from the same brain region yield a shear modulus in the same range. All three ex vivo tests display no significant age-dependency
3.9.7 Does Our Brain Stiffness Change During Disease?
Neurodegenerative diseases involve remodeling of the brain’s microstructure. Expectedly, this also leads to changes in the mechanical properties of the tissue. Most insightful in this respect are studies that compare healthy and diseased brain tissue properties in vivo. A limitation of in vivo measurements via magnetic resonance elastography, as already discussed in Sect. 3.9.6, is that they do not necessarily reflect changes in the local stiffness of the solid phase, including cells and extracellular matrix, but rather changes in the overall integrity of brain tissue, including changes in fluid transport and wave propagation. Magnetic resonance elastography shows that brain tissue softens in multiple sclerosis [156, 182], Alzheimer’s disease [65, 124], and demyelination in general [149]. Interestingly, these softening effects scale with disease stage [86]. This points towards an exciting new application of magnetic resonance elastography as a diagnostic tool to diagnose and quantify disease progression. Interestingly, however, while the stiffness seems to be a sensitive marker for tau-pathology, neuronal loss, and inflammation, this is not the case for amyloid-pathology [65].
3.9.8 Does Our Brain Stiffness Change After Death?
An important unanswered question remains how brain tissue properties measured ex vivo compare to the tissue response in vivo. Several experimental setups have been designed to tackle this issue. Indentation experiments on rats, for instance, showed that the shear modulus obtained in vivo is about 31% higher than that obtained in vitro [118, 152]. Similarly, in situ indentation yielded approximately 30 to 50% higher shear moduli than ex vivo indentation [63]. These observations agree well with measurements using ultrasound elastography, where the shear modulus in vivo was about 47% higher than that given by ex vivo measurements [105]. A recent study using magnetic resonance elastography confirms that porcine brain tissue appears stiffer in vivo than ex vivo at frequencies of 100 and 125 Hz [72]. At lower frequencies, however, they found closer agreement between ex vivo and in vivo measurements. Contrary to these finding, other magnetic resonance elastography studies found an increase in shear moduli immediately after death [164], by up to 58% within only three minutes [176]. The origin of this rapid change within such a short period of time is likely of biochemical nature, but has not yet been explored to full extent.
3.9.9 Does Brain Stiffness Change Post Mortem?
Besides death itself, the post mortem storage time could potentially affect experimental results on brain tissue properties. Fig. 23 shows that, when kept intact and hydrated, bovine brain slices maintained their mechanical characteristics from nanoindentation throughout the entire testing period of five days post mortem. Also, in the time window of human brain experiments in Figs. 7, 8, 9, 10 and 11, we could not observe a notable change in tissue stiffness between samples that were tested first and last. This agrees well with a recent study using ultrasound elastography on Japanese big-ear rabbits, which reported that the change in mechanical properties is negligible at least within 1 hour after death [105]. In contrast, previous studies on porcine brain tissue have revealed a slight increase in tissue stiffness beginning 6h post mortem [61, 126]. This may be attributed to the fact that these studies were performed upon continued mechanical loading. During the experiments in Fig. 23, however, we minimized exposure to mechanical testing to clearly separate the effects of mechanical history discussed in Sects. 3.3–3.5 and post-mortem time. We conclude that if the experimental conditions are carefully chosen and the tissue is kept hydrated at all times, the degeneration process of post mortem brains is rather negligible. If brain tissue is stored without any liquid medium, however, the bio-molecular interactions and the mechanical strength of brain tissue deteriorate with prolonged storage duration, for instance due to the degeneration of myelin sheaths and the vacuolization of cristae [183].
Temporal variation of gray and white matter moduli. The consistent moduli within five days post mortem reveal that brain slices are virtually insensitive to the time of preservation. The stiffness increases moderately with indentation depth, from black dots to white dots. Gray matter, left, is consistently softer than white matter, right. Black horizontal lines indicate the mean; gray zones indicate the standard deviation. Adapted from [20]
3.9.10 Is Brain Stiffness Temperature-Dependent?
Most ex vivo experiments have been performed at room temperature. It is important to understand, how a rise in temperature from room to body temperature in the in vivo situation will affect the mechanical response of brain tissue. A recent study using ultrasound shear wave elastography on rabbit brains indicates that brain tissue stiffness decreases approximately linearly when the temperature increases from room temperature to body temperature, stays relatively constant in the range from 35 to \(42\,^{\circ }\hbox {C}\), and then rises again [104]. This is an interesting finding as according to those results, the stiffness of brain tissue seems to be constant exactly for the range of temperatures that may occur in vivo. It agrees well with dynamic shear tests on murine brain tissue in which brain tissue was stiffer at \(22\,^{\circ }\hbox {C}\) than at \(37\,^{\circ }\hbox {C}\) [133]. Uniaxial compression experiments on porcine brain tissue, on the contrary, show a slight but insignificant increase in stiffness from room temperature at \(22\,^{\circ }\hbox {C}\) to body temperature at \(37\,^{\circ }\hbox {C}\) [138].
4 Modeling Aspects
Computational modeling allows us to analyze and predict the behavior of human brain tissue under a variety of loading conditions. However, the value of a numerical prediction critically depends on the adequate choice of constitutive models. In the following, we will systematically propose mathematical formulations to capture the specific characteristics of brain tissue behavior discussed in Sect. 3.
The complexity of the tissue response depends on the loading conditions and so does the appropriate modeling approach. Different constitutive relations may be needed for the same material depending on the particular application. In this section, we will introduce constitutive relations of increasing complexity to capture the elastic, viscoelastic, and poroelastic behavior of brain tissue. We will then make application-specific suggestions towards selecting an appropriate model in the subsequent Sect. 5.
Due to the high compliance of brain tissue and the distinct nonlinearity of the tissue response, even for strains of only 1% as discussed in Sect. 3.1, we limit ourselves to constitutive models using the nonlinear field theory of mechanics. To characterize the kinematics of finite deformation, we introduce the deformation map \(\varvec{\varphi }\), which maps position \(\varvec{X}\) from the undeformed, unloaded configuration, \({\mathcal {B}}_0 \in {\mathbb {R}}^3\), to its new position \(\varvec{x} = \varvec{\varphi } \, (\varvec{X}, t)\) in the deformed, loaded configuration—the current placement of the body at time t, \({\mathcal {B}}_t\) [78]. We further introduce the deformation gradient \(\varvec{F}(\varvec{X},t)=\nabla _{\varvec{X}} \varvec{\varphi }(\varvec{X},t)\) to map undeformed line elements to deformed line elements, where \(\varvec{X}\) and \(\varvec{x}\) denote the position vectors in the unloaded reference and loaded spatial configurations, respectively. The principal stretches \(\lambda _{\mathrm{a}}\), \(\mathrm{a}=\{1,2,3\}\) are the square roots of the eigenvalues of the left and right Cauchy–Green tensors defined by \(\varvec{b}=\varvec{F} \cdot \varvec{F}^{\mathrm{t}}\) and \(\varvec{C}=\varvec{F}^{\mathrm{t}} \cdot \varvec{F}\), respectively.
4.1 Hyperelasticity of Brain Tissue
In a first step, we focus on the time-independent response of brain tissue, neglecting viscous or porous contributions, namely the experimental findings presented in Sects. 3.1 and 3.2 . The main time-independent characteristics are nonlinearity and compression-tension asymmetry. We postulate the existence of a strain-energy function \(\psi (\varvec{F})\), which is defined per unit reference volume and only depends on the deformation gradient \(\varvec{F}\). We note that several previous studies proposed fiber-reinforced material models for brain tissue, where the strain energy not only depends on the deformation gradient \(\varvec{F}\) but also on the fiber direction \(\varvec{f}_0\) [6, 37, 52, 60, 66, 128, 181]. Based on our experimental findings in Sect. 3.7, however, we suggest that the elastic behavior of brain tissue is, to a first approximation, isotropic. Nonetheless, the anisotropy induced by the orientation of nerve fibers may be important for other mechanical processes including damage or diffusion [175].
4.1.1 Hyperelastic Constitutive Modeling
Several phenomenological, isotropic strain-energy functions have been proposed to describe the constitutive behavior of brain tissue [22, 41, 92, 108, 137]. Most of these models were originally developed for much stiffer materials such as polymers [130] and calibrated using a single loading mode only [92, 108, 137]. Here, we evaluate the capability of previously proposed isotropic hyperelastic constitutive models to capture the time-independent response of human brain tissue under multiple loading conditions. We consider three special cases of the generalized Ogden model with the strain-energy function [130], i.e.,
where the constitutive parameters \(\alpha _i\) correspond to the strain-magnitude-sensitive nonlinear characteristics of the tissue. The classical shear modulus, known from the linear theory, is given by \(\mu =1/2\sum _{i=1}^{n} \mu _i \, \alpha _i\) [78]. Firstly, we adopt the neo-Hookean model with \(\alpha _1=2\) and \(\mu _1=\mu\), i.e.,
Secondly, we use the Mooney-Rivlin model with \(\alpha _1=2\), \(\mu _1=C_1=\mu -C_2\), \(\alpha _2=-2\) and \(\mu _2=C_2\) according to
and thirdly we reformulate the one-term Ogden model in terms of the classical shear modulus \(\mu =\alpha _1\mu _1/2\) and the parameter \(\alpha =\alpha _1\), i.e.,
In addition, we consider an exponential strain-energy function proposed by Demiray [42] as
and a rapidly strain-stiffening material model proposed by Gent [64],
Following standard arguments of continuum thermodynamics, we can express the Piola stress tensor \(\varvec{P}\) as the derivative of the strain-energy function \(\psi\) with respect to the deformation gradient \(\varvec{F}\) [78]. Assuming that brain tissue deforms homogeneously and isochorically with the incompressibility constraint \(\det \varvec{F} =1\), we may provide an analytical prediction of the Piola stresses
where \(\varvec{n}_{\mathrm{a}}\) and \(\varvec{N}_{\mathrm{a}}\) are the eigenvectors of the left and right Cauchy–Green strain tensors and p serves as a Lagrange multiplier [78]. We can then compare the stresses predicted by the model to experimentally observed responses.
4.1.2 Parameter Identification
Figure 24 illustrates the performance of the hyperelastic constitutive models (2) to (6) to represent the conditioned brain tissue response in different regions, including the cortex, basal ganglia, corona radiata, and corpus callosum during multiple loading conditions, uniaxial compression, uniaxial tension, and simple shear, simultaneously. Table 1 summarizes the resulting region-specific material parameters.
Different hyperelastic strain-energy functions (neo-Hooke, Mooney-Rivlin, Demiray, Gent, and Ogden) calibrated with the average conditioned experimental data of brain tissue under multiple loading modes, simple shear, compression, and tension, from four different brain regions: cortex, basal ganglia, corona radiata, and corpus callosum. Adapted from [22]
As the shear response of brain tissue deviates from linearity, even for small amounts of shear, neither the neo-Hookean nor the Mooney-Rivlin material model are able to satisfactorily represent the experimental data, which is clearly visible in Fig. 24. Only the one-term Ogden model is able to represent all loading modes simultaneously. It not only predicts the nonlinearity but also inherently captures the compression-tension asymmetry with a notably softer response in tension than in compression. These characteristics are controlled by the material parameter \(\alpha\): the higher the absolute value of \(\alpha\), the higher the degree of nonlinearity; if \(\alpha >2\), tensile stresses are stiffer than compressive stresses, if \(\alpha <2\), we observe the opposite. We note that in simple shear, shear stresses predicted by the constitutive model are independent of the sign of \(\alpha\). This implies that we have to be cautious when calibrating the constitutive model exclusively with simple shear data [22].
Combined compression/tension-shear loadings: sinusoidal simple shear superimposed on axial stretch \(\lambda =1.0\), 0.95, 0.9, 0.85, 0.8, 0.75, 1.05, 1.1, 1.15, 1.2, and 1.25. Average experimental ‘elastic’ shear stress versus amount of shear for white matter brain tissue (left) compared to analytical (center) and numerical (right) prediction
Figure 25 further demonstrates that the modified one-term Ogden model is able to capture the inherent characteristic of brain tissue that shear stresses increase significantly with increasing superimposed axial compression but only slightly with increasing axial tension, with coefficients of determination \(R^2>0.91\). This behavior is a logical outcome of the compression-tension asymmetry of brain tissue and can only be captured by one of the strain-energy functions compared in Fig. 24: the modified one-term Ogden model with a negative nonlinearity parameter \(\alpha\).
Calibrating the constitutive model with combined compression/tension-shear data yields a similar value for the shear modulus \(\mu\) as simultaneously calibrating the model with the data from multiple uniaxial loading modes in Table 1, bottom. However, the absolute value of \(\alpha\) is much lower. A high absolute value of \(\alpha\) with \(\alpha \approx -20\) yields unrealistically high shear stresses for high compressive or tensile pre-strain in the combined loading case. In contrast, a low absolute value of \(\alpha\) with \(\alpha \approx -7\) is not capable of representing the nonlinearity of the shear stress versus amount of shear curve reasonably well. For the sequence of multiple uniaxial loading modes in Fig. 24, the load is limited to 10% strain in compression and tension, and 20% in shear, to not damage the tissue during the course of the experiment. Due to the distinct nonlinearity of the stress-strain curve, even for those relatively small strains, the value \(\alpha\) obtained from uniaxial experiments would predict unrealistically high stresses for larger strains. This demonstrates that the one-term Ogden model, a phenomenological model in nature, can easily predict an unrealistic behavior when exceeding the deformation range used for parameter identification.
