Abstract
Baculoviruses are successful microbial control agents used in the biological control of agricultural pest species, especially in the order Lepidoptera. The beet armyworm, Spodoptera exigua is a popular agricultural pest in the world. S exigua larvae, which are active in the all-summer period, cause economic losses by damaging many crops in agricultural production areas. This article aims to analyze the full genome of Spodoptera exigua multiple nucleopolyhedroviruses from Iran (SeMNPV-IR) and to determine the geographical difference between the strains at the genomic level. The full genome of SeMNPV-IR is 135.764 base pairs in length that contained 136 open reading frames (ORFs), and 43.92% G + C content. The seven homologous repeated (hr) regions were identified. In the results of genome-wide phylogenetic analysis, it was determined that the SeMNPV-IR genome isolated from Iran was interestingly close to the genome of the US and Korea isolates. However, there are significant differences in the two hypothetical (Orf 83 and Orf 104) genes. The SeMNPV-IR has a unique homolog repeat region (hr1, 96 bp) that is not found in other SeMNPV genomes, and it also differs in terms of the hr2 region. In silico restriction endonuclease analysis by StuI and SacII enzymes show that there were significant differences between all geographic isolates of SeMNPV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Baculoviruses have a positive-stranded double-stranded DNA genome and are one of the most successful insect viruses that have been used in biological control for many years (Herniou et al. 2011). They are an environmentally friendly microbial agent that does not harm any living thing other than insects and therefore can be used safely in biological control (Krieg et al. 1980; Entwistle et al. 1983). In addition, the use of baculoviruses as an expression vector in the production of recombinant protein for the diagnosis of diseases and vaccine development with bacmid (bacterial artificial chromosomes containing the baculovirus genome) technology is one of its most important features (O’Reilly et al. 1992; Miller et al. 1998).
Baculoviruses are subject to various classifications according to both their nucleo-capsid structure and the host group they infect. Baculoviruses with polyhedrin protein are called nucleopolyhedrovirus, and those with granulin protein are called granuloviruses. Nucleopolyhedroviruses are determined to have a single or multiple nucleocapsid structure as a result of transmission electron microscopy examinations (Ackermann and Smirnoff 1983). Although the majority of baculoviruses infect the Lepidoptera group (Alphabaculoviruses and Betabaculoviruses), there are also isolates that infect Hymenoptera (Gamabaculoviruses) and Diptera (Deltabaculoviruses) (Jehle et al. 2006).
In recent years, molecular comparison of baculovirus isolates has been done at the whole genome level. In particular, genomic differences between baculovirus isolates obtained from dissimilar geographical regions are understood by next-generation sequencing and bioinformatics analysis. Thus, detailed information about the genome features of baculoviruses was obtained and it was determined that they contain approximately 900 different genes (Ferrelli et al. 2012). There are 38 conserved genes common to all baculoviruses (Garavaglia et al. 2012; Javed et al. 2017). However, other genes and repeat regions in the genome cause significant differences between isolates. In particular, geographic variants isolated from the same insect have different virulence and host spectrum. The reason for these differences will only be revealed by performing genome analyses and comparing the isolates with each other at the genome level. In addition, genome analyses of baculoviruses will enable the identification of new baculovirus genes that were not known until now (Van Oers et al. 2005).
Multiple genotypic variants have emerged in baculoviruses, even in individually infected larvae. Restriction fragment length polymorphism analysis (RFLP), which has been used since old times, is a very convenient method for the detection of geographic variation (Erlandson 2009). In addition, the emergence of new, reliable, and inexpensive DNA sequencing strategies that have emerged in recent years has allowed these assays to be performed more effortlessly and more reliably for the efficient detection and characterization of genotypic variants in and within geographic and temporal isolates of baculovirus species in silico. The results of the comparative analysis of geographic strains showed that especially homologous repeat regions and baculoviral repeat origin genes cause potential recombination events that cause genetic variability (Eroglu et al. 2020).
Spodoptera exigua is an important agricultural pest that originates from Southeast Asia and has spread all over the world thanks to its high migration-adaptation ability (Xia-Ling et al. 2011). During the larval period, polyphagous feed causes great economic loss in more than 50 plant species. It especially prefers important cultural plants such as sugar beet, corn, cotton, lettuce, sunflower, and tomato. If these larvae, which consume all the leaves until only the veins remain in the plants, are not combated, the product loss reaches 100% (Smits et al. 1987; Eroglu 2022).
