Abstract
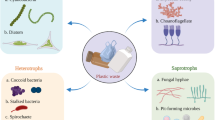
The daily use of plastics presents a serious pollution issue due to their extremely slow degradation. Microplastics and the biofilm that grows on plastics (i.e., the plastisphere) are important subsets of plastic wastes. Many studies have been conducted to reveal the structures of the plastispheres, the driving factors for the formation of the plastisphere, and the ability of the plastispheres to degrade plastics in a variety of water bodies. However, the plastispheres related to wastewater are understudied. In this study, we used a microcosmic strategy to study the evolution of the plastispheres associated with microplastics (MPs) over time in wastewater. We found that plastic materials and water sources did not actively select and shape the plastispheres at an early stage, but the active selection for a unique niche of the plastisphere occurred after 14 d of growth. In addition, we confirmed that the alkB gene was densely present, and metagenomics showed some additional chemical reactions, which suggests that MPs are consumed by the microbes in the plastispheres. Additionally, metagenomics identified some metagenome-assembled genomes (MAGs) associated with high-density polyethylene (HDPE) and polyethylene terephthalate (PET). The identification of HDPE-associated MAGs and PET-associated MAGs further supports the notion that the selection for a unique niche of the plastisphere is driven by plastic materials and water sources (in this study, after 14 d of growth). Our discoveries bring new views on the behavior of the wastewater-associated plastisphere, especially how long it takes a wastewater plastisphere to form.

Similar content being viewed by others
References
Aronesty E (2013). Comparison of sequencing utility programs. The Open Bioinformatics Journal, 7(1875–0362): 1–8
Bhagwat G, Zhu Q, O’Connor W, Subashchandrabose S, Grainge I, Knight R, Palanisami T (2021). Exploring the Composition and functions of plastic microbiome using whole-genome sequencing. Environmental Science & Technology, 55(8): 4899–4913
de Tender C A, Devriese L I, Haegeman A, Maes S, Ruttink T, Dawyndt P (2015). Bacterial community profiling of plastic litter in the belgian part of the North Sea. Environmental Science & Technology, 49(16): 9629–9638
Duan J, Bolan N, Li Y, Ding S, Atugoda T, Vithanage M, Sarkar B, Tsang D C W, Kirkham M B (2021). Weathering of microplastics and interaction with other coexisting constituents in terrestrial and aquatic environments. Water Research, 196: 117011
Eriksen M, Mason S, Wilson S, Box C, Zellers A, Edwards W, Farley H, Amato S (2013). Microplastic pollution in the surface waters of the Laurentian Great Lakes. Marine Pollution Bulletin, 77(1–2): 177–182
Estahbanati S, Fahrenfeld N L (2016). Influence of wastewater treatment plant discharges on microplastic concentrations in surface water. Chemosphere, 162: 277–284
Guo X P, Yang Y, Lu D P, Niu Z S, Feng J N, Chen Y R, Tou F Y, Garner E, Xu J, Liu M, Hochella M F JJr (2018). Biofilms as a sink for antibiotic resistance genes (ARGs) in the Yangtze Estuary. Water Research, 129: 277–286
Harrison J P, Schratzberger M, Sapp M, Osborn A M (2014). Rapid bacterial colonization of low-density polyethylene microplastics in coastal sediment microcosms. BMC Microbiology, 14(1): 232
Kang D D, Li F, Kirton E, Thomas A, Egan R, An H, Wang Z (2019). MetaBAT 2: An adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ, 7: e7359
Kazour M, Terki S, Rabhi K, Jemaa S, Khalaf G, Amara R (2019). Sources of microplastics pollution in the marine environment: Importance of wastewater treatment plant and coastal landfill. Marine Pollution Bulletin, 146: 608–618
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner F O (2013). Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Research, 41(1): e1
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6): 1547–1549
Lagarde F, Olivier O, Zanella M, Daniel P, Hiard S, Caruso A (2016). Microplastic interactions with freshwater microalgae: Hetero-aggregation and changes in plastic density appear strongly dependent on polymer type. Environmental Pollution (Barking, Essex: 1987), 215: 331–339
Lechner A, Keckeis H, Lumesberger-Loisl F, Zens B, Krusch R, Tritthart M, Glas M, Schludermann E (2014). The Danube so colourful: A potpourri of plastic litter outnumbers fish larvae in Europe’s second largest river. Environmental Pollution (Barking, Essex: 1987), 188(100): 177–181
Lee C, Kim J, Shin S G, Hwang S (2008). Monitoring bacterial and archaeal community shifts in a mesophilic anaerobic batch reactor treating a high-strength organic wastewater. FEMS Microbiology Ecology, 65(3): 544–554
Li X, Chen L, Mei Q, Dong B, Dai X, Ding G, Zeng E Y (2018). Microplastics in sewage sludge from the wastewater treatment plants in China. Water Research, 142: 75–85
Long A M, Hou S, Ignacio-Espinoza J C, Fuhrman J A (2021). Benchmarking microbial growth rate predictions from metagenomes. ISME Journal, 15(1): 183–195
Lu J, Zhang Y, Wu J, Luo Y (2019). Effects of microplastics on distribution of antibiotic resistance genes in recirculating aquaculture system. Ecotoxicology and Environmental Safety, 184: 109631
McCormick A, Hoellein T J, Mason S A, Schluep J, Kelly J J (2014). Microplastic is an abundant and distinct microbial habitat in an urban river. Environmental Science & Technology, 48(20): 11863–11871