Consistent with these findings, all studies in the literature that considered both compression and tension experiments, reported that only the one or two-term Ogden models could satisfactorily represent the material response [58, 121, 165]. These studies proposed \(\mu =1.0\) kPa for cyclic compression-tension experiments on human white matter tissue [58], \(\mu =0.8\) kPa and \(\alpha =-4.7\) for mixed porcine brain tissue [121], and \(\mu =0.3\)-0.7 kPa and \(\alpha =-7.0\) when extrapolating tensile porcine white matter data to compression [165]. The lower absolute values for \(\alpha\) can be attributed to strains of \(30\%\) and more.
In contrast, Studies considering each loading mode individually found excellent agreement between experimental data of mixed porcine brain tissue and the Demiray, Gent, and Ogden strain-energy functions [22, 137, 139, 140]. Similarly, a study based on indentation data reported that polynomial, Yeoh, and one-term Ogden models agreed well with experimental data using an inverse finite elements analysis [92]. A more recent study on nanoindentation experiments, in contrast, suggests that the neo-Hookean model best represents indentation data [108].
This emphasizes that, due to the highly complex mechanical response of brain tissue, constitutive models derived from a single loading mode are not necessarily valid for different loading conditions. We conclude that the one-term Ogden model is able to capture the mechanical response of human brain tissue under multiaxial loading modes. However, particular caution is necessary when determining the parameter \(\alpha\): The compression-tension asymmetry pre-supposes a negative sign for \(\alpha\) and high absolute values yield unrealistically high stresses for large strains and multiaxial loading cases.
4.1.3 (In)homogeneous Deformation
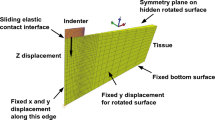
Effects of inhomogeneous deformation. Finite element simulations reveal the inhomogeneous deformation and stress states during simple shear as well as uniaxial compression and tension loadings, which can be observed during experiments (a). Analytically determined material parameters lead to an overestimation of compressive and tensile stresses and an underestimation of shear stresses when used in finite element simulations (b)
The results presented in Figs. 24 and 25, as well as in Table 1, are based on the assumption that brain tissue deforms homogeneously during uniaxial compression and tension as well as in simple shear. However, due to the high compliance of brain tissue and the fact that the upper and lower faces of the specimens are glued to the specimen holders during testing, the deformation actually displays certain inhomogeneities. Figure 26 contrasts the actual deformation of the specimens during simple shear, compression, and tension loadings with finite element simulations using the analytically determined parameters in Table 1.
Figure 26 indicates that material parameters calibrated for a homogeneous response tend to underestimate the shear stresses and overestimate the compressive and tensile stresses compared to using inhomogeneous finite element simulations, here exemplary shown for the neo-Hookean and the one-term Ogden model. Especially regarding the one-term Ogden model, however, the model predictions mostly remain within the standard deviations of the experiments.
Figure 25 demonstrates another effect resulting from the inhomogeneous deformation state. Using the analytically calibrated material parameters to numerically simulate combined loading conditions yields lower shear stresses than those predicted analytically, which is in accordance with the results in Fig. 24. However, the simulations capture the qualitative effect that shear stresses not only increase with superimposed compressive strains, but also slightly increase with superimposed tensile strains in accordance with the experimental results in Fig. 25. Taken together, the numerical results in Figs. 25 and 26 emphasize the importance of using an inverse parameter identification scheme to determine appropriate material parameters for computational simulations of brain tissue behavior in the future [167].
In addition to an inverse parameter identification, numerical simulations are valuable to optimize experimental procedures or testing protocols. They allow us—in advance—to evaluate the sensitivity of material parameters towards certain loading conditions, which will help to explicitly design experiments that are suitable for accurate and unambiguous parameter identification.
Numerical study on the influence of specimen geometry and specimen height h on the recorded tissue response. Stresses show a significant increase for small specimen heights. A rectangular cross-sectional area leads to a direction-dependent response in simple shear, which vanishes when the specimen height approaches zero
One effect that can not be captured analytically is the effect of specimen geometry. Figure 27 illustrates how the specimen geometry, height and cross-sectional area, affect the recorded stresses. Stresses significantly increase when specimens become too thin as the deformation inhomogeneities at the fixed faces gain in influence. For compression and tension loadings, a specimen height of approximately 5 mm seems optimal to ensure a consistent response. For simple shear loadings, the effect of specimen height is noticeable independent of the height; however, with increasing height the effect of the cross-sectional area becomes more prominent.
4.1.4 (In)compressibility
The results presented in Fig. 24 and Table 1 are based on the assumption that brain tissue is incompressible, motivated by the high water content of approximately 80%. This assumption may be adequate for impact situations [88, 99]; however, especially when considering the time-independent, quasi-static response of brain tissue relevant for extremely slow processes such as brain development, cerebrospinal fluid may escape through the ventricular system, as discussed in Sect. 3.5. This will lead to a slight compressibility, which we can model by adding a volumetric contribution to the modified one-term Ogden model [78, 130],
where \(\psi ^{\mathrm{Ogd}}_{{{{\mathrm{iso}}}}{}}\) describes the isochoric response, \(\psi ^{\mathrm{Ogd}}_{{{{\mathrm{vol}}}}{}}\) describes the purely volumetric response, \({\tilde{\lambda }}_{\mathrm{a}}=J^{-1/3}{\lambda }_\mathrm{a}\) with \(\mathrm{a}=\{1,2,3\}\) are the volume-preserving parts of the principal stretches, and \(\kappa\) denotes the bulk modulus.
To demonstrate the influence of compressibility on the brain tissue response during unconfined experiments, we used the material parameters calibrated analytically under the assumption of incompressibility in Table 1 in a finite element setting and varied the Poisson’s ratio \(\nu\) from 0.3 to 0.49. We ensured that the results for \(\nu =0.49\) were not affected by locking effects when using linear finite elements through a comparison with the results using mixed finite elements to deal with quasi-incompressibility. Figure 28 shows how tissue compressibility affects the response during unconfined compression, tension, and simple shear. Expectedly, independent of the loading mode, a decrease in the Poisson’s ratio also leads to a decrease in tissue stresses. Interestingly, a recent study argues that a different compressibility in compression than in tension leads to the experimentally observed compression-tension asymmetry [167]. From a physical perspective, such an effect could be attributed to poro-elastic effects, which will be discussed in more detail in Sect. 4.3.
4.1.5 Conclusions and Future Perspectives
The one-term Ogden model inherently captures the main characteristics of the time-independent response of brain tissue, the nonlinearity and compression-tension asymmetry. It is capable of representing multiple loading modes simultaneously. However, material parameters calibrated analytically assuming incompressibility and a homogeneous deformation will tend to overestimate compressive and tensile stresses and underestimate shear stresses when used in finite element simulations. To address these limitations, we recommend calibrating the model parameters with sophisticated inverse identification schemes, which capture inhomogeneous deformation states and several loading modes simultaneously. A remaining open challenge is to identify a single model that captures a wide range of strains.
Depending on the loading conditions, it may be appropriate to model human brain tissue as an incompressible or compressible solid [88, 99]. If movement of free flowing cerebrospinal fluid into the ventricles and the subarachnoid space is possible, for instance during slow processes such as brain development, brain tissue effectively changes its local volume [148]. Therefore a slight compressibility with a Poisson’s ratio of 0.45 to 0.49 seems appropriate [26]. In addition, parameters should be calibrated using the conditioned, drained tissue response. In impact situations, in contrast, the fluid offers resistance and contributes to tissue stiffness. In these case, the tissue may behave incompressibly and parameters should be calibrated using the unconditioned tissue response.
4.2 Finite Viscoelasticity
While hyperelasticity can describe the nonlinearity and compression-tension asymmetry as outlined in Sects. 3.1 and 3.2 , it cannot capture the characteristic time-dependent effects, the highly hysteretic behavior of Sect. 3.3 [22, 58, 121], the significant strain-rate-dependence of Sect. 3.4, and the characteristic conditioning as pointed out in Sect. 3.5. The time-dependent nature of brain tissue is associated with various physical mechanisms and time scales: The motion of fluid within the solid network of cells and extracellular matrix introduces a poroelastic behavior [48, 58], whereas intracellular interactions between the cytoplasm, nucleus, and the cell membrane trigger a viscoelastic response [84]. Importantly, when tested below a critical threshold, the conditioning behavior of brain tissue is fully recoverable, as we conclude from Fig. 15: No damage or permanent softening occurs, which justifies to model the time-dependent response of brain tissue using visco- or poro-viscoelasticity instead of irreversible damage or the Mullin’s effect [58].
In this section, we will extend the hyperelastic model presented in Sect. 4.1 to account for time-dependent, viscoelastic effects. The simplest and most popular approach to characterize the time-dependent behavior of brain tissue is to use a Prony series [20, 57, 100, 120, 121, 134, 139]. A Prony series approach is equivalent to a generalized Maxwell model for linear viscoelasticity in relaxation type loading [89]. However, the Prony series approach has two major limitations: it is restricted to linear elasticity and is computationally expensive [41]. To account for the large deformability of brain tissue discussed in Sect. 2.1, instead, we consider a class of viscoelastic models within the general setting of finite deformation continuum mechanics. We multiplicatively decompose the deformation gradient into elastic and inelastic parts [153], additively decompose the free-energy function into an equilibrium and non-equilibrium parts [154], and introduce internal variables to account for the rate-dependent behavior [142].
4.2.1 Kinematics of Finite Viscoelasticity
To model the viscoelastic nature of brain tissue, we decompose the deformation gradient \(\varvec{F}\) into elastic and viscous parts,
where i denotes the parallel arrangement of m viscoelastic elements [153], as exemplary shown for two viscoelastic elements in Fig. 29. We can then introduce the spatial velocity gradient as
Multiplicative decomposition model, where \(\varvec{F}\) is associated with the main elastic network characterized by material parameters \(\mu _{\infty }\) and \(\alpha _{\infty }\), \(\varvec{F}^{{{{\mathrm{v}}}}{}}_1\) is the viscous damper associated with fluid flow inside the cell characterized by the viscosity \(\eta _1\) with the corresponding hyperelastic spring \(\varvec{F}^{{{{\mathrm{e}}}}{}}_1\) with parameters \(\mu _1\) and \(\alpha _1\), and \(\varvec{F}^{{{{\mathrm{v}}}}{}}_2\) is the viscous damper associated with fluid flow within the solid network of cells and extracellular matrix characterized by the viscosity \(\eta _2\) with the corresponding hyperelastic spring \(\varvec{F}^{{{{\mathrm{e}}}}{}}_2\) with parameters \(\mu _2\) and \(\alpha _2\). Adapted from [23]
and decompose it additively into elastic parts, \(\varvec{l}_i^{\mathrm{e}} = \dot{\varvec{F}}{}_{i}^{\mathrm{e}} \cdot (\varvec{F}_{i}^{\mathrm{e}}){}^{-1}\), and viscous parts, \(\varvec{l}_i^{\mathrm{v}} = \varvec{F}_i^{\mathrm{e}} \cdot \dot{\varvec{F}}{}_i^{\mathrm{v}} \cdot (\varvec{F}_{i}^{\mathrm{v}}){}^{-1} \cdot (\varvec{F}_i^{\mathrm{e}}){}^{-1}\). From the deformation gradient, we determine the left Cauchy–Green tensor \(\varvec{b}\) and its spectral representation in terms of the principal stretches \(\lambda _{\mathrm{a}}\) and the Eulerian eigenvectors \(\varvec{n}_{\mathrm{a}}\),
It is convenient to introduce the elastic left Cauchy–Green tensor for each mode, i.e.
with eigenvalues \((\lambda _{i\,\mathrm a}^{\mathrm{e}})^2\) and eigenvectors \(\varvec{n}_{i\,\mathrm a}^{\mathrm{e}}\), which are, in general, not identical to the eigenvectors of the total left Cauchy–Green tensor, \(\varvec{n}_{i\,\mathrm a}^{\mathrm{e}} \ne \varvec{n}_{\mathrm{a}}\). The material time derivative of the elastic left Cauchy–Green tensor \(\varvec{b}^\mathrm{e}_i\) is
which introduces its Lie-derivative, i.e.
along the velocity field of the material motion.
4.2.2 Viscoelastic Constitutive Modeling
Hysteresis versus maximum stress during the first and third loading cycles for all specimens tested in shear, compression, and tension. The data indicate that the dissipated energy increases linearly with the maximum recorded stress. Adapted from [23]
Motivated by the experimental findings discussed in Sect. 3.7, we assume an isotropic, incompressible material response for both the elastic and the viscoelastic behavior. We introduce the viscoelastic free-energy function \(\psi\) as the sum of three terms, an equilibrium part \(\psi ^{\mathrm{eq}}\) in terms of the total principal stretches \(\lambda _{\mathrm{a}}\), a non-equilibrium part \(\psi ^{\mathrm{neq}} = \sum _{i=1}^{m} \psi _{i}\) in terms of the \(i=1,\ldots ,m\) elastic principal stretches \(\lambda _{i\mathrm{a}}^\mathrm{e}\), and a term \(p \, [\,J-1\,]\) that enforces the incompressibility constraint \(J-1=0\) via the Lagrange multiplier p [78], i.e.
Similarly, we introduce the stress power \({\mathcal {P}}\) as the sum of an equilibrium part \({\mathcal {P}}^{\mathrm{eq}}=\varvec{\tau }^\mathrm{eq}:\varvec{l}\) in terms of the equilibrium Kirchoff stress \(\varvec{\tau }^{\mathrm{eq}}\) and a non-equilibrium part \({\mathcal {P}}^\mathrm{neq}=\varvec{\tau }^{\mathrm{neq}}:\varvec{l}\) in terms of the non-equilibrium Kirchoff stress \(\varvec{\tau }^{\mathrm{neq}} = \sum _{i=1}^{m} \varvec{\tau }_{i}\), i.e.