In our previous studies, a baculovirus isolate was detected as a result of the examination of S. exigua larvae collected from sugar beet cultivation areas in Iranian (Darsouei et al. 2017). Whole genome analysis of S. exigua baculovirus isolates previously obtained only from the USA, Spain, UK, Korea, and China was performed. However, SeMNPV-IR isolated a very different locality (Razavi Khorasan region located between Southwest Asia and Central Asia) from other SpexNPV strains for which complete genome analysis was performed. This study, it was aimed to perform a whole genome analysis of this different geographical isolate (SeMNPV-IR) and to compare it with other all Spodoptera nucleopolyhedrovirus genomes in the database.
Materials and methods
Virus source and DNA extraction
The virus isolate was obtained from Spodoptera exigua larvae found in sugar beet plantations in Mashhad, Iran between 2014 and 2015 (Darsouei et al. 2017). After the infected larvae were collected from the field by Darsouei et al. and brought to the laboratory, virus production was carried out in healthy S. exigua larvae. Infected larvae were purified by the classical cheesecloth and sucrose gradient method (Munoz et al. 1997; Eroglu et al. 2018, 2019). The polyhedral inclusion bodies (PIB) need to be removed before proceeding with the DNA extraction. So, firstly 300 µl of pure virus suspension (1 × 108 PIB/mL) and 300 µl 3 × DAS buffer (0.3 M Na2CO3, 0.5 M NaCl, 0.03 M EDTA; pH 10.5) were mixed and rotated for 30 min to remove the virus particles from PIBs. Then, centrifuged at 20.300 g, 25℃ for 30 min, and removed the supernatant. DNA extraction was performed from the naked virus solution according to the instructions of the EcoPURE Genomic DNA kit. The DNA concentration measured by nanodrop was 68 ng/µl and the purity ratio A260/280 ≥ 1.8.
Genomic DNA sequencing and genome assembly
Genomic DNA was sequenced at MG Bioinformatic company using Illumina NovaSeq generates raw images utilizing (NovaSeq 2 × 150 PE) for system control. First, raw sequence reads were checked using the FastQC program (Andrews 2010). Since the sequencing results were of the quality and depth to form mitogenome sequences, the extraction of low-quality-score sequences from the data was not performed. More than 90% of the sequences in all sequence reads meet the Q30 criterion (error rate 0.001). No positional deviation was observed between sequence reads in terms of nucleotide content. Two different methods were used together to select and assemble the sequences of the genome from the high-volume sequence information of which sequencing was performed de novo and preliminary quality analysis was performed. First, raw sequence reads were checked using the FastQC program. Since the sequencing results were of the quality and depth to form genome sequences, the extraction of low-quality-score sequences from the data was not performed. After that, NOVOPlasty 2.7.2 (Dierckxsens et al. 2017) algorithms, which are used to assemble the sequences of the genome in all data, were run in the TRUBA computer cluster. Both methods were successful in acquiring genome data de novo, resulting in almost identical genome sequences. The genome obtained using these two tools was transferred to the Geneious Prime (Biomatters Ltd.) program and the accuracy of the genome obtained using the “map read to reference” feature on the raw sequences was checked.
Genome annotation and genomic relationship analyzes
Accurate determination of gene boundaries (annotation) of the genome formed by combining contigs is required for further analysis. Annotation is the process of defining the start, ending, and transcription chains (heavy or light) of gene regions, the locations of repetitive regions, and structural features such as the origin of transcription and replication.
In genome annotation of SeMNPV-IR, the detection of all genes and homologous repeat sequences in the genome was performed using the Benchling Biology Software (https://benchling.com) as described in our previous study (Eroglu et al. 2020). The SeMNPV (Accession number: AF169823), the first Spodoptera exigua nucleopolyhedrovirus whose genome was analyzed, was chosen as the reference genome (Ijkel et al. 1999). The amino acid sequences encoded by 38 core genes concerning all Spodoptera nucleopolyhedrovirus genomes and SeMNPV-IR genome were aligned in BioEdit (7.1.3.0). In phylogenetic analysis, the Jones-Taylor-Thornton (JTT) model with 1000 bootstrap in the Maximum Likelihood method to generate a phylogeny was used by the MEGA11 program. The restriction endonuclease profile of the SeMNPV-IR was compared to other Spodoptera exigua NPV genomes (SeMNPV, SeMNPV-K1, SeMNPV VT-SeAl2, and SeMNPV VT-SeOx4) in terms of StuI and SacII enzymes as in silico by using the Benchling tool (https://benchling.com).
