Abstract
Introduction
Antibody response plays a fundamental role in the natural history of infectious disease. A better understanding of the immune response in patients with SARS-CoV-2 infection could be important for identifying patients at greater risk of developing a more severe form of disease and with a worse prognosis.
Methods
We performed a cross-sectional analysis to determine the presence and the levels of both anti-SARS-CoV-2 IgG and IgA in a cohort of hospitalized patients with confirmed infection at different times in the natural history of the disease. Patients enrolled when admitted at the emergency department were prospectively followed up during hospital stay.
Results
Overall, 131 patients were considered with a total of 237 samples processed. Cross-sectional analysis showed that seroconversion for IgA seems to occur between days 6 and 15, while IgG response seems to occur slightly later, peaking at day 20 after symptoms onset. Both IgA and IgG were maintained beyond 2 months. Severe patients showed a higher IgA response compared with mild patients when analyzing optical density (8.3 versus 5.6, p < 0.001). Prospective analysis conducted on 55 patients confirmed that IgA appear slightly earlier than IgG. After stratifying for the severity of disease, both the IgA and IgG responses were more vigorous in severe cases. Moreover, while IgG tended to stabilize, there was a relevant decline after the first month of IgA levels in mild cases.
Conclusion
IgA and IgG antibody response is closely related, although seroconversion for IgA occurs earlier. Both IgA and IgG are maintained beyond 2 months. Severe patients showed a more vigorous IgA and IgG response. IgA levels seem to decline after 1 month since the onset of symptoms in mild cases. Our results should be interpreted with cautions due to several limitations in our study, mainly the small number of cases, lack of data on viral load and clinical setting.
Similar content being viewed by others
Introduction
At the end of December 2019, a cluster of patients suffering from a new kind of pneumonia of unknown etiology has been observed in Wuhan (China) [1]; subsequently, it was attributed to a new virus (SARS-CoV-2), a beta-coronavirus phylogenetically similar to the causative agent of SARS, which determines the respiratory disease then called COVID-19 [2].
From December 2019 to today, the spread of SARS-CoV-2 infection has taken on the size of a pandemic [3]. According to the latest WHO data, to date, more than 100 million cases and two million deaths from COVID-19 have been confirmed worldwide [4], challenging the healthcare systems all over the world.
Moreover, although mortality from SARS-CoV-2 was found to be lower than that from SARS and MERS, COVID-19 deaths are greater in absolute number due to the higher transmissibility of this new virus [5].
As a result, public health services around the world have implemented different strategies for rapid identification of both infected and asymptomatic carriers to prevent virus transmission.
To date, direct detection of SARS-CoV-2 virus RNA throughout real-time reverse-transcriptase–polymerase-chain-reaction (rtPCR) assay from specimen of upper or lower airways has become the standard method for diagnosing SARS-CoV-2 infection in microbiology laboratories worldwide, although these tests have several limitations and many false negatives have been reported [6].
Recently, detection of antibodies specific to SARS-CoV-2 has also been recognized as deterministic evidence for confirmed SARS-CoV-2 infection [6]. However, the role and the pattern of antibody response in SARS-CoV-2 infection are not yet fully understood and many gaps are still present [7].
Neutralizing antibodies play a fundamental role in the clearance of viruses and are used as a gold standard for the evaluation of immunization in many viral infections.
Previous studies carried out during the SARS epidemic showed that the antibody titer and the rate of seroconversion were associated with the severity of the disease. In particular, in the study conducted by Lee et al. [8], seroconversion was observed around the 16th day and the peak of IgG was reached at 4 weeks; moreover, the rapidity of seroconversion (< 16 days) was associated with the need for hospitalization in intensive care unit (ICU), while high IgG antibody titers were associated with the need for ventilation and hospitalization in ICU. Other studies carried out during SARS epidemic, showed that patients with milder forms of disease had lower antibody titers [9,10,11,12], suggesting that, as observed in other infectious diseases (such as Dengue), an important humoral response may be associated with an exaggerated immune response and generate a cytokine storm [13,14,15,16], which can contribute to disease progression.