We can then evaluate the dissipation inequality \({\mathcal {D}} ={\mathcal {P}} - {\dot{\psi }} \ge 0\) in terms of the individual equilibrium and non-equlibrium contributions as
With the assumption of isotropy, we can rewrite the non-equilibrium stress power, \({\mathcal {P}}^{\mathrm{neq}} = \sum _{i=1}^m \!\! {[}{\varvec{\tau }}_i \cdot (\varvec{b}^{\mathrm{e}}_i)^{-1} {]}\!\! : \frac{1}{2}{} [\, \dot{\varvec{b}}{}^{\mathrm{e}} - {\mathscr {L}}_v \varvec{b}^{\mathrm{e}}_i \,]\), in terms of the Lie derivative of the elastic left Cauchy–Green tensor (14), and obtain the following explicit representation of the dissipation inequality
The first term of (18) vanishes when we define the equilibrium Kirchhoff stress tensor as
and the second term vanishes for non-equilibrium Kirchhoff stresses, i.e.
This leaves the reduced dissipation inequalities for each individual mode i, which is
It remains to specify the equilibrium and non-equilibrium parts of the free energy, \(\psi ^{\mathrm{eq}}\) and \(\psi ^{\mathrm{neq}}\), and the evolution of the internal variables \(\varvec{b}_i^{\mathrm{e}}\). For the equilibrium energy, we adopt a modified one-term Ogden model parameterized in terms of the total stretches \(\lambda _{{{{\mathrm{a}}}}{}}\) [130], which best represented the hyperelastic response of human brain tissue in Sect. 4.1, i.e.
with the equilibrium shear modulus \(\mu _{\infty }\) and the equilibrium compression-tension asymmetry parameter \(\alpha _{\infty }\) [22]. The derivative in (19) then becomes
For the non-equilibrium energy, we adopt the same Ogden strain-energy function
which introduces two additional parameters \(\mu _i\) and \(\alpha _i\) for each mode i, and is now parameterized in terms of the deviatoric elastic principal stretches \({{\tilde{\lambda }}}^\mathrm{e}_{i\,\mathrm{a}} =(J^{\mathrm{e}}_i)^{-1/3} \lambda ^{\mathrm{e}}_{i\,\mathrm{a}}\), the square roots of the eigenvalues of the isochoric part of the elastic left Cauchy–Green tensor \(\tilde{\varvec{b}}^\mathrm{e}_{i}=(J^{\mathrm{e}}_i){}^{-2/3} \varvec{b}^{\mathrm{e}}_i\), with \(J^{\mathrm{e}}_i= \det \varvec{F}^{\mathrm{e}}_{i}\). The derivatives in (20) then become
where \(\mathrm{a}, \mathrm{b}, \mathrm{c} = \{ 1, 2, 3 \}\) and \(\mathrm{a} \ne \mathrm{b}\), \(\mathrm{a} \ne \mathrm{c}\), and \(\mathrm{b} \ne \mathrm{c}\). Finally, we need to specify the temporal evolution of the viscoelastic kinematics. To a priori satisfy the reduced dissipation inequality (21) [142, 154, 170], and motivated by the linear relation between hysteresis and maximum stress during our experiments, as illustrated in Fig. 30, we choose the evolution equations for the internal variables \(\varvec{b}^{\mathrm{e}}_i\) as linear functions of the Kirchhoff stresses \({\varvec{\tau }}_i\) [18, 79], i.e.
This introduces one additional parameter for each mode i, the viscosity \(\eta _i>0\), or, when scaled with the corresponding shear modulus \(\mu _i\), the associated relaxation time, \(\tau _i = {\eta _i}/{\mu _i}\) [23, 142]. Since the internal variables \(\varvec{b}^{\mathrm{e}}_i\) are a linear function of the deviatoric Kirchhoff stress tensors \({\varvec{\tau }}_i\), the elastic deformation always remains volume preserving, \(J_i^{\mathrm{e}}=1\), and thus \(J_i^{\mathrm{v}}=J/J_i^{\mathrm{e}} = 1\). To advance the internal variables in time [70, 80, 142], we may adopt an implicit or explicit time integration scheme [23]. Finally, we calculate the Piola stress \(\varvec{P}\) as the right-sided pull back of the Kirchhoff stress \(\varvec{\tau }\), i.e.
We note that we may decompose the viscous parts of the spatial velocity gradient, \(\varvec{l}_i^{\mathrm{v}} = \varvec{F}_i^{\mathrm{e}} \cdot \dot{\varvec{F}}{}_i^{\mathrm{v}} \cdot (\varvec{F}_{i}^{\mathrm{v}}){}^{-1} \cdot (\varvec{F}_i^{\mathrm{e}}){}^{-1}\), into their symmetric and skew-symmetric parts, \(\varvec{l}_i^{\mathrm{v}}=\varvec{d}_i^{\mathrm{v}}+\varvec{w}_i^{\mathrm{v}}\,\), in terms of the viscous stretch rates, \(\varvec{d}_i^{\mathrm{v}} =\frac{1}{2}{} \, [ \, \varvec{l}_i^{\mathrm{v}} + (\varvec{l}_i^\mathrm{v})^{\mathrm{t}} \,]\), and the viscous spin rates, \(\varvec{w}_i^{\mathrm{v}} =\frac{1}{2}{} \, [ \, \varvec{l}^{\mathrm{v}}_i - (\varvec{l}^\mathrm{v}_i)^{\mathrm{t}} \,]\). If we now reformulate the reduced dissipation inequalities (21) using the symmetry of \(\varvec{\tau }_i\), we obtain
A comparison with equation (21) yields an alternative expression for the evolution equations for the internal variables (26), expressed in terms of the viscous stretch rates \(\varvec{d}_i^{\mathrm{v}}\), i.e.
which has been proposed by [80]. If all the rheological elements in the model were linear, it would reduce to the generalized Maxwell model, which is equivalent to the Prony series in a classical relaxation experiment [41, 155].
Representative conditioning behavior for cyclic loading with step-wise increasing maximum strain. Shear/nominal stress versus amount of shear/stretch behavior for specimens from the corona radiata; experiment (top row) and model prediction (bottom row). At a stretch level of 0.9, the difference between the first and second cycles indicates pronounced pre-conditioning. When increasing the stretch level to 0.8, the first curve initially follows the pre-conditioned behavior up until 0.9; beyond 0.9, the curve again displays pronounced pre-conditioning between the first and second cycles. The model is capable of predicting this history-dependence at different load levels. For simple shear loading, the qualitative model prediction slightly deviates from the experimental observations
Figure 31 demonstrates that the proposed viscoelastic constitutive model is capable of capturing the experimentally observed pronounced pre-conditioning during the first loading cycle and the moderate conditioning effects during all subsequent cycles [23]. The model also predicts the experimentally observed successive softening when increasing the strain in a step-wise fashion [23], as discussed in Sect. 3.5. In simple shear, however, the model predictions deviate from the experimental data, especially for the initial loading segment. Furthermore, the model predicts larger residual stresses upon completing the first loading cycle of each stretch level than observed in the experiment. This difference is likely caused by the pore fluid that squeezes out of the sample when it is first loaded [58]. Our monophasic viscoelastic model can only implicitly capture these porous effects [11]. To accurately model the fluid flow within the tissue, we will introduce a biphasic poro-viscoelastic model [33, 48, 68] in the next section to further contrast and compare viscous and porous effects in human brain tissue.
Numerical study on the influence of the strain rate \({{\dot{\epsilon }}}\) on the model predicted tissue response during cyclic simple shear, compression and tension experiments. The maximum stresses increase with increasing strain rate. For intermediate strain rates, the response is highly hysterestic; for extremely fast or slow rates, the hysteresis closes
Figure 32 demonstrates the effect of the strain rate on the observed stress response. The viscoelastic model nicely captures the experimentally observed increase in the maximum stresses with increasing strain rate, as discussed in Sect. 3.4. At both ends of the time scale spectrum—for extremely slow and extremely fast loadings—the hysteresis closes and the loading and unloading paths are almost identical. For intermediate time scales, as those performed during the experiments shown in Figs. 9 and 13, the response is notably hysteretic.
4.2.3 Parameter Identification
We have shown that the finite viscoelastic model can qualitatively capture the most important time-dependent characteristics of brain tissue including hysteresis, strain-rate-dependence, and conditioning. In order to use the proposed model in finite element simulations, we need to provide material parameters that also quantitatively capture the tissue response. Instead of following the common approach of previous studies [57, 120, 137] and calibrating the elastic parameters using only the initial loading path during quasi-static experiments and the viscous parameters using only the stress relaxation behavior, we integrate data from simple shear, unconfined compression, tension, shear relaxation, and compression relaxation experiments [22] to holistically characterize the overall behavior of brain tissue [23]. Notably, all tests were performed on one and the same samples to avoid that inter-specimen variations affect the results. By simultaneously considering the data of all five tests, we identified material parameters that characterize the behavior of the exact same tissue samples under all five loading modes. We further use the entire loading history for parameter identification—including unloading during cyclic experiments and the loading ramp for stress relaxation experiments [23].
Figures 33 and 34 demonstrate that the finite viscoelastic constitutive model with two viscoelastic elements is capable of representing the average experimental data during the first and third loading cycles associated with the unconditioned and conditioned response, respectively, for the four different brain regions, the cortex, the basal ganglia, the corona radiata, and the corpus callosum [23]. Tables 2 and 3 summarize the corresponding region-specific material parameters. The model slightly underestimates the shear response, especially during the initial loading segment, and slightly overestimates the stresses under tensile loading but nicely captures the relaxation behavior over the entire loading history.
Simultaneous identification of viscoelastic material parameters for the finite viscoelastic constitutive model with two generalized Maxwell elements and a single nonlinearity parameter \(\alpha =\alpha _i=\alpha _{\infty }\) for all five loading conditions. Average experimental data (solid lines) during the first loading cycle associated with the unconditioned response with standard deviations in four regions, cortex, basal ganglia, corona radiata, and corpus callosum with corresponding constitutive model (dashed lines) calibrated using data from all loading modes simultaneously, see Table 2. Adapted from [25]
Simultaneous identification of viscoelastic material parameters for the finite viscoelastic constitutive model with two generalized Maxwell elements and a single nonlinearity parameter \(\alpha =\alpha _i=\alpha _{\infty }\) for all five loading conditions. Average experimental data (solid lines) during the third loading cycle associated with the conditioned response with standard deviations in four regions, cortex, basal ganglia, corona radiata, and corpus callosum with corresponding constitutive model (dashed lines) calibrated using data from all loading modes simultaneously, see Table 3. Adapted from [25]
Figure 35 contrasts the shear moduli of the unconditioned and conditioned tissue responses in four brain regions [23]. Table 4 summarizes the corresponding characteristic time constants \(\tau _i=\eta _i\, /\, \mu _i\) near thermodynamic equilibrium. Shear moduli are generally lower for the conditioned than for the unconditioned response, which may be attributed to the contribution of free flowing fluid, which only offers resistance during initial loading but not during subsequent cycles.
The equilibrium shear modulus \(\mu _{\infty }\) displays similar regional trends for the unconditioned and conditioned responses. It is highest in the cortex, lowest in the corpus callosum, and comparable in basal ganglia and corpus callosum. Interestingly, we observe the same regional trends for the shear modulus \(\mu _2\) of the conditioned response in the Maxwell element with particularly high time constants \(\tau _2\). In contrast, the shear moduli \(\mu _1\) corresponding to lower time constants \(\tau _1\) are generally higher than \(\mu _{\infty }\) and \(\mu _2\), and show slightly different regional trends: White matter regions, corona radiata and corpus callosum, have stiffened relative to gray matter regions, cortex and basal ganglia.
Shear moduli calibrated with the unconditioned and conditioned tissue response in four brain regions: the cortex (C), the basal ganglia (BG), the corona radiata (CR), and the corpus callosum (CC). Adapted from [23]
It is interesting to note that the conditioned response displays higher time constants \(\tau _2\) than the unconditioned response, although we have calibrated both parameter sets with exactly the same stress relaxation experiments. This indicates that the cyclic experiments have a pronounced effect on the viscoelastic parameter identification. Comparing the unconditioned and conditioned behavior suggests that we can attribute the lower time scale \(\tau _1\), to the viscous component of the solid phase, and the higher time scale \(\tau _2\), to porous effects of the fluid phase: At short time scales, fluid moves inside the cell; at intermediate time scales, fluid moves through the porous solid skeleton of cells and extracellular matrix, as sketched in Fig. 29 [19]. When using the conditioned response of the third cycle for our calibration, we intentionally neglect the porous effect and, accordingly, the higher time constant adopts significantly larger values than for the unconditioned response. This agrees well with a previous study that has reported a pre-conditioned viscosity of 60 kPas in unconfined compression tests of porcine brain [135].
4.2.4 Conclusions and Future Perspectives
We have proposed a finite viscoelastic model that combines the hyperelastic Ogden model with two viscoelastic elements and can, in addition to the experimentally observed compression-tension asymmetry and nonlinearity, capture time-dependent effects including hysteresis according to Sect. 3.3, and the successive softening for stepwise loading according to Sect. 3.5. Importantly, we have shown that we can provide a single set of material parameters that simultaneously fits cyclic loadings and the early and late relaxation behavior of brain tissue. We have provided separate parameter sets for the unconditioned response using the first loading cycle and for the conditioned response using the third loading cycle. Expectedly, the unconditioned tissue response is markedly stiffer than the conditioned response. We would like to point out that there is no right or wrong set of parameters: Depending on the application of interest–for example the interpretation of an ex vivo test or the prediction of an in vivo response–either the unconditioned or conditioned parameter set seems to be more physiological.