Results
Genome organization of SeMNPV-IR
The whole genome of the SeMNPV-IR isolate was sequenced and registered at NCBI (Accession number: OP562161). The genome size of SeMNPV-IR was detected to be 135.764 kb. There are a total of 136 ORFs in the genome, of which 75 are clockwise and 61 are counterclockwise with 43.92% GC content. All ORFs of SeMPN-IR were compared to reference baculovirus genomes (Autographa californica MNPV, Cydia pomonella GV, and Spodoptera exigua MNPV) and close to the relationship as per phylogenetic analysis (SeMPNV-K1, VTSe-Ox4, and VTSe-Al2). The location of ORFs in the SeMPV-IR genome with the genomes indicated in Table 1 and their amino acid similarity ratios are given. In addition, 38 core genes found in baculoviruses are indicated by light blue color in Table 1.
ORFs contents
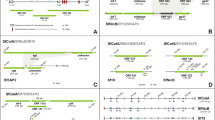
The ORFs in the genome of baculoviruses are divided into seven main classes according to their replication, oral infectivity, transcription, apoptosis, auxiliary, structural, and unknown functions (Herniou et al. 2003). In the genome map of the SeMNPV-IR isolate detailed in Fig. 1, the functions of the genes are represented by different colors.
Eleven genes are responsible for replication (me-53, helicase, i.e.-1, dutpase, dnapol, dbp, lef-1, lef-2, lef-3, lef-7, and lef-11), twelve genes for transcription (lef-4, lef-5, lef-6, lef-8, lef-9, lef-10, met, vlf-1, p47, 39k, i.e.-0, and pkip-1), nineteen genes for structural (polyhedrin, p10, odv-ec27, odv-e18, gp-41, vp-39, p78, p6.9, pk-1, fusion protein, p24, gp16, gp37, calyx, odv-e25, odv-e28, two odv-e66 and vp1034), nine genes for auxiliary (cathepsin, chitinase, egt, alk-exo, fgf, sod, ptp-2, arif-1 and ubiqutin), ten genes for oral infectivity (pif0 -pif9), three genes for apoptosis (iap-2 and two iap-3), thirty-two genes for unknown and forty genes for hypothetical. Baculovirus repeat (bro) genes are absent in the SeMNPV-IR genome as in other SeMNPV genomes.
In addition to protein-coding genes, homologous repeat (hr) sequences in the genome are also very common in baculoviruses. These sequences, which can be in different numbers in the genome, are generally responsible for increased gene expression and DNA replication (Cochran et al. 1982; Kool et al. 1995). SeMNPV-IR has 7 repeat regions in its genome, ranging in length from 96 to 806 base pairs (Table 1).
Genomic relationship analyzes
In order to determine the phylogenetic closeness of SeMNPV genomes isolated from different geographical regions, the amino acid sequences of 38 core genes belonging to the genomes were obtained from the NCBI database. The localities and access numbers from which the genomes used in this analysis were obtained are given in the S1 table. As a result of the analysis, the SeMNPV isolate obtained from the Mashhad region of Iran showed akin to the SeMNPV genomes isolated from the USA and Korea (Fig. 2).
In silico Restriction Fragment Length Polymorphism (RFLP).
To compare the genomes of geographic SeMNPV isolates, in silico analysis was performed using the Benchling online tool. Digestion of the SeMNPV-IR genome with the StuI and SacII restriction endonucleases produced 6 and 11 fragments, respectively. The RFLP results showed that the genome of the Iranian isolate differed from the USA, UK, Spain, and Korea isolates (Fig. 3; Table 2). RFLP profiles typically differ among geographic isolates of baculoviruses. These differences are usually due to the presence or absence of non-protein-coding repeat sequences found in the genome (Smith and Croock, 1988; Munoz et al. 1999).