Nauendorf A, Krause S, Bigalke N K, Gorb E V, Gorb S N, Haeckel M, Wahl M, Treude T (2016). Microbial colonization and degradation of polyethylene and biodegradable plastic bags in temperate fine-grained organic-rich marine sediments. Marine Pollution Bulletin, 103(1–2): 168–178
Oberbeckmann S, Osborn A M, Duhaime M B (2016). Microbes on a bottle: substrate, season and geography influence community composition of microbes colonizing marine plastic debris. PLoS One, 11(8): e0159289
Ogonowski M, Schür C, Jarsén Å, Gorokhova E (2016). The effects of natural and anthropogenic microparticles on individual fitness in daphnia magna. PLoS One, 11(5): e0155063
Pham D N, Clark L, Li M (2021). Microplastics as hubs enriching antibiotic-resistant bacteria and pathogens in municipal activated sludge. Journal of Hazardous Materials Letters, 2: 100014
Pinnell L J, Turner J W (2019). Shotgun metagenomics reveals the benthic microbial community response to plastic and bioplastic in a coastal marine environment. Frontiers in Microbiology, 10: 1252
Pinto M, Polania Zenner P, Langer T M, Harrison J, Simon M, Varela M M, Herndl G J (2020). Putative degraders of low-density polyethylene-derived compounds are ubiquitous members of plastic-associated bacterial communities in the marine environment. Environmental Microbiology, 22(11): 4779–4793
Roux S, Enault F, Bronner G, Debroas D (2011). Comparison of 16S rRNA and protein-coding genes as molecular markers for assessing microbial diversity (Bacteria and Archaea) in ecosystems. FEMS Microbiology Ecology, 78(3): 617–628
Seemann T (2014). Prokka: rapid prokaryotic genome annotation. Bioinformatics (Oxford, England), 30(14): 2068–2069
Son H F, Cho I J, Joo S, Seo H, Sagong H Y, Choi S Y, Lee S Y, Kim K J (2019). Rational protein engineering of thermo-stable PETase from ideonella sakaiensis for highly efficient PET degradation. ACS Catalysis, 9(4): 3519–3526
Toyofuku M, Inaba T, Kiyokawa T, Obana N, Yawata Y, Nomura N (2016). Environmental factors that shape biofilm formation. Bioscience, Biotechnology, and Biochemistry, 80(1): 7–12
Wu X, Pan J, Li M, Li Y, Bartlam M, Wang Y (2019). Selective enrichment of bacterial pathogens by microplastic biofilm. Water Research, 165: 114979
Wu Y, Simmons B A, Singer S W (2016). MaxBin 2.0: An automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics (Oxford, England), 32(4): 605–607
Yin X, Jiang X, Chai B, Li L, Yang Y, Cole J R, Tiedje J M, Zhang T (2018). ARGs-OAP v2.0 with an expanded SARG database and Hidden Markov Models for enhancement characterization and quantification of antibiotic resistance genes in environmental metagenomes. Bioinformatics (Oxford, England), 34(13): 2263–2270
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K (2016). A bacterium that degrades and assimilates poly(ethylene terephthalate). Science, 351(6278): 1196–1199
Yuan B, McLachlan M S, Roos A M, Simon M, Strid A, de Wit C A (2021). Long-chain chlorinated paraffins have reached the arctic. Environmental Science & Technology Letters, 8(9): 753–759
Zeng L, Li H, Wang T, Gao Y, Xiao K, Du Y, Wang Y, Jiang G (2013). Behavior, fate, and mass loading of short chain chlorinated paraffins in an advanced municipal sewage treatment plant. Environmental Science & Technology, 47(2): 732–740
Zeng L, Wang T, Ruan T, Liu Q, Wang Y, Jiang G (2012). Levels and distribution patterns of short chain chlorinated paraffins in sewage sludge of wastewater treatment plants in China. Environmental Pollution, 160(1): 88–94
Zettler E R, Mincer T J, Amaral-Zettler L A (2013). Life in the “plastisphere”: Microbial communities on plastic marine debris. Environmental Science & Technology, 47(13): 7137–7146
Zhao Z, Baltar F, Herndl G J (2020). Linking extracellular enzymes to phylogeny indicates a predominantly particle-associated lifestyle of deep-sea prokaryotes. Science Advances, 6(16): eaaz4354
Acknowledgements
Our research was funded by the National Key Research and Development Program of China (No. 2021YFA1202500), the National Natural Science Foundation of China (No. 42177357), the Guangdong Basic and Applied Research Foundation (China) (No. 2021A1515012191), and the Shenzhen Science and Technology Innovation Committee (China) (No. JCYJ20210324104412033). We thank the Center for Computational Science and Engineering at the Southern University of Science and Technology (China) (SUSTech) and the core research facilities at SUSTech for providing excellent resources and services. Moreover, we thank Dr. Chuanlun Zhang from the Department of Ocean Science and Engineering at SUSTech for lending QuantStudio qPCR.
Author information
Authors and Affiliations
Corresponding author
Additional information
Highlights
• Wastewater MPs exhibited resistomes and therefore health threats.
• High density of alkB gene indicates both HDPE and PET can be utilized by microbes.
• Plastics and waters actively selected and shaped the plastispheres over time.
• A broader phylogenetic spectrum of MHET-degrading microorganisms was annotated.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Xia, Y., Zhang, X., Zhang, M. et al. Plastic materials and water sources actively select and shape wastewater plastispheres over time. Front. Environ. Sci. Eng. 16, 145 (2022). https://doi.org/10.1007/s11783-022-1580-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11783-022-1580-1