Through our viscoelastic parameter identification, we have obtained insights that help us to better understand the individual time-dependent contributions of the solid and fluid phases [58]. The first viscoelastic element corresponds to time constants on the order of 1 to 3 seconds and can be associated with the viscoelasticity of individual cells, while the second viscoelastic element corresponds to time constants between 20 and 100 seconds and can represent the movement of fluid within the solid skeleton of cells and extracellular matrix. Such insights become especially important when we attempt to interpret and model brain tissue as a biphasic material [33, 48, 57], which will be the subject of the next section.
While our simultaneous analysis of multiple loading conditions has significantly improved the parameterization compared to previous viscoelastic models based on a single loading mode [13, 80, 135], the asymmetry predicted by our model is still less pronounced than observed in our experiments. Furthermore, the parameters are based on the assumption of a homogeneous deformation state. To improve the fit of the model, an inverse parameter identification scheme could be used in the future to potentially address these concerns. Further open challenges regarding the viscoelastic modeling of brain tissue remain the discrepancies between model and experiment under shear loading and the overestimation of the tensile stresses.
4.3 Finite Poro-viscoelasticity
To explicitly model the biphasic nature of brain tissue, we can combine the finite viscoelastic model for brain tissue introduced in Sect. 4.2 with the theory of porous media [47]. Importantly, a poro-elastic model alone would not be sufficient to model the hysteretic behavior of brain tissue [39].
4.3.1 Kinematics of Poro-viscoelasticity
Kinematics of the material configuration \({\mathcal {B}}_0\), and the spatial configurations \({\mathcal {B}}_t\) and \({\mathcal {B}}_{\tau }\) of a biphasic material body within the context of finite poro-elasticity [39]
We assimilate brain tissue to a biphasic material consisting of an immiscible aggregate of a solid skeleton (S) fully saturated with pore fluid (F). Both components are assumed to be incompressible. The solid phase represents the network of cells and extracellular matrix—including the fluid trapped inside the cells and within the extracellular matrix—while the pore fluid exclusively represents the free-flowing fluid in brain tissue. Figure 36 illustrates that, at a given time t, the constituents simultaneously occupy the same spatial position \(\varvec{x}\) in the current configuration \({\mathcal {B}}_t\). The primary unknowns of finite poro-viscoelasticity are the solid deformation \(\varvec{x}=\varvec{\varphi }(\varvec{X}_{\!S},t)\) and the fluid pore pressure p. The strong form of the governing equations in the reference configuration reads
where the Piola stress \(\varvec{P}(\varvec{F}_{\!S},p)\) and the seepage velocity in the reference configuration \(\varvec{W}(\varvec{F}_{\!S},p )\) are functions of the solid deformation gradient \(\varvec{F}_{\!S}=\partial \varvec{x} / \partial \varvec{X}_{\!S}\), and the fluid pore pressure p, \(\varvec{B}_0\) is the body force vector in the reference configuration, and \(J_{\!S}= \mathrm {det} \varvec{F}_{\!S}\) is the solid Jacobian. The seepage velocity describes the velocity of the fluid phase with respect to the deforming solid. Following the considerations in the previous section, we involve finite viscoelasticity by decomposing the solid deformation gradient into elastic and viscous parts, \(\varvec{F}_{\!S}=\varvec{F}_{\!S}^{\mathrm{{e}}} \cdot \varvec{F}_{\!S}^{\mathrm{{v}}}\), and introduce the elastic left Cauchy–Green tensor \(\varvec{b}^{\mathrm{{e}}} =\varvec{F}_{\!S}^{\mathrm{{e}}} \cdot (\varvec{F}_{\!S}^{\mathrm{{e}}})^{{{{\mathrm{t}}}}{}} = \sum _{\mathrm{a}=1}^3 (\lambda _{\mathrm{a}}^{\mathrm{{e}}})^2 \varvec{n}_\mathrm{a}^{\mathrm{{e}}} \otimes \varvec{n}_{\mathrm{a}}^{\mathrm{{e}}}\), with eigenvalues \((\lambda _{\mathrm{a}}^{\mathrm{{e}}})^2\) and eigenvectors \(\varvec{n}_\mathrm{a}^{\mathrm{{e}}}\), \(\mathrm{a}=1,2,3\).
4.3.2 Constitutive Modeling—Solid Component
We further additively split the strain-energy function of the solid skeleton into an equilibrium part \(\psi ^{\mathrm{{eq}}}\) and a non-equilibrium part \(\psi ^{\mathrm{{neq}}}\), \(\psi _{\!S} = \psi ^{\mathrm{{eq}}} + \psi ^{\mathrm{{neq}}}\). The total free-energy function then reads \(\psi = \psi ^{\mathrm{{eq}}}(\lambda _i) + \psi ^{\mathrm{{neq}}}(\lambda _i^{\mathrm{{e}}}) + U(J_{\!S})\), where \(U(J_{\!S})\) is an extension function that describes the compressibility effects of the poroelastic material, including the concept of a compaction point. We select a function that introduces the Lamé parameter \(\lambda\) of the solid component, \(U(J_{\!S}) = \lambda [ 1 - n_{S0} ]^2 [[J_{\!S}-1]/[1-n_{S0} ] - \text {ln} ( [J_{\!S}-n_{S0}]/[1-n_{S0}])]\), where \(n_{S0}\) is the initial porosity of the biphasic material, i.e., the volume fraction of the solid component in the reference configuration at time \(t_0\) [47]. The overall Piola stress that represents the stress state in the solid is
where \(\varvec{I}\) is the unit tensor, and the volumetric part of the extension function is \(\varvec{\tau }^{\mathrm{vol}} = \lambda [1-n_{S0}]^2 [J_{\!S}/[1-n_{S0}]- J_{\!S}/[J_{\!S}-n_{S0}]] \; \varvec{I}\).
4.3.3 Constitutive Modeling—Fluid Component
In addition to the constitutive equations for the solid, we need to specify a material model for the fluid defined by the seepage velocity \(\varvec{W} = \varvec{w} \cdot \text{ cof }\varvec{F}\). Standard approaches have used Darcy’s law to describe the motion of the pore fluid with respect to the deforming solid component in terms of the changing pressure gradients in the biphasic material and its deformation-dependent permeability [47, 113]. In this case, the seepage velocity is defined as
where \(\mu _F\) and \(\rho _F\) are the effective fluid viscosity and fluid density, \(\varvec{K}_S\) is the intrinsic permeability tensor, and \(\varvec{g}\) is the vector of gravity acceleration. For example, we can assume an isotropic, deformation-dependent intrinsic permeability tensor \(\varvec{K}_S=( [J_S - n_{S0} ] / [ 1 - n_{S0} ] )^{\kappa }\varvec{K}_{S0}\), where the initial intrinsic permeability tensor \(\varvec{K}_{S0}=K_{S0} \, \varvec{I}\) would reduce to the weighted identity tensor when the material permeability is isotropic [113]. Unfortunately, our current experimental data are not sufficient to adequately calibrate the poro-viscoelastic constitutive model. To truly calibrate a poro-viscoelastic model, we would have to perform experiments with drained and undrained conditions [58] to quantify the—potentially region- and loading-mode-dependent—permeability of brain tissue and to clearly differentiate between viscous and porous effects.
4.3.4 Conclusions and Future Perspectives
While the finite viscoelastic constitutive model introduced in Sect. 4.2 can implicitly capture poro-elastic effects through its two independent time scales, a poro-elastic model alone is insufficient to describe the hysteretic behavior of brain tissue [99]. The advantage of using a more complex biphasic poro-viscoelastic model is that it allows us to explicitly control the physics of the fluid as it moves within the solid skeleton of cells and extracellular matrix. Notably, we could even further advance the poro-viscoelastic constitutive model by adding components such as the blood, which flows inside the small brain capillaries [48]. In this case, brain tissue is permeated by two viscous pore liquids, the interstitial fluid and the blood. An important drawback concerning the poro-viscoelasticity of brain tissue is that, to date, the poro-elasticity of brain tissue has not yet been well characterized experimentally. Consolidation tests performed on human white matter brain tissue have shown that brain tissue behaves similiarly to filled elastomers [58]. However, the data available in the literature are not sufficient to adequately calibrate the poro-viscoelastic model presented in Sect. 4.3. Well designed experiments will be essential to quantify region-specific permeability of brain tissue and to differentiate between viscous and porous effects. This will be critical when choosing the appropriate constitutive model for a different biomedical application.
5 Application-Specific Considerations
Computational modeling can form a cornerstone in improving diagnostics and treatment of brain tissue injury and disease. However, reliable, clinically relevant predictions of brain tissue behavior critically depend on the appropriate choice of constitutive models and material parameters for the numerical simulations. Importantly, the required model complexity depends on the application of interest and the corresponding length- and time-scales. Brain deforms differently during trauma at high rates, surgery at medium rates, and brain development at low rates. To save computational cost, we should evaluate efficiency versus accuracy, and choose a model that is as complex as necessary but as simple as possible. In the following, we present selected examples in which numerical simulations can valuably assist medical diagnostics and treatment of neurological disorders. We discuss the relevant length- and time-scales and propose an appropriate modeling approach. We intend to demonstrate how the evaluation of relevant scales, as summarized in Fig. 6, can help to choose the appropriate mechanical model for a certain application.
5.1 Brain Development
Cortical folding is a classical hallmark of most mammalian brains and closely correlates with brain function and dysfunction. During development, the initially smooth surface evolves into an elaborately convoluted pattern. Growing evidence suggests that mechanical instabilities emerging from differential growth between a faster growing outer gray matter, the cortex, and a slower growing inner white matter, the subcortex, play a major role in brain morphogenesis. We can model brain growth by complementing the kinematics of finite deformation introduced in Sect. 4 by the kinematics of finite growth [144]. We multiplicatively decompose the deformation gradient \(\varvec{F}\) into an elastic part \(\varvec{F}^{{{{\mathrm{e}}}}{}}\) and a growth part \(\varvec{F}^{{{{\mathrm{g}}}}{}}\) [144], \(\varvec{F} = \nabla _{\varvec{{X}}} \varvec{\varphi } = \varvec{F}^{{{{\mathrm{e}}}}{}} \cdot \varvec{F}^{{{{\mathrm{g}}}}{}}\). A key challenge is to prescribe the growth tensor \(\varvec{F}^{{{{\mathrm{g}}}}{}}\) and its evolution in time to realistically mimic the underlying biological processes [3] in the developing brain. In the simplest case, growth is purely morphogenetic, evolving in time irrespective of the mechanical environment [117]. In more complex cases, growth is mechanically driven, for instance by hypertension in arteries [98] or by hyperstretch in skin [159].
Inspired by cellular mechanisms during brain development [27], we assume that both cortex and subcortex grow isotropically with \(\varvec{F}^{{{{\mathrm{g}}}}{}} = \vartheta \; \varvec{I}\), where \(\vartheta\) is a scalar-valued growth multiplier. The grown volume \(J^{{{{\mathrm{g}}}}{}} = \text{ det } \, \varvec{F}^{{{{\mathrm{g}}}}{}} = \vartheta ^3\) is identical to the growth multiplier cubed \(\vartheta ^{{{{\mathrm{3}}}}{}}\). We further assume that cortical growth is purely morphogenetic [2]—independent of mechanical stress or strain [4]—growing linearly in time at the growth rate \(G_{{{{\mathrm{c}}}}{}}\) = const, and that subcortical growth is stretch-dependent at a growth rate \(G_{{{{\mathrm{s}}}}{}}\), as illustrated in Fig. 37 [21, 26]. This yields the evolution of the cortical growth multiplier, \({\dot{\vartheta }}_{{{{\mathrm{c}}}}{}} = G_{{{{\mathrm{c}}}}{}}\) and the subcortical growth multiplier, \({\dot{\vartheta }}_{{{{\mathrm{s}}}}{}} = G_{{{{\mathrm{s}}}}{}} \left< J^{{{{\mathrm{e}}}}{}} - J^{{{{\mathrm{0}}}}{}} \right> = G_{{{{\mathrm{s}}}}{}} \left<J \,/\, \vartheta ^3 - J^{{{{\mathrm{0}}}}{}} \right>\). The term in the Macaulay brackets \(\left<J^{\mathrm{e}} - J^{\mathrm{0}}\right >\) activates growth only if the elastic volume stretch \(J^{{{{\mathrm{e}}}}{}}\) exceeds the threshold value \(J^{{{{\mathrm{0}}}}{}}\), i.e., when axons are stretched beyond their physiological limit [43]. Taken together, the mechanical model of brain growth enables us to bridge the scales from accumulation and growth of individual cells into the growth kinematics of cortex and subcortex, and ultimately cortical folding [67].