Discussion
The new generation sequencing methods that emerged after the completion of the human genome project allowed the genomes of many organisms to be elucidated. Viruses are organisms on the fringes of life that differentiate quite easily at the genome level and can even have great variation among geographic isolates of the same virus. Variations in baculoviruses between different geographic isolates from the same insect have been a topic of interest for us virologists for many years. The fact that these differences can be easily analyzed at the genome level today has enabled the studies to be carried to advanced dimensions. Most studies of geographic genetic diversity among baculoviruses have been done for Alphabaculoviruses that infect lepidopteran larval populations (Erlandson 2009). Comparison of geographic isolates and band differences seen in RFLP often reveal differences in virulence versus native and alternative hosts (Laviña-Caoili et al. 2001).
Spodoptera exigua larvae are a popular agricultural pest that damages many agricultural crops and causes economic loss all over the world (Lasa et al. 2007). Although many studies have been conducted on the use of baculoviruses in the control of the pest (Takatsuka et al. 2003; Nathan and Kalaivani 2006; Widiawati et al. 2021), the search for isolates with local, higher virulence and more effectiveness is still continuing. Kondo et al. (1994) after examining 11 nucleopolyhedrovirus-infected S. exigua larvae collected from Shiga, Japan divided the isolates into two groups according to the shape difference seen in both polyhedra (cubic and icosahedral). In addition, the RFLP results supported that the isolates should be divided into two groups. They reported that while the RFLP results of the isolates in the first group were similar to Authographa californica NPV, the bands of the isolates in the second group were similar to Spodoptera exigua NPV California isolates. Murillo et al. (2006) isolated seven S. exigua NPVs with both phenotypic and genotypic variation from greenhouse fields in Spain. As a result of polymerase chain reactions, it has been shown that a single genotype is dominant in some isolates and two or three different genotypes are mixed in others. They have been reported that this situation affects both the killing rate and the pathogenicity of the isolates. There are general differences between geographic isolates of baculoviruses in terms of host spectrum, virulence rate, and mortality rate on larvae (Allaway and Payne 1984). These differences are undoubtedly due to the variations found in the genome (Cory et al. 2005). To date, genome analysis of SeMNPV isolates from the USA, Korea, UK, Spain, and China has been performed (Ijkel et al. 1999; Theze et al. 2014). In this study, the genome of SeMNPV-IR isolated from Razavi Khorasan Province of Mashhad in eastern Iran was analyzed. The region is an interesting geographical region located in the middle of three countries which countries Iran, Turkmenistan, and Afghanistan. In this study, genomic features of SeMNPV-IR were described in detail and compared with other Spodoptera exigua NPV genomes (SeMNPV, SeMNPV-K1, SeMNPV VT-SeAl2, and SeMNPV VT-SeOx4). The SeMNPV-IR genome has 40 hypothetical genes and most of these hypothetical genes are also found in other SeMNPV genomes as homologs. However, two of them (Orf 83 and Orf 104) are not found in UK and Spain isolates (Theze et al. 2014), while they are present in the genomes of USA (Ijkel et al. 1999) and Korea isolates (unpublished) (Table 1). The similarity rate between Orf 83 in the SeMNPV-IR genome and the homologous gene found in the genome of the USA and Korea isolates is 98%, while this rate is 89% for Orf 104.
Homologous repeat regions in baculoviruses are responsible both transcriptional enhancers and for progeny virus production and are regions that vary widely among isolates (McClintock and Dougherty 1988; Sun 2015). In these regions, the A + T ratio is quite high, and some baculoviruses may be absent while may be high numbers in some baculovirus genomes (Luque et al. 2001; Wang et al. 2016). The SeMNPV-IR has 7 homolog repeat regions (hr1-hr7). Hr1 (96 bp) is not found in other SeMNPV genomes. Besides, the base length of hr2 (355 bp) is considerably shorter than that of other SeMNPVs (1131 and 1347 bp), with a similarity rate of 92% (Table 3).
Phylogenetic relationship analysis demonstrated that the Iran isolate of SeMNPV clustered near the isolates from USA and Korea.