Continuum model for cortical and subcortical growth. The cortex, the gray matter, grows morphogenetically at a constant rate \(G_{{{{\mathrm{c}}}}{}}\). Cortical growth induces subcortical deformation, which triggers subcortical growth. The subcortex, the white matter, grows at a stretch-dependent rate as \(G_{{{{\mathrm{s}}}}{}} \left<J^{{{{\mathrm{e}}}}{}} - J^{0}\right>\), where \(G_{{{{\mathrm{s}}}}{}}\) mimics the axon elongation rate and \(\left<J^{{{{\mathrm{e}}}}{}}-J^{{{{\mathrm{0}}}}{}}\right>\) activates growth only, if the elastic volume stretch \(J^{{{{\mathrm{e}}}}{}}\) exceeds its baseline value \(J^{{{{\mathrm{0}}}}{}}\). Adapted from [28]
Variety of mammalian brains. Mammalian brains hugely vary in size, shape, and convolutional complexity but only little in cortical thickness (top row), adapted from [179]. Numerical simulation of a growing shell on a spheroidal substrate for increasing ratio of elliptical radius to cortical thickness R / t (bottom row). Folding complexity increases with absolute size
Figure 38 demonstrates that the model explains why the surface-to-volume ratio in different mammalian brains increases disproportionally with brain size and why brains of larger mammals tend to be more folded than brains of smaller animals [179]. From a mechanics point of view, this phenomenon is a merely geometric effect and independent of phylogenetic effects—contrary to the common notion in developmental biology [179]. The results in Fig. 38 demonstrate the great opportunities of using mechanical models to understand brain development. However, the emerging folding pattern is the result of an instability problem, which, by its very nature, is highly sensitive to the constitutive model and material parameters of finite element simulations.
Sensitivity of the evolving surface morphology during brain development towards the constitutive model used for brain tissue. With increasing absolute value of the nonlinearity parameter \(\alpha\) from top to bottom, as introduced in Sect. 4.1, the wavelength increases and the primary pattern transitions from sinusoidal wrinkles to cusped sulci
Concerning the choice of the appropriate modeling approach, brain folding occurs over weeks and months of gestation and we can safely assume that the cerebrospinal fluid has time to escape through the ventricular system and that its contributions to the tissue stiffness do not change markedly during the simulation period. Consequently, we propose to use an isotropic hyperelastic constitutive model calibrated with the conditioned experimental response. However, we need to pay caution with regard to the material parameters since extremely large deformations will occur in the bottom of sulci—the valleys of cortical folds. If we use the material parameters calibrated using experimental data up to 10% compressive and tensile strains, and 20% shear strains from Table 1, the model will predict unrealistically high stresses in the valleys of emerging folds and unrealistic folding patterns as evident in the bottom row of Fig. 39. This is a natural result of the corresponding nonlinearity parameter \(\alpha =-20\). Even for a slightly reduced material nonlinearity with \(\alpha =-7\) in the center row, according to the parameters identified using combined compression/tension-shear experiments, the folding pattern still deviates from the one observed in the real brain. Surprisingly, we obtain the most realistic patterns for the neo-Hookean material model with \(\alpha =2\) in the top row of Fig. 39. These results emphasize that we need to be careful when using material parameters for deformation states that exceed the experimentally investigated ranges of strains. For the application of cortical folding, where high compression occurs locally in the developing sulci, the neo-Hookean model indeed seems to be the best choice. Alternatively, in a more physiological model, we could account for the continuous adaption of the living brain tissue to mechanical loading—for example, motivated by the accumulated cell death in the bottom of the sulci—which gradually reduces the tissue stresses in these regions.
5.2 Neurodegenerative Diseases
Spatio-temporal progression of toxic protein across the brain for different initial seeding regions associated with a, b Alzheimer’s disease, c Parkinson’s disease, and d amyotrophic lateral sclerosis. Adapted from [175]
Because of demographic changes worldwide, neurodegeneration will, without a doubt, become a major challenge in medicine and public health. More than 45 million people are living with dementia today and this number is expected to triple by 2050 [1]. A major challenge of neurodegenerative diseases is that the initial symptoms of cognitive decline become noticeable only one or two decades after the first pathological abnormalities have started to affect the brain [166]. For several decades, researchers have tried to establish early predictive indicators of neurodegeneration and identify common pathological themes of neurodegenerative disorders. Computational simulations are now increasingly recognized as a powerful tool to correlate biochemical and biomechanical events and provide early biomarkers for neurodegenerative disorders [175].
Figure 40 shows how we can use a physics-based reaction-diffusion model to identify common underlying schemes in the pathogenesis of Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis [174]. The underlying simulation combines the classical Fisher-Kolmogorov equation [55], \({\dot{c}} = \text{ Div } ( \varvec{D} \cdot \nabla c) + \alpha \, c \, [\,1-c\,]\) for the toxic protein concentration c with an anisotropic diffusion model, \(\varvec{D} = d^{\mathrm{{ext}}} \varvec{I} + d^{\mathrm{{axn}}} \, \varvec{n} \otimes \varvec{n}\), with a pronounced diffusion along the axonal direction \(\varvec{n}\). Here \(\alpha\) is the growth rate of toxic protein, and \(d^{\mathrm{{ext}}}\) and \(d^{\mathrm{{axn}}}\) are the diffusion parameters associated with extracellular diffusion and axonal transport, respectively. The model correctly predicts amyloid-\(\beta\) deposits and tau inclusions in Alzheimer’s disease, \(\alpha\)-synuclein inclusions in Parkinson’s disease, and TDP-43 inclusions in amyotrophic lateral sclerosis and displays excellent agreement with the histological patterns observed in diseased human brains [87]. When integrated across the brain, the concentration profiles result in biomarker curves that display a striking similarity with the sigmoid shape and qualitative timeline of clinical biomarker models [82]. Regions of high toxic protein concentration in Fig. 40 are likely correlated with neuronal death and tissue atrophy, which we can model as negative tissue growth, \(\varvec{F} = \varvec{F}^{{{{\mathrm{e}}}}{}} \cdot \varvec{F}^{{{{\mathrm{a}}}}{}}\). To model the biochemical effects of neurodegeneration, we could introduce the atrophy tensor \(\varvec{F}^{{{{\mathrm{a}}}}{}}\) as a function of the toxic protein concentration c [175]. To model the biomechanical effects of neurodegeneration, we recommend using a finite hyperelastic model and assume that viscoelastic and poro-elastic effects are negligible on the relevant time scales on the order of several decades [75]. While the computational modeling of neurodegeneration is still in its infancy, there seems to be a general agreement that a more quantitative understanding of the spatio-temporal spreading of neurodegenerative diseases is necessary to establish a prognostic timeframe of disease progression. This could have important clinical implications, ranging from more accurate estimates of the socioeconomic burden of neurodegeneration to a more informed design of clinical trials and pharmacological intervention.
5.3 Hydrocephalus
Hydrocephalus is a medical condition in which excess cerebrospinal fluid builds up in the ventricles of the brain. The excess fluid can increase the size of the ventricles and, with it, the intracranial pressure. Normal pressure hydrocephalus develops gradually in time and manifests itself in enlarged ventricles, while the intracranial pressure remains at normal physiological levels [73]. The common treatment strategy is cerebrospinal fluid flow diversion and surgical placement of a shunt to drain excess fluid into the abdomen. The improvement after such a neurosurgical shunt procedure varies greatly with a morbidity of about 40–50% [102]. Therefore, there is an urgent need for numerical simulations that assist the design of better diagnostic and treatment protocols.
Efforts towards this direction have employed mostly two-dimensional finite element models using poro-elasticity with a linear elastic solid skeleton [88, 103, 125], or the multiple-network poroelastic theory [161]. Regarding the conclusions in Sect. 4.3, we infer that it may be important to also take viscoelastic effects into account, which has been done in [34]. Most recent works even propose a triphasic mechano-electrochemical theory [45]. This model additionally accounts for a change in the ionic concentrations of the ventricular cerebrospinal fluid in the absence of an elevated intracranial pressure as a possible cause for the onset of normal pressure hydrocephalus. Interestingly, a three-dimensional finite element study found no major advantage by using a biphasic instead of a single phase model for brain tissue [46].
Based on the different time scales–rather rapid in hydrocephalus in young children versus gradual in normal pressure hydrocephalus in the elderly–either a poro-viscoelastic or a purely poro-elastic model could be more appropriate versus more efficient. It remains to carefully evaluate the influence of viscous and porous effects to decide which model to use. Intracranial pressure monitoring is critical in many neurological conditions. The current gold standard is to monitor the intracranial pressure via catheter transducer systems that are inserted into the cranium. This process is highly invasive and there is hope that computational simulations can help estimate the intracranial pressure less invasively.
5.4 Drug Delivery
A promising treatment approach for neurological disorders is the infusion of therapeutic agents, generally known as convection-enhanced drug delivery [16]. Infusion catheters are placed through small holes in the skull directly into the extra-vascular space of the brain parenchyma. The pressure gradient generated by external medical pumps initiates an interstitial fluid flow and, therewith, the distribution of the therapeutic agents. Various poro-elastic mechanical models have been developed to simulate this problem analytically [122] or numerically [32, 48, 155]. Drug delivery is a typical example where it seems indispensible to model the fluid phase of brain tissue explicitly by using a biphasic constitutive model based on the findings in Sect. 4.3 and due to the fact that infusion of therapeutic agents occurs at intermediate time scales, it might also be important to include viscous effects by using a poro-viscoelastic model.
5.5 Neurosurgery
During neurosurgical procedures, brain tissue can temporarily experience strains and stresses that are high enough to induce tissue damage. In this respect, simulation tools capable of predicting the level of stress and strain can be highly valuable to optimize surgical procedures [99]. A typical example is decompressive craniectomy, a traditional but controversial surgical procedure that removes part of the skull to allow an injured and swollen brain to expand outward and release an elevated intracranial pressure [93]. Despite a marked increase in popularity, a decompressive craniectomy remains a compromise between maximizing control of the intracranial pressure and minimizing side effects to the surrounding tissue [69]. To date, the precise criteria when to perform a decompressive craniectomy, the optimal timing of treatment, the optimal location and size of the skull opening, and the long-term functional outcome remain unclear [171]. Studies suggest that mechanical strains beyond the physiological level are associated with its undesired, high failure rates. However, the precise strain fields induced by the craniectomy are unknown.
Decompressive craniectomy. Displacements and maximum principal strains for unilateral and frontal flaps with left and right, left, and right hemispherical swelling. Swelling causes maximum principal strains of up to 30% localized around the opening. Reprinted with permission from [178]
Figure 41 shows finite element simulations of a decompressive craniectomy performed on a personalized head model from magnetic resonance images [178]. The simulations compare the two most common types of craniectomy, a unilateral flap and a frontal flap. For each case, either both hemispheres or the left or right hemisphere of the brain are swollen and part of the skull is removed to allow the brain to bulge outward and release the intracranical swelling pressure. Clearly, the underlying swelling is a biochemical process associated with the absorption of fluid, and we can best model this phenomenon using a poro-viscoelastic approach. While swelling takes place on the order of hours, the clinical procedure of skull opening takes place on the order of minutes on which both porous and viscous effects can become relevant. The simulations in Fig. 41 reveal three potential failure mechanisms associated with the procedure: axonal stretch in the center of the bulge, axonal compression at the edge of the opening, and axonal shear around the opening [171]. Strikingly, for a swelling of only 10%, axonal strain, compression, and shear reach local maxima of up to 30%, and exceed the reported functional and morphological damage thresholds on the order of 20% [8]. Computational simulations of surgical procedures can help quantify brain deformation, tissue strain, axonal stretch, and shear with the goal to identify high-risk regions for brain damage on a personalized basis [56]. While computational modeling is beyond clinical practice in neurosurgery today, simulations of neurosurgical procedures have the potential to rationalize surgical process parameters including timing, location, and size, and provide standardized guidelines for clinical decision making and neurosurgical planning.
5.6 Traumatic Brain Injury
A major public health concern are the consequences of neurotrauma with over 2 million people affected by traumatic brain injury each year [111]. During traumatic brain injury, external mechanical load leads to damage of the highly delicate brain tissue and ultimately loss of brain function [116]. Clinically, traumatic brain injury can be classified into mild, moderate, and severe, and repetitive mild traumatic brain injuries are now more commonly referred to as chronic traumatic encephalopathy [75]. While computational models can help estimate injury risk and advance injury prevention [38], correct predictions of the deformation field during impact critically depends on the accurate representation of the nonlinear, loading-mode-, and region-dependent stress-strain relationship of brain tissue. This is especially challenging since impact loading involves extremely high strain rates. It is not surprising that previous studies have shown that simulation results largely depend on both the choice of boundary conditions, for instance concerning the brain-skull-interface, and the choice of the constitutive model for brain tissue, as illustrated in Fig. 42 [184]. If only interested in the instant of the impact, which can occur at strain rates as high as 100/s, we can assume that the hysteresis loop closes, as shown in Fig. 32, and use a finite hyperelastic model. Importantly, however, the corresponding parameters should be calibrated using the unconditioned tissue response at equally high rates. Contrary to slow processes such as brain development or neurodegeneration, during traumatic brain injury, the fluid will not drain and will contribute to the tissue stiffness. In accordance with the considerations in Sect. 4.1.2, the Ogden hyperelastic constitutive model has shown best agreement with experimental results [74, 168]. Interestingly, numerical studies further indicate that using a linear viscoelastic model may overestimate the deformations within the brain and, consequently, overestimate the injury risk if strain-based injury criteria are used [168]. In turn, a study investigating traumatic spinal cord injury in rats found that the compressible Ogden model with a two-term Prony series [110] achieved good agreement with impactor weight-drop experiments.
Cumulative maximum principal strains during high-rate impact using different material models for brain tissue (top). Radial-circumferential shear strains resulting from in vivo head rotation (bottom). Reprinted with permission from [184]
Taken together, time-dependent effects are critically important, especially on the small time scales of moderate to severe traumatic brain injury. More important than choosing between an elastic, a viscoelastic, or a poro-viscoelastic model is the use of appropriate high strain rate experiments to accurately calibrate the model. In addition to the correct prediction of tissue strains and stresses through numerical simulations, a major research effort in the coming years will be to accurately determine reliable tolerance criteria for cell death and tissue damage to improve injury diagnosis and advance injury prevention. Another major direction in the coming years will be the exploration of the effects of multiple mild repeated injuries to the head with a quest to better understand how tissue damage accumulates over time and how we can best prevent neurodegeneration in chronic traumatic encephalopathy.