The genomes of SeMNPV of Iran, USA, Korea, UK, and Spain isolates were digested in silico with StuI and SacII enzymes (Fig. 3; Table 3). While there are 6, 8, 7, 7, and 7 cut regions for the StuI enzyme in genomes, there are 11, 12, 12, 12, and 12 cut regions for the SacII enzyme, respectively. In terms of both enzymes, it was Iran isolate that had the least cut sites.
As a result of the comparison of all protein-coding regions and homologous repeat regions in the genome, significant differences were observed between SeMNPV genomes, especially in two hypothetical regions (Orf 83, Orf 104) and two homologous repeat regions (hr1, hr2). Genomic variation among geographic variants from the same insect species isolated is thought to be largely due to hr regions.
In this study, whole genome analysis of SeMNPV-IR was performed. The elucidation of the function of two hypothetical genes in the genome of this isolate, whose genome is in a very different location (Razavi Khorasan) from the previously sequenced SeMNPV samples, provides a basis for further investigation. The clarification of the functions of these genes by conducting different studies will contribute greatly to the known data about the genome of baculoviruses.
References
Ackermann HW, Smirnoff WA (1983) A morphological investigation of 23 baculoviruses. J Invertebr Pathol 41:269–280. https://doi.org/10.1016/0022-2011(83)90244-6
Allaway GP, Payne CC (1984) Host range and virulence of five baculoviruses from lepidopterous hosts. Ann Appl Biol 105:29–37
Andrews S (2010) FastQC-A Quality Control application for FastQ files. Babraham, UK, Babraham Bioinformatics
Barrera GP, Belaich MN, Patarroyo MA, Villamizar LF, Ghiringhelli PD (2015) Evidence of recent interspecies horizontal gene transfer regarding nucleopolyhedrovirus infection of Spodoptera frugiperda. BMC Genomics 16:1–18. https://doi.org/10.1186/s12864-015-2218-5
Ben Tiba S, Laarif A, Wennmann JT, Bouslama T, Jehle JA (2022) Biological activity and genome composition of a tunisian isolate of Spodoptera littoralis nucleopolyhedrovirus (SpliNPV-Tun2). Egypt J Biol Pest Control 32:1–15. https://doi.org/10.1186/s41938-022-00568-w
Breitenbach JE, El-Sheikh ESA, Harrison RL, Rowley DL, Sparks ME, Gundersen-Rindal DE, Popham HJ (2013) Determination and analysis of the genome sequence of Spodoptera littoralis multiple nucleopolyhedrovirus. Virus Res 171:194–208. https://doi.org/10.1016/j.virusres.2012.11.016
Chen Y, Qi B, Zheng G, Zhang Y, Deng F, Wan F, Li C (2019) Identification and genomic sequence analysis of a new Spodoptera exigua multiple nucleopolyhedrovirus, SeMNPV-QD, isolated from Qingdao, China. J Invertebr Pathol 160:8–17. https://doi.org/10.1016/j.jip.2018.11.006
Cochran MA, Carstens EB, Eaton BT, Faulkner P (1982) Molecular cloning and physical mapping of restriction endonuclease fragments of Autographa californica nuclear polyhedrosis virus DNA. J Virol 41:940–946 pmid:16789219
Cory JS, Green BM, Paul RK, Hunter-Fujita F (2005) Genotypic and phenotypic diversity of a baculovirus population within an individual insect host. J Invertebr Pathol 89:101–111. https://doi.org/10.1016/j.jip.2005.03.008
Entwistle PF, Adams PHW, Evans HF, Rivers CF (1983) Epizootiology of a nuclear polyhedrosis virus (Baculoviridae) in European spruce sawfly (Gilpinia hercyniae): spread of disease from small epicentres in comparison with spread of baculovirus diseases in other hosts.Journal of Applied ecology473–487