6 Conclusions and Future Perspectives
6.1 Recommendations
The response of brain tissue–more than any other tissue in our body–is hugely sensitive to the length and time scales during loading. This is mainly a result of the ultrasoft, gel-like nature of brain tissue, which implies that biochemical effects on very small length and time scales can have tremendous effects on the overall behavior. At first glance, it appears that previous experimental findings are largely contradictory; in reality, these studies tested different “shades” of brain, as illustrated in Fig. 6.
To capture the highly complex mechanical response of brain tissue, several hyper-, visco-, poro-, or poro-viscoelastic constitutive models have been proposed. In this review, we have systematically introduced and discussed models of different level of complexity. Our goal was to show that the selection of an appropriate model, and with it an appropriate set of material parameters, hugely depends on the application of interest: Slow processes such as brain development may be modeled using finite hyperelasticity, moderately fast processes such as brain surgery are predominated by visco- and poro-elastic effects, and extremely fast processes such as brain injury are dominated by the behavior at very high rates.
Concerning material parameters, we have to keep in mind that parameters calibrated from a single experimental loading mode will likely not represent the physiological response of the brain under complex deformation states occurring in vivo. Similarly, parameters for the simulation of slow processes should be calibrated using the conditioned tissue response, while parameters for fast processes should be based on the unconditioned response, ideally probed at equally high rates. Using the appropriate set of material parameters for a particular application is critical because simulated strains outside the calibrated strain range can easily under- or over-estimate the mechanical effect on the brain, and, with it, safety criteria and injury risk. We conclude that due to the inherent bio-chemo-mechanical complexity of the brain, it is critically important to design experiments hand in hand with the modeling approach, and, ideally, even with the clinical application in mind.
6.2 Challenges and Perspectives
We conclude by listing our top ten most important steps that we consider critical to push the frontiers in understanding the mechanics of the human brain:
-
To date, there is not a single constitutive model with a single set of material parameters capable of correctly predicting the response of brain tissue for small and large strains, as well as fast and slow loading rates. Especially for extreme loading situations, associated with the applications in Sect. 5, this drawback should be tackled in the future, through a combination of well-designed experiments and careful modeling.
-
A challenge in characterizing the constitutive behavior of the brain is its pronounced microstructural heterogeneity. First steps towards capturing local variations in tissue stiffness have been made by using phenomenological models and calibrating regional material parameters. A true step forward would be to design microstructurally motivated constitutive models, which would inherently capture regional heterogeneities at a finer resolution.
-
Closely related to the previous point, to date, it is not clear, which microstructural components control macroscopic tissue mechanics. While most microstructural investigations have focused on the cellular components—the functional elements of brain tissue—from a mechanics point of view, the role of the extracellular matrix should be investigated more thoroughly. Notably, however, load bearing elements in the extracellular matrix of other soft tissues such as collagen do not play a major role in the brain, which explains its incredible compliance compared to other tissues.
-
Another open question remains, how the in vivo properties of brain tissue compare to the reported ex vivo parameters. Hard tissues have been extensively studied because their mechanical properties change very slowly and only marginally from live to dead. Soft tissues like tendons, ligaments, or arteries, change more drastically but over the past decade, we have learned how to preserve them appropriately post mortem. Ultrasoft tissues like the brain can undergo drastic and rapid changes, both biochemically and biomechanically. To fully understand their in vivo behavior, it will be key to intelligently combine in vivo and ex vivo experiments to develop and calibrate reliable mechanical models for brain tissue in the future.
-
In addition to the appropriate choice of the constitutive model and the appropriate evaluation of strains, it is essential to accurately capture the boundary conditions during finite element simulations of brain tissue behavior. Especially in the context of high impact loading, during moderate to severe traumatic brain injury, the brain-skull interface and internal structures like the falx will significantly affect the deformation profile, and, with it the strains and stresses across the brain. The appropriate choice of boundary conditions needs to be carefully addressed in the future.
-
Once we understand the constitutive behavior of healthy brain tissue, a next critical step is to predict the risk of axonal injury and tissue damage, and, ultimately, the progression of neurodegenerative disease. It will be essential to perform well-designed experiments to quantify the time course of events during neurodegeneration, and to identify reliable failure criteria under multiaxial loading conditions. Again, these failure criteria will likely depend on the loading mode, as well as the time and length scale of loading.
-
An important phenomenon that has not been widely considered yet is the fact that brain tissue is not passive but that it contains living cells that actively respond to mechanical stimuli. Brain cells sense and respond to their mechanical environment. Understanding the role of mechanosensing and mechanotransduction is critical in directing migration paths of nervous cells or tumor cells within nervous tissue and in guiding the remodeling of the tissue microstructure.
-
Understanding the mechanobiology of brain tissue requires a deep biochemical knowledge of the mechanical stimuli that can modulate the response of the brain. Recent studies have highlighted the importance of mechanical stiffness as a guiding principle in neurodevelopment and neurodegeneration. Manipulating the mechanical microenvironment and recreating developmental conditions could hold the key to enhance axonal sprouting, trigger axonal regrowth, and simulate remyelination and provide a new path towards new treatment strategies for neuronal regeneration.
-
To achieve the long-term goal of using mechanical models for clinical applications, a major challenge is the numerical implementation of sophisticated, highly coupled multiscale and multiphysics problems. When simulating whole brain phenomena, the design of efficient and robust algorithms will be critical to make these models applicable and useful for the community at large.
-
Finally, validating the models, parameters, and simulations is the most critical step in building confidence in the computational modeling of the brain. A validation across the scales, both in space and time, will require concerted efforts of biologists, engineers, and clinical researchers. Large data sets of healthy and diseased brains have already been collected and are being made available to the broad public. Extracting valuable information from these data sets will likely require tools of machine learning to extract valuable features and correlations in space and time. Rather than using machine learning tools as a black box, without any prior information, our mechanical understanding of the brain can define important constraints for these tools to reduce the parameter space and ensure that the results are physically and physiologically meaningful.
With the immense progress in biomedicine, imaging, and computation, we now have the tools to address these frontiers. If we succeed in tackling these challenges, mechanical modeling and computational simulation of the brain will be highly valuable to advance everyday clinical practice by assisting diagnosis and treatment of neurological diseases.
References
Alzheimer’s Disease International (2016) Improving healthcare for people living with dementia. Coverage, quality and costs now and in the future. World Alzheimer Report
Amar MB, Goriely A (2005) Growth and instability in elastic tissues. J Mech Phys Solids 53(10):2284–2319
Ambrosi D, Ateshian G, Arruda E, Cowin S, Dumais J, Goriely A, Holzapfel GA, Humphrey J, Kemkemer R, Kuhl E et al (2011) Perspectives on biological growth and remodeling. J Mech Phys Solids 59(4):863–883
Ambrosi D, Mollica F (2002) On the mechanics of a growing tumor. Int J Eng Sci 40(12):1297–1316
Antonovaite N, Beekmans SV, Hol EM, Wadman WJ, Iannuzzi D (2018) Structure-stiffness relation of live mouse brain tissue determined by depth-controlled indentation mapping. arXiv preprint arXiv:1802.02245
Arbogast KB, Margulies SS (1999) A fiber-reinforced composite model of the viscoelastic behavior of the brainstem in shear. J Biomech 32(8):865–870
Awdry WV (1946) Thomas the tank engine. Edmund Ward Ltd, Leicester
Bain AC, Meaney DF (2000) Tissue-level thresholds for axonal damage in an experimental model of central nervous system white matter injury. J Biomech Eng 122(6):615–622
Barnes JM, Przybyla L, Weaver VM (2017) Tissue mechanics regulate brain development, homeostasis and disease. J Cell Sci 130(1):71–82
Bayly PV, Clayton EH, Genin GM (2012) Quantitative imaging methods for the development and validation of brain biomechanics models. Annu Rev Biomed Eng 14:369–396
Bilston LE (2011) Neural tissue biomechanics. Springer, Heidelberg
Bilston LE (2018) Soft tissue rheology and its implications for elastography: challenges and opportunities. NMR Biomed 31(10):e3832
Bilston LE, Liu Z, Phan-Thien N (2001) Large strain behaviour of brain tissue in shear: some experimental data and differential constitutive model. Biorheology 38(4):335–345
Blümcke I, Vinters HV, Armstrong D, Aronica E, Thom M, Spreafico R (2009) Malformations of cortical development and epilepsies. Epileptics Disord 11(3):181–193
Blumenthal NR, Hermanson O, Heimrich B, Shastri VP (2014) Stochastic nanoroughness modulates neuron-astrocyte interactions and function via mechanosensing cation channels. Proc Natl Acad Sci U S A 111(45):16124–16129
Bobo RH, Laske DW, Akbasak A, Morrison PF, Dedrick RL, Oldfield EH (1994) Convection-enhanced delivery of macromolecules in the brain. Proc Natl Acad Sci U S A 91(6):2076–2080
Bock AS, Olavarria JF, Leigland LA, Taber EN, Jespersen SN, Kroenke CD (2010) Diffusion tensor imaging detects early cerebral cortex abnormalities in neuronal architecture induced by bilateral neonatal enucleation: an experimental model in the ferret. Front Syst Neurosci 4:149
Boyce MC, Parks DM, Argon AS (1988) Large inelastic deformation of glassy polymers. Part i: rate dependent constitutive model. Mech Mater 7:15–33
Budday S, Andres S, Walter B, Steinmann P, Kuhl E (2017) Wrinkling instabilities in soft bilayered systems. Philos Trans A Math Phys Eng Sci 375:1–11
Budday S, Nay R, de Rooij R, Steinmann P, Wyrobek T, Ovaert TC, Kuhl E (2015) Mechanical properties of gray and white matter brain tissue by indentation. J Mech Behav Biomed Mater 46:318–330
Budday S, Raybaud C, Kuhl E (2014) A mechanical model predicts morphological abnormalities in the developing human brain. Sci Rep 4:5644. https://doi.org/10.1038/srep05644
Budday S, Sommer G, Birkl C, Langkammer C, Haybaeck J, Kohnert J, Bauer M, Paulsen F, Steinmann P, Kuhl E, Holzapfel GA (2017) Mechanical characterization of human brain tissue. Acta Biomater 48:319–340
Budday S, Sommer G, Haybaeck J, Steinmann P, Holzapfel GA, Kuhl E (2017) Rheological characterization of human brain tissue. Acta Biomater 60:315–329
Budday S, Sommer G, Paulsen F, Holzapfel G, Steinmann P, Kuhl E (2018) Region- and loading-specific finite viscoelasticity of human brain tissue. Proc Appl Math Mech 18:1–2
Budday S, Sommer G, Steinmann P, Holzapfel GA, Kuhl E (2017) Viscoelastic parameter identification of human brain tissue. J Mech Behav Biomed Mater 74:463–476
Budday S, Steinmann P, Kuhl E (2014) The role of mechanics during brain development. J Mech Phys Solids 72:75–92
Budday S, Steinmann P, Kuhl E (2015) Physical biology of human brain development. Front Cell Neurosci 9:257
Budday S, Steinmann P, Kuhl E (2015) Secondary instabilities modulate cortical complexity in the mammalian brain. Philos Mag 95(28–30):3244–3256
Chatelin S, Constantinesco A, Willinger R (2010) Fifty years of brain tissue mechanical testing: from in vitro to in vivo investigations. Biorheology 47(5–6):255–276
Chatelin S, Vappou J, Roth S, Raul J-S, Willinger R (2012) Towards child versus adult brain mechanical properties. J Mech Behav Biomed Mater 6:166–173