Darsouei R, Karimi J, Hosseini M, Ghadamyari M (2017) Immune defence components of spodoptera exigua larvae against entomopathogenic nematodes and symbiotic bacteria. Biocontrol Sci Technol 27:867–885. https://doi.org/10.1080/09583157.2017.1350632
Dierckxsens N, Mardulyn P, Smits G (2017) Nucleic Acids Res 45:e18–e18. https://doi.org/10.1093/nar/gkw955. NOVOPlasty: de novo assembly of organelle genomes from whole genome data
Erlandson MA (2009) Genetic variation in field populations of baculoviruses: mechanisms for generating variation and its potential role in baculovirus epizootiology. Virol Sin 24:458–469. https://doi.org/10.1007/s12250-009-3052-1
Eroglu GB, Demir I, Demirbag Z (2018) A novel alphabaculovirus isolated from the cotton bollworm, Helicoverpa armigera (hubner) (Lepidoptera: Noctuidae): characterization and pathogenicity. Biologia 73:545–551. https://doi.org/10.2478/s11756-018-0053-2
Eroglu GB, Nalcacioglu R, Demirbag Z (2019) A new helicoverpa armigera nucleopolyhedrovirus isolate from Heliothis peltigera (Denis & Schiffermuller) (Lepidoptera: Noctuidae) in Turkey. Turkish J Biology 43(5):340–348. https://doi.org/10.3906/biy-1902-64
Eroglu GB, Inan C, Nalcacioglu R, Demirbag Z (2020) Genome sequence analysis of a Helicoverpa armigera single nucleopolyhedrovirus (HearNPV-TR) isolated from Heliothis peltigera in Turkey. PLoS ONE 15:e0234635. https://doi.org/10.1371/journal.pone.0234635
Eroglu GB (2022) Pest Insects and Their Biological Control. In: Abdurakhmonov IY (editör). Cotton [Internet]. London: IntechOpen; [cited 2023 Jan 17]. Available from: https://www.intechopen.com/chapters/80016 doi: https://doi.org/10.5772/intechopen.101980
Escasa SR, Harrison RL, Mowery JD, Bauchan GR, Cory JS (2019) The complete genome sequence of an alphabaculovirus from Spodoptera exempta, an agricultural pest of major economic significance in Africa. PLoS ONE 14:e0209937. https://doi.org/10.1371/journal.pone.0209937
Ferrelli ML, Berretta MF, Belaich MN, Ghiringhelli PD, Sciocco-Cap A, Romanowski V (2012) In: Garcia ML, Romanowsky V, editors. The baculoviral genome. Yugoslavia
Garavaglia MJ, Miele SAB, Iserte JA, Belaich MN, Ghiringhelli PD (2012) The ac53, ac78, ac101 and ac103 are newly discovered core genes in the family Baculoviridae. J Virol 86:12069–12079
Harrison RL, Puttler B, Popham HJ (2008) Genomic sequence analysis of a fast-killing isolate of Spodoptera frugiperda multiple nucleopolyhedrovirus. J Gen Virol 89:775–790. https://doi.org/10.1099/vir.0.83566-0
Harrison RL, Rowley DL (2019) Complete genome sequence of an alphabaculovirus from the southern armyworm, Spodoptera eridania. Microbiol Resource Announcements 8:e01277–e01218. https://doi.org/10.1128/MRA.01277-18
Herniou EA, Olszewski JA, Cory JS, O’Reilly DR (2003) The genome sequence and evolution of baculoviruses. Ann Rev Entomol 48:211. https://doi.org/10.1146/annurev.ento.48.091801.112756
Herniou EA, Arif BM, Becnel JJ, Blissard GW, Bonning B, Harrison R, Vlak JM (2011) Baculoviridae. Virus taxonomy: ninth report of the International Committee on Taxonomy of Viruses 163–173
Kondo A, Yamamoto M, Takashi S, Maeda S (1994) Isolation and characterization of nuclear polyhedrosis viruses from the beet armyworm Spodoptera exigua (Lepidoptera: Noctuidae) found in Shiga, Japan. Appl Entomol Zool 29(1):105–111. https://doi.org/10.1303/aez.29.105
IJkel WFJ, Van Strien EA, Heldens JGM, Broer R, Zuidema D, Goldbach RW, Vlak JM (1999) Sequence and organization of the Spodoptera exigua multicapsid nucleopolyhedrovirus genome. J Gen Virol 80:3289–3304. https://doi.org/10.1099/0022-1317-80-12-3289
Javed MA, Biswas S, Willis LG, Harris S, Pritchard C, Van Oers MM, Donly BC, Erlandson MA, Hegedus DD, Theilmann DA (2017) Autographa californica multiple nucleopolyhedrovirus ac83 is a per os infectivity factor (PIF) protein required for occlusion-derived virus (ODV) and budded virus nucleocapsid assembly as well as assembly of the PIF complex in ODV envelopes. J Virol 91:e02115–02116. https://doi.org/10.1128/JVI.02115-16
Jehle JA, Blissard GW, Bonning BC, Cory JS, Herniou EA, Rohrmann GF et al (2006) On the classification and nomenclature of baculoviruses: a proposal for revision. Arch Virol 151:1257–1266. https://doi.org/10.1007/s00705-006-0763-6
Kool M, Ahrens CH, Vlak JM, Rohrmann GF (1995) Replication of baculovirus DNA. J Gen Virol 76:2103–2118 pmid:7561748
Krieg A, Franz JM, Gröner A, Huber J, Miltenburger HG (1980) Safety of entomopathogenic viruses for control of insect pests. Environ Conserv 7:158–160
Lasa R, Pagola I, Ibanez I, Belda JE, Williams T, Caballero P (2007) Efficacy of Spodoptera exigua multiple nucleopolyhedrovirus as a biological insecticide for beet armyworm control in greenhouses of southern Spain. Biocontrol Sci Technol 17:221–232. https://doi.org/10.1080/09583150701211335
Laviña-Caoili BA, Kamiya K, Kawamura S, Ikeda M, Kobayashi M (2001) Comparative in vitro analysis of geographic variants of nucleopolyhedrovirus of Spodoptera litura isolated from China and the Philippines. J Insect Biotechnol Sericology 70(3):199–209. https://doi.org/10.11416/jibs2001.70.199
Luque T, Finch R, Crook N, O’Reilly DR, Winstanley D (2001) The complete sequence of the Cydia pomonella granulovirus genome. J Gen Virol 82:2531–2547. https://doi.org/10.1099/0022-1317-82-10-2531
McClintock JT, Dougherty EM (1988) Restriction mapping of Lymantria dispar nuclear polyhedrosis virus DNA: localization of the polyhedrin gene and identification of four homologous regions. J Gen Virol 69:2303–2312 pmid:2842437
Miller LK, Kaiser WJ, Seshagiri S (1998) Baculovirus regulation of apoptosis. Seminars in Virology 8:445–452
Munoz D, Vlak JM, Caballero P (1997) In vivo recombination between two strains of the genus nucleopolyhedrovirus in its natural host, Spodoptera exigua. Appl Environ Microbiol 63:3025–3031 pmid:9251191. https://doi.org/10.1128/aem.63.8.3025-3031.1997
Munoz D, Murillo R, Krell PJ, Vlak JM, Caballero P (1999) Four genotypic variants of a Spodoptera exigua nucleopolyhedrovirus (Se-SP2) are distinguishable by a hypervariable genomic region. Virus Res 59:61–74 pmid:10854166. https://doi.org/10.1016/S0168-1702(98)00125-7
Murillo R, Elvira S, Muñoz D, Wiliams T, Caballero P (2006) Genetic and phenotypic variability in Spodoptera exigua nucleopolyhedrovirus isolates from greenhouse soils in southern Spain. Biol Control 38:157–165. https://doi.org/10.1016/j.biocontrol.2005.12.001
Nathan SS, Kalaivani K (2006) Combined effects of azadirachtin and nucleopolyhedrovirus (SpltNPV) on Spodoptera litura Fabricius (Lepidoptera: Noctuidae) larvae. Biol Control 39:96–104. https://doi.org/10.1016/j.biocontrol.2006.06.013
O’Reilly DR, Miller LK, Luckow VA (1992) Baculovirus expression vectors, Laboratory Manual. Oxford University Press, New York
Pang Y, Yu J, Wang L, Hu X, Bao W, Li G, Yang H (2001) Sequence analysis of the Spodoptera litura multicapsid nucleopolyhedrovirus genome. Virology 287:391–404. https://doi.org/10.1006/viro.2001.1056
Simon O, Palma L, Beperet I, Munoz D, Lopez-Ferber M, Caballero P, Williams T (2011) Sequence comparison between three geographically distinct Spodoptera frugiperda multiple nucleopolyhedrovirus isolates: detecting positively selected genes. J Invertebr Pathol 107:33–42. https://doi.org/10.1016/j.jip.2011.01.002