Chen F, Zhou J, Li Y, Wang Y, Li L, Yue H (2015) Mechanical properties of porcine brain tissue in the coronal plane: interregional variations of the corona radiata. Ann Biomed Eng 43(12):2903–2910
Chen X, Sarntinoranont M (2007) Biphasic finite element model of solute transport for direct infusion into nervous tissue. Ann Biomed Eng 35(12):2145–2158
Cheng S, Bilston LE (2007) Unconfined compression of white matter. J Biomech 40(1):117–124
Cheng S, Bilston LE (2010) Computational model of the cerebral ventricles in hydrocephalus. J Biomech Eng 132(5):054501
Christ AF, Franze K, Gautier H, Moshayedi P, Fawcett J, Franklin RJ, Karadottir RT, Guck J (2010) Mechanical difference between white and gray matter in the rat cerebellum measured by scanning force microscopy. J Biomech 43(15):2986–2992
Clayton EH, Genin GM, Bayly PV (2012) Transmission, attenuation and reflection of shear waves in the human brain. J R Soc Interface 9(76):2899–2910
Cloots R, Van Dommelen J, Nyberg T, Kleiven S, Geers M (2011) Micromechanics of diffuse axonal injury: influence of axonal orientation and anisotropy. Biomech Model Mechanobiol 10(3):413–422
Cloots RJ, Van Dommelen J, Kleiven S, Geers M (2013) Multi-scale mechanics of traumatic brain injury: predicting axonal strains from head loads. Biomech Model Mechanobiol 12(1):137–150
Comellas E, Steinmann P (2017) A non-linear poroelastic constitutive model for the computational modelling of brain tissue behaviour. In: XIV international conference on computational plasticity
Darvish K, Crandall JR (2001) Strain conditioning in the dynamic viscoelastic response of brain tissue. In: Proceedings of the 2001 bioengineering conference ASME, pp 893–894
de Rooij R, Kuhl E (2016) Constitutive modeling of brain tissue: current perspectives. Appl Mech Rev 68(1):010801
Demiray H (1972) A note on the elasticity of soft biological tissues. J Biomech 5(3):309–311
Dennerll TJ, Lamoureux P, Buxbaum RE, Heidemann SR (1989) The cytomechanics of axonal elongation and retraction. J Cell Biol 109(6):3073–3083
Donnelly B, Medige J (1997) Shear properties of human brain tissue. J Biomech Eng 119(4):423–432
Drapaca CS, Fritz JS (2012) A mechano-electrochemical model of brain neuro-mechanics: application to normal pressure hydrocephalus. Int J Numer Anal Mod Ser B 1:82–93
Dutta-Roy T, Wittek A, Miller K (2008) Biomechanical modelling of normal pressure hydrocephalus. J Biomech 41(10):2263–2271
Ehlers W, Eipper G (1999) Finite elastic deformations in liquid-saturated and empty porous solids. Transp Porous Media 34(1):179–191
Ehlers W, Wagner A (2015) Multi-component modelling of human brain tissue: a contribution to the constitutive and computational description of deformation, flow and diffusion processes with application to the invasive drug-delivery problem. Comput Methods Biomech Biomed Eng 18(8):861–879
Elkin BS, Ilankovan A, Morrison B (2010) Age-dependent regional mechanical properties of the rat hippocampus and cortex. J Biomech Eng 132(1):011010
Estes M, MacElhaney JH (1970) Response of brain tissue to compressive loading. In: American society of mechanical engineers biomechanical and human factors conference
Fallenstein G, Hulce VD, Melvin JW (1969) Dynamic mechanical properties of human brain tissue. J Biomech 2(3):217–226
Feng Y, Lee C-H, Sun L, Ji S, Zhao X (2017) Characterizing white matter tissue in large strain via asymmetric indentation and inverse finite element modeling. J Mech Behav Biomed Mater 65:490–501
Feng Y, Okamoto RJ, Namani R, Genin GM, Bayly PV (2013) Measurements of mechanical anisotropy in brain tissue and implications for transversely isotropic material models of white matter. J Mech Behav Biomed Mater 23:117–132
Finan JD, Elkin BS, Pearson EM, Kalbian IL, Morrison B III (2012) Viscoelastic properties of the rat brain in the sagittal plane: effects of anatomical structure and age. Ann Biomed Eng 40(1):70–78
Fisher RA (1937) The wave of advance of advantageous genes. Ann Eugen 7:355–369
Fletcher TL, Kolias AG, Hutchinson PJA, Sutcliffe MPF (2014) The wave of advance of advantageous genes. PLoS ONE 9:e102131
Forte AE, Gentleman SM, Dini D (2016) On the characterization of the heterogeneous mechanical response of human brain tissue. Biomech Model Mechanobiol 16:907–920
Franceschini G, Bigoni D, Regitnig P, Holzapfel GA (2006) Brain tissue deforms similarly to filled elastomers and follows consolidation theory. J Mech Phys Solids 54(12):2592–2620
Galford JE, McElhaney JH (1970) A viscoelastic study of scalp, brain, and dura. J Biomech 3(2):211–221
Garcia-Gonzalez D, Jérusalem A, Garzon-Hernandez S, Zaera R, Arias A (2018) A continuum mechanics constitutive framework for transverse isotropic soft tissues. J Mech Phys Solids 112:209–224
Garo A, Hrapko M, Van Dommelen J, Peters G (2007) Towards a reliable characterisation of the mechanical behaviour of brain tissue: the effects of post-mortem time and sample preparation. Biorheology 44(1):51–58
Gefen A, Gefen N, Zhu Q, Raghupathi R, Margulies SS (2003) Age-dependent changes in material properties of the brain and braincase of the rat. J Neurotrauma 20(11):1163–1177
Gefen A, Margulies SS (2004) Are in vivo and in situ brain tissues mechanically similar? J Biomech 37:1339–1352
Gent A (1996) A new constitutive relation for rubber. Rubber Chem Technol 69(1):59–61
Gerischer LM, Fehlner A, Köbe T, Prehn K, Antonenko D, Grittner U, Braun J, Sack I, Flöel A (2018) Combining viscoelasticity, diffusivity and volume of the hippocampus for the diagnosis of alzheimer’s disease based on magnetic resonance imaging. NeuroImage Clin 18:485–493
Giordano C, Kleiven S (2014) Connecting fractional anisotropy from medical images with mechanical anisotropy of a hyperviscoelastic fibre-reinforced constitutive model for brain tissue. J R Soc Interface 11(91):20130914
Goriely A, Budday S, Kuhl E (2015) Chapter two-neuromechanics: from neurons to brain. Adv Appl Mech 48:79–139
Goriely A, Geers MG, Holzapfel GA, Jayamohan J, Jérusalem A, Sivaloganathan S, Squier W, van Dommelen JA, Waters S, Kuhl E (2015) Mechanics of the brain: perspectives, challenges, and opportunities. Biomech Model Mechanobiol 14(5):931–965
Goriely A, Weickenmeier J, Kuhl E (2016) Stress singularities in swelling soft solids. Phys Rev Lett 117:138001
Govindjee S, Reese S (1997) A presentation and comparison of two large deformation viscoelastic models. J Eng Mater Technol 119(5):251–255
Green MA, Bilston LE, Sinkus R (2008) In vivo brain viscoelastic properties measured by magnetic resonance elastography. NMR Biomed 21(7):755–764
Guertler CA, Okamoto RJ, Schmidt JL, Badachhape AA, Johnson CL, Bayly PV (2018) Mechanical properties of porcine brain tissue in vivo and ex vivo estimated by mr elastography. J Biomech 69:10–18
Hakim S, Adams R (1965) The special clinical problem of symptomatic hydrocephalus with normal cerebrospinal fluid pressure: Observations on cerebrospinal fluid hydrodynamics. J Neurol Sci 2(4):307–327
Hardy WN, Foster CD, Mason MJ, Yang KH, King AI, Tashman S (2001) Investigation of head injury mechanisms using neutral density technology and high-speed biplanar x-ray. Stapp Car Crash J 45:337–368
Harris TC, de Rooij R, Kuhl E (2019) The shrinking brain: cerebral atrophy following traumatic brain injury. Ann Biomed Eng. https://doi.org/10.1007/s10439-018-02148-2
Harrison P (1999) The neuropathology of schizophrenia: a critical review of the data and their interpretation. Brain 1(122):593–624
Hiscox LV, Johnson CL, McGarry MD, Perrins M, Littlejohn A, van Beek EJ, Roberts N, Starr JM (2018) High-resolution magnetic resonance elastography reveals differences in subcortical gray matter viscoelasticity between young and healthy older adults. Neurobiol Aging 65:158–167
Holzapfel GA (2000) Nonlinear solid mechanics: a continuum approach for engineering science, vol 24. Wiley, Chichester
Holzapfel GA, Simo JC (1996) A viscoelastic constitutive model for continuous media at finite thermomechanical changes. Int J Solids Struct 33:3019–3034
Hrapko M, Van Dommelen J, Peters G, Wismans J (2006) The mechanical behaviour of brain tissue: large strain response and constitutive modelling. Biorheology 43(5):623–636
Iwashita M, Kataoka N, Toida K, Kosodo Y (2014) Systematic profiling of spatiotemporal tissue and cellular stiffness in the developing brain. Development 141(19):3793–3798
Jack CR, Holtzman DM (2013) Biomarker modeling of Alzheimer’s disease. Neuron 80:1347–1358
Jannesar S, Allen M, Mills S, Gibbons A, Bresnahan JC, Salegio EA, Sparrey CJ (2018) Compressive mechanical characterization of non-human primate spinal cord white matter. Acta Biomater 74:260–269
Jérusalem A, Dao M (2012) Continuum modeling of a neuronal cell under blast loading. Acta Biomater 8(9):3360–3371
Jin X, Zhu F, Mao H, Shen M, Yang KH (2013) A comprehensive experimental study on material properties of human brain tissue. J Biomech 46(16):2795–2801
Johnson CL, Telzer EH (2017) Magnetic resonance elastography for examining developmental changes in the mechanical properties of the brain. Dev Cogn Neurosci 33:176–181
Jucker M, Walker LC (2013) Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 501:45–51
Kaczmarek M, Subramaniam RP, Neff SR (1997) The hydromechanics of hydrocephalus: steady-state solutions for cylindrical geometry. Bull Math Biol 59(2):295–323
Kaliske M, Rothert H (1997) Formulation and implementation of three-dimensional viscoelasticity at small and finite strains. Comput Mech 19(3):228–239
Kansal A, Torquato S, Harsh G, Chiocca E, Deisboeck T (2000) Simulated brain tumor growth dynamics using a three-dimensional cellular automaton. J Theor Biol 203(4):367–382
Karimi A, Navidbakhsh M (2014) An experimental study on the mechanical properties of rat brain tissue using different stress-strain definitions. J Mater Sci Mater Med 25(7):1623–1630
Kaster T, Sack I, Samani A (2011) Measurement of the hyperelastic properties of ex vivo brain tissue slices. J Biomech 44(6):1158–1163
Kolias AG, Kirkpatrick PJ, Hutchinson P (2013) Decompressive craniectomy: past, present and future. Nat Rev Neurol 9:405–415
Koser DE, Moeendarbary E, Hanne J, Kuerten S, Franze K (2015) CNS cell distribution and axon orientation determine local spinal cord mechanical properties. Biophys J 108(9):2137–2147
Koser DE, Moeendarbary E, Kuerten S, Franze K (2018) Predicting local tissue mechanics using immunohistochemistry, p 358119. bioRxiv
Koser DE, Thompson AJ, Foster SK, Dwivedy A, Pillai EK, Sheridan GK, Svoboda H, Viana M, da F Costa L, Guck J et al (2016) Mechanosensing is critical for axon growth in the developing brain. Nat Neurosci 19(12):1592
Kruse SA, Rose GH, Glaser KJ, Manduca A, Felmlee JP, Jack CR, Ehman RL (2008) Magnetic resonance elastography of the brain. Neuroimage 39(1):231–237
Kuhl E, Maas R, Himpel G, Menzel A (2007) Computational modeling of arterial wall growth. Attempts towards patient-specific simulations based on computer tomography. Biomech Model Mechanobiol 6(5):321–331
Kyriacou SK, Mohamed A, Miller K, Neff S (2002) Brain mechanics for neurosurgery: modeling issues. Biomech Model Mechanobiol 1(2):151–164
Labus KM, Puttlitz CM (2016) Viscoelasticity of brain corpus callosum in biaxial tension. J Mech Phys Solids 96:591–604
Li X, von Holst H, Kleiven S (2011) Influence of gravity for optimal head positions in the treatment of head injury patients. Acta Neurochir 153(10):2057–2064
Lin S, Hung C (1979) Complications of cerebrospinal fluid shunts for hydrocephalus. A review of 204 cases. J Formos Med Assoc 78(3):258–266
Linninger AA, Sweetman B, Penn R (2009) Normal and hydrocephalic brain dynamics: the role of reduced cerebrospinal fluid reabsorption in ventricular enlargement. Ann Biomed Eng 37(7):1434–1447
Liu Y-L, Li G-Y, He P, Mao Z-Q, Cao Y (2017) Temperature-dependent elastic properties of brain tissues measured with the shear wave elastography method. J Mech Behav Biomed Mater 65:652–656
Liu Y-L, Liu D, Xu L, Su C, Li G-Y, Qian L-X, Cao Y (2018) In vivo and ex vivo elastic properties of brain tissues measured with ultrasound elastography. J Mech Behav Biomed Mater 83:120–125
Lu Y-B, Franze K, Seifert G, Steinhäuser C, Kirchhoff F, Wolburg H, Guck J, Janmey P, Wei E-Q, Käs J et al (2006) Viscoelastic properties of individual glial cells and neurons in the cns. Proc Natl Acad Sci U S A 103(47):17759–17764
MacManus D, Pierrat B, Murphy JG, Gilchrist M (2015) Dynamic mechanical properties of murine brain tissue using micro-indentation. J Biomech 48(12):3213–3218
MacManus DB, Murphy JG, Gilchrist MD (2018) Mechanical characterisation of brain tissue up to 35% strain at 1, 10, and 100/s using a custom-built micro-indentation apparatus. J Mech Behav Biomed Mater 87:256–266
MacManus DB, Pierrat B, Murphy JG, Gilchrist MD (2017) Region and species dependent mechanical properties of adolescent and young adult brain tissue. Sci Rep 7(1):13729
Maikos JT, Qian Z, Metaxas D, Shreiber DI (2008) Finite element analysis of spinal cord injury in the rat. J Neurotrauma 25(7):795–816