Simon O, Palma L, Williams T, Lopez-Ferber M, Caballero P (2012) Analysis of a naturally-occurring deletion mutant of Spodoptera frugiperda multiple nucleopolyhedrovirus reveals sf58 as a new per os infectivity factor of lepidopteran-infecting baculoviruses. J Invertebr Pathol 109:117–126. https://doi.org/10.1016/j.jip.2011.10.010
Smith IRL, Crook NE (1988) In vivo isolation of baculovirus genotypes. Virology 166:240–244 pmid:2842948
Smits PH, Van de Vrie M, Vlak JM (1987) Nuclear polyhedrosis virus for control of Spodoptera exigua larvae on glasshouse crops. Entomol Exp Appl 43:73–80
Sun X (2015) History and current status of development and use of viral insecticides in China. Viruses 7:306–319 pmid:25609304. https://doi.org/10.3390/v7010306
Takatsuka J, Okuno S, Nakai M, Kunimi Y (2003) Genetic and biological comparisons of ten geographic isolates of a nucleopolyhedrovirus that infects Spodoptera litura (Lepidoptera: Noctuidae). Biol Control 26:32–39. https://doi.org/10.1016/S1049-9644(02)00113-5
Theze J, Cabodevilla O, Palma L, Williams T, Caballero P, Herniou EA (2014) Genomic diversity in european Spodoptera exigua multiple nucleopolyhedrovirus isolates. J Gen Virol 95:2297–2309. https://doi.org/10.1099/vir.0.064766-0
Van Oers MM, Abma-Henkens MHC, Herniou EA, de Groot JCW, Peters S, Vlak JM (2005) Genome sequence of Chrysodeixis chalcites nucleopolyhedrovirus, a baculovirus with two DNA photolyase genes. J Gen Virol 86:2069–2080. https://doi.org/10.1099/vir.0.80964-0
Wang J, Zhu Z, Zhang L (2016) Genome sequencing and analysis of Catopsilia pomona nucleopolyhedrovirus: a distinct species in group I Alphabaculovirus. PLoS ONE 11:e0155134 pmid:27166956. https://doi.org/10.1371/journal.pone.0155134
Widiawati H, Sukirno S, Sumarmi S, Purwanto H, Soesilohadi RH, Sudaryadi I (2021) UV Protectant Ability of Attacus atlas L. (Lepidoptera: Saturniidae) Sericin Extract to Increase Nucleopolyhedrovirus Effectiveness against Beet Army Worm, Spodoptera exigua (Hübner) (Lepidoptera: Noctuidae). In 7th International Conference on Biological Science pp. 82–89 Atlantis Press. https://doi.org/10.2991/absr.k.220406.013
Wolff JLC, Valicente FH, Martins R, de Castro Oliveira JV, de Andrade Zanotto PM (2008) Analysis of the genome of Spodoptera frugiperda nucleopolyhedrovirus (SfMNPV-19) and of the high genomic heterogeneity in group II nucleopolyhedroviruses. J Gen Virol 89:1202–1211. https://doi.org/10.1099/vir.0.83581-0
Xia-Ling Z, Cong XP, Wang XP, Lei CL (2011) A review of geographic distribution, overwintering and migration in Spodoptera exigua Hübner (Lepidoptera: Noctuidae). J Entomol Res Soc 13:39–48
Author information
Authors and Affiliations
Contributions
Data curation: Gozde Busra Eroglu.
Funding acquisition: Gozde Busra Eroglu, Javad Karimi.
Software: Gozde Busra Eroglu.
Validation: Javad Karimi.
Writing – original draft: Gozde Busra Eroglu.
Writing – review & editing: Javad Karimi.
Corresponding author
Ethics declarations
Conflict of Interest
All authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Additional file 1: Table S1.
Detail Information on the Spodoptera nucleopolyhedrovirus and out-group enomes used in the phylogenetic analyzes.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Eroglu, G.B., Karimi, J. Whole genome analysis of a novel Spodoptera exigua nucleopolyhedrovirus isolate (SeMNPV-IR) to Iran. Biologia 78, 2563–2574 (2023). https://doi.org/10.1007/s11756-023-01399-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-023-01399-2