Majdan M, Plancikova D, Brazinova A, Rusnak M, Nieboer D, Feigin V, Maas A (2016) Epidemiology of traumatic brain injuries in europe: a cross-sectional analysis. Lancet Public Health 1(2):e76–e83
Mariappan YK, Glaser KJ, Ehman RL (2010) Magnetic resonance elastography: a review. Clin Anat 23(5):497–511
Markert B (2007) A constitutive approach to 3-d nonlinear fluid flow through finite deformable porous continua. Transp Porous Media 70(3):427
McCracken PJ, Manduca A, Felmlee J, Ehman RL (2005) Mechanical transient-based magnetic resonance elastography. Magn Reson Med 53(3):628–639
McIlvain G, Schwarb H, Cohen NJ, Telzer EH, Johnson CL (2018) Mechanical properties of the in vivo adolescent human brain. Dev Cogn Neurosci 34:27–33
Meaney DF, Morrison B, Bass CD (2014) The mechanics of traumaic brain injury: a review of what we know and what we need to know for reducing its societal burden. J Biomech Eng 136(2):021008
Menzel A, Kuhl E (2012) Frontiers in growth and remodeling. Mech Res Commun 42:1–14
Miller K (1999) Constitutive model of brain tissue suitable for finite element analysis of surgical procedures. J Biomech 32(5):531–537
Miller K (2011) Biomech Brain. Springer, Berlin
Miller K, Chinzei K (1997) Constitutive modelling of brain tissue: experiment and theory. J Biomech 30(11):1115–1121
Miller K, Chinzei K (2002) Mechanical properties of brain tissue in tension. J Biomech 35(4):483–490
Morrison PF, Laske DW, Bobo H, Oldfield EH, Dedrick RL (1994) High-flow microinfusion: tissue penetration and pharmacodynamics. Am J Physiol Regul Integr Comp Physiol 266(1):R292–R305
Murphy MC, Huston J III, Jack CR Jr, Glaser KJ, Senjem ML, Chen J, Manduca A, Felmlee JP, Ehman RL (2013) Measuring the characteristic topography of brain stiffness with magnetic resonance elastography. PLoS One 8(12):e81668
Murphy MC, Jones DT, Jack Jr CR, Glaser KJ, Senjem M L, Manduca A, Felmlee J P, Carter R E, Ehman R L, Huston III J (2016) Regional brain stiffness changes across the Alzhseimer’s disease spectrum. NeuroImage Clin 10:283–290
Nagashima T, Tamaki N, Matsumoto S, Horwitz B, Seguchi Y (1987) Biomechanics of hydrocephalus: a new theoretical model. Neurosurgery 21(6):898–904
Nicolle S, Lounis M, Willinger R (2004) Shear properties of brain tissue over a frequency range relevant for automotive impact situations: new experimental results. Stapp Car Crash J 48:239–258
Nie X, Sanborn B, Weerasooriya T, Chen W (2013) High-rate bulk and shear responses of bovine brain tissue. Int J Impact Eng 53:56–61
Ning X, Zhu Q, Lanir Y, Margulies SS (2006) A transversely isotropic viscoelastic constitutive equation for brainstem undergoing finite deformation. J Biomech Eng 128(6):925–933
Nordahl CW, Dierker D, Mostafavi I, Schumann CM, Rivera SM, Amaral DG, Van Essen DC (2007) Cortical folding abnormalities in autism revealed by surface-based morphometry. J Neurosci 27(43):11725–11735
Ogden R (1972) Large deformation isotropic elasticity-on the correlation of theory and experiment for incompressible rubberlike solids. Proc R Soc A 326(1567):565–584
Papazoglou S, Hirsch S, Braun J, Sack I (2012) Multifrequency inversion in magnetic resonance elastography. Phys Med Biol 57(8):2329
Pervin F, Chen WW (2009) Dynamic mechanical response of bovine gray matter and white matter brain tissues under compression. J Biomech 42(6):731–735
Pogoda K, Chin L, Georges PC, Byfield FJ, Bucki R, Kim R, Weaver M, Wells RG, Marcinkiewicz C, Janmey PA (2014) Compression stiffening of brain and its effect on mechanosensing by glioma cells. New J Phys 16(7):075002
Prange MT, Margulies SS (2002) Regional, directional, and age-dependent properties of the brain undergoing large deformation. J Biomech Eng 124(2):244–252
Prevost TP, Balakrishnan A, Suresh S, Socrate S (2011) Biomechanics of brain tissue. Acta Biomater 7:83–95
Qian L, Zhao H, Guo Y, Li Y, Zhou M, Yang L, Wang Z, Sun Y (2018) Influence of strain rate on indentation response of porcine brain. J Mech Behav Biomed Mater 82:210–217
Rashid B, Destrade M, Gilchrist MD (2012) Mechanical characterization of brain tissue in compression at dynamic strain rates. J Mech Behav Biomed Mater 10:23–38
Rashid B, Destrade M, Gilchrist MD (2012) Temperature effects on brain tissue in compression. J Mech Behav Biomed Mater 14:113–118
Rashid B, Destrade M, Gilchrist MD (2013) Mechanical characterization of brain tissue in simple shear at dynamic strain rates. J Mech Behav Biomed Mater 28:71–85
Rashid B, Destrade M, Gilchrist MD (2014) Mechanical characterization of brain tissue in tension at dynamic strain rates. J Mech Behav Biomed Mater 33:43–54
Razavi MJ, Zhang T, Liu T, Wang X (2015) Cortical folding pattern and its consistency induced by biological growth. Sci Rep 5:14477. https://doi.org/10.1038/srep14477
Reese S, Govindjee S (1998) A theory of finite viscoelasticity and numerical aspects. Int J Solids Struct 35(26–27):3455–3482
Richman DP, Stewart RM, Hutchinson J, Cavincss VS Jr (1975) Mechanical mode of brain convolutional deve lopment. Science 189:18–21
Rodriguez EK, Hoger A, McCulloch AD (1994) Stress-dependent finite growth in soft elastic tissues. J Biomech 27(4):455–467
Sack I, Beierbach B, Wuerfel J, Klatt D, Hamhaber U, Papazoglou S, Martus P, Braun J (2009) The impact of aging and gender on brain viscoelasticity. Neuroimage 46(3):652–657
Sack I, Streitberger K-J, Krefting D, Paul F, Braun J (2011) The influence of physiological aging and atrophy on brain viscoelastic properties in humans. PloS one 6(9):e23451
Samadi-Dooki A, Voyiadjis GZ, Stout RW (2018) A combined experimental, modeling, and computational approach to interpret the viscoelastic response of the white matter brain tissue during indentation. J Mech Behav Biomed Mater 77:24–33
Sarron J-C, Blondeau C, Guillaume A, Osmont D (2000) Identification of linear viscoelastic constitutive models. J Biomech 33(6):685–693
Schregel K, née Tysiak EW, Garteiser P, Gemeinhardt I, Prozorovski T, Aktas O, Merz H, Petersen D, Wuerfel J, Sinkus R (2012) Demyelination reduces brain parenchymal stiffness quantified in vivo by magnetic resonance elastography. Proc Natl Acad Sci U S A 109(17):6650–6655
Shreiber DI, Hao H, Elias RA (2009) Probing the influence of myelin and glia on the tensile properties of the spinal cord. Biomech Model Mechanobiol 8(4):311–321
Shuck L, Advani S (1972) Rheological response of human brain tissue in shear. J Basic Eng 94(4):905–911
Shulyakov AV, Cenkowski SS, Buist RJ, Del Bigio MR (2011) Age-dependence of intracranial viscoelastic properties in living rats. J Mech Behav Biomed Mater 4(3):484–497
Sidoroff F (1974) Nonlinear viscoelastic model with intermediate configuration. J Mec 13(4):679–713
Simo J (1992) Algorithms for static and dynamic multiplicative plasticity that preserve the classical return mapping schemes of the infinitesimal theory. Comput Methods Appl Mech Eng 99(1):61–112
Støverud KH, Darcis M, Helmig R, Hassanizadeh SM (2012) Modeling concentration distribution and deformation during convection-enhanced drug delivery into brain tissue. Transp Porous Media 92(1):119–143
Streitberger K-J, Sack I, Krefting D, Pfüller C, Braun J, Paul F, Wuerfel J (2012) Brain viscoelasticity alteration in chronic-progressive multiple sclerosis. PloS One 7(1):e29888
Tallinen T, Chung JY, Rousseau F, Girard N, Lefèvre J, Mahadevan L (2016) On the growth and form of cortical convolutions. Nat Phys 12(6):588–593
Tamura A, Hayashi S, Watanabe I, Nagayama K, Matsumoto T (2007) Mechanical characterization of brain tissue in high-rate compression. J Biomech Sci Eng 2(3):115–126
Tepole AB, Ploch CJ, Wong J, Gosain AK, Kuhl E (2011) Growing skin: a computational model for skin expansion in reconstructive surgery. J Mech Phys Solids 59(10):2177–2190
Thibault KL, Margulies SS (1998) Age-dependent material properties of the porcine cerebrum: effect on pediatric inertial head injury criteria. J Biomech 31(12):1119–1126
Tully B, Ventikos Y (2011) Cerebral water transport using multiple-network poroelastic theory: application to normal pressure hydrocephalus. J Fluid Mech 667:188–215
Tyler WJ (2012) The mechanobiology of brain function. Nat Rev Neurosci 13(12):867
Van Dommelen J, Van der Sande T, Hrapko M, Peters G (2010) Mechanical properties of brain tissue by indentation: interregional variation. J Mech Behav Biomed Mater 3:158–166
Vappou J, Breton E, Choquet P, Willinger R, Constantinesco A (2008) Assessment of in vivo and post-mortem mechanical behavior of brain tissue using magnetic resonance elastography. J Biomech 41(14):2954–2959
Velardi F, Fraternali F, Angelillo M (2006) Anisotropic constitutive equations and experimental tensile behavior of brain tissue. Biomech Model Mechanobiol 5(1):53–61
Verner S, Garikipati K (2013) Amyloid \(\beta\) deposition, neurodegeneration, and cognitive decline in sporadic Alzheimer’s disease: a prospective cohort study. Lancet Neurol 12:357–367
Voyiadjis GZ, Samadi-Dooki A (2018) Hyperelastic modeling of the human brain tissue: effects of no-slip boundary condition and compressibility on the uniaxial deformation. J Mech Behav Biomed Mater 83:63–78
Wang F, Han Y, Wang B, Peng Q, Huang X, Miller K, Wittek A (2018) Prediction of brain deformations and risk of traumatic brain injury due to closed-head impact: quantitative analysis of the effects of boundary conditions and brain tissue constitutive model. Biomech Model Mechanobiol 17(4):1165–1185
Wang X, Studholme C, Grigsby PL, Frias AE, Carlson VCC, Kroenke CD (2017) Folding, but not surface area expansion, is associated with cellular morphological maturation in the fetal cerebral cortex. J Neurosci 37(8):1971–1983
Weber G, Anand L (1990) Finite deformation constitutive equations and a time integration procedure for isotropic, hyperelastic-viscoplastic solids. Comput Methods Appl Mech Eng 79(2):173–202
Weickenmeier J, Butler C, Young P, Goriely A, Kuhl E (2017) The mechanics of decompressive craniectomy: perssonalized simulations. Comput Methods Appl Mech Eng 314:180–195
Weickenmeier J, de Rooij R, Budday S, Ovaert TC, Kuhl E (2017) The mechanical importance of myelination in the central nervous system. J Mech Behav Biomed Mater 76:119–124
Weickenmeier J, de Rooij R, Budday S, Steinmann P, Ovaert T, Kuhl E (2016) Brain stiffness increases with myelin content. Acta Biomater 42:265–272
Weickenmeier J, Jucker M, Kuhl E, Goriely A (2019) A physics-based model explains the prion-like features of neurodegeneration in Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis. J Mech Phys Solids 124:264–281
Weickenmeier J, Kuhl E, Goriely A (2018) Multiphysics of prion-like diseases: progression and atrophy. Phys Rev Lett 121:158101
Weickenmeier J, Kurt M, Ozkaya E, de Rooij R, Ovaert T, Ehman R, Butts Pauly K, Kuhl E (2018) Brain stiffens post mortem. J Mech Behav Biomed Master 84:88–98
Weickenmeier J, Kurt M, Ozkaya E, Wintermark M, Butts Pauly K, Kuhl E (2018) Magnetic resonance elastography of the brain: a comparison between pigs and humans. J Mech Behav Biomed Mater 77:702–710
Weickenmeier J, Saez P, Butler C, Young P, Goriely A, Kuhl E (2017) Bulging brains. J Elast 129(1–2):197–212
Welker W, Johnson IJ, Noe A (2014) Comparative mammalian brain collections. http://brainmuseum.org. Accessed 11 Oct 2014
Whittall KP, Mackay AL, Graeb DA, Nugent RA, Li DK, Paty DW (1997) In vivo measurement of t2 distributions and water contents in normal human brain. Magn Reson Med 37(1):34–43
Wright RM, Post A, Hoshizaki B, Ramesh KT (2013) A multiscale computational approach to estimating axonal damage under inertial loading of the head. J Neurotrauma 30(2):102–118
Wuerfel J, Paul F, Beierbach B, Hamhaber U, Klatt D, Papazoglou S, Zipp F, Martus P, Braun J, Sack I (2010) Mr-elastography reveals degradation of tissue integrity in multiple sclerosis. Neuroimage 49(3):2520–2525
Zhang W, Liu L-F, Xiong Y-J, Liu Y-F, Yu S-B, Wu C-W, Guo W (2018) Effect of in vitro storage duration on measured mechanical properties of brain tissue. Sci Rep 8(1):1247
Zhao W, Choate B, Ji S (2018) Material properties of the brain in injury-relevant conditions-experiments and computational modeling. J Mech Behav Biomed Mater 80:222–234
Acknowledgements
This study was kindly supported by the German Research Foundation grant STE 544/50, the Emerging Fields Initiative (EFI) by the FAU, and the STAEDTLER Stiftung to P.S. and S.B., as well as by the Emerging Talents Initiative (ETI) by the FAU and the German Research Foundation grant BU 3728/1-1 to S.B. We further greatly acknowledge the support by the National Science Foundation grant CMMI 1727268 “Understanding neurodegeneration across the scales”, the Stanford Bio-X IIP seed grant “Molecular mechanisms of chronic traumatic encephalopathy” and the Humboldt Research Award to E.K.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Budday, S., Ovaert, T.C., Holzapfel, G.A. et al. Fifty Shades of Brain: A Review on the Mechanical Testing and Modeling of Brain Tissue. Arch Computat Methods Eng 27, 1187–1230 (2020). https://doi.org/10.1007/s11831-019-09352-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11831-019-09352-w