Despite many similarities to SARS-CoV infection, little information is currently available on the role of the antibody response in the evolution of SARS-CoV-2 disease. Moreover, most of the studies available at the moment explore the characteristic only of IgM and IgG response to SARS-CoV-2 infection [17,18,19,20], while little information are available about all the other mechanisms involved in the immune response, such as, for example, the role of IgA antibodies.
In the past, several studies have pointed out the crucial role of IgA antibodies as an effective defense against respiratory infections [21] and previous studies on SARS-CoV have shown similar kinetics of IgA, IgM, and IgG [13, 22]. It is well known that mucosal immunity plays a key role in limiting infections by respiratory pathogens [21], and preliminary data hint that IgA may be an essential part of immune response against SARS-CoV-2 as highlighted from other authors [23,24,25]. Thus, a better understanding of IgA response is needed for SARS-CoV-2.
A better understanding of the immune response in patients with SARS-CoV-2 infection could be important for the development of an effective vaccine and for identifying patients at greater risk of developing a more severe form of disease and with a worse prognosis [17].
In our study we investigate the characteristics of IgA and IgG antibody response in a cohort of patients with confirmed infection from SARS-COV-2 admitted to the Emergency Department (ED) of a referral COVID Hospital in Italy.
Methods
We conducted an observational, single-center study in a tertiary ED of a university hospital located in Rome, Italy, currently serving as a referral center for COVID-19.
Aim the of the study
Aim of the study was to describe the pattern of IgG and IgA antibody response against SARS-CoV-2 in hospitalized individuals.
The study involved two kind of analysis:
-
1.
A cross-sectional analysis which had the aim to evaluate the presence and levels of specific IgG and IgA antibodies at different times in the natural history of the disease in patients referred to the ED or hospitalized in our institution with confirmed SARS-CoV-2 infection in a considered time frame;
-
2.
A prospective observational analysis with the aim to evaluate among patients referred to the ED with confirmed SARS-CoV-2 infection, the presence and level of IgA and IgG antibodies at the time of diagnosis and at different time points after the onset of symptoms.
Furthermore, we assessed whether antibody levels are associated with the severity of infection.
Patient enrollment and study population
Enrollment procedures for both parts of the study are detailed below.
For the cross-sectional analysis, all patients who were hospitalized between May 1st and May 31st in any of the COVID-19-dedicated wards of our institution (either in the ED, in COVID-19-dedicated general medical wards, in COVID-19-dedicated ICU or in our institution’s supervised residential care) were evaluated for inclusion in the present study; patients fulfilling enrollment criteria underwent a single blood draw for IgA and IgG testing. For each patient, the date of symptoms onset was recorded.
For the prospective analysis all patients presenting to our institution’s ED between May, 1st and May, 31st who had a clinical suspicion of COVID-19 were evaluated for inclusion in the study; patients who fulfilled the inclusion criteria were enrolled in the study; for each patient, the day of symptoms onset, as recalled by the patient, was registered and a blood sample for IgA and IgG measurement was drawn.
Subsequently, patients hospitalized in any of the COVID-19-dedicated wards of our institution (either in COVID-19-dedicated general medical wards, in COVID-19-dedicated ICU or in our institution’s supervised residential care) underwent blood draw during the course of their illness at 7, 14, 21 days, 1 and 2 months since the admission.
Such data were analyzed independently from data obtained for the cross-sectional part of the study.
Furthermore, all data point obtained for the prospective enrollment were pooled with those obtained for the cross-sectional enrollment before cross-sectional analysis.
Inclusion criteria were the following:
-
suggestive clinical presentation (dyspnea, fever, cough, coryza; for patients with a pre-existing chronic respiratory condition, worsening dyspnea or worsening respiratory failure; chest imaging at X-ray, CT or lung ultrasound of interstitial involvement or multiple consolidations suspected for COVID-19);
-
microbiological confirmation of the diagnosis of SARS-CoV2 infection, defined as direct detection of SARS-CoV-2 virus RNA throughout rtPCR assay on nasopharyngeal swab performed at the time of access to the ED, in the preceding 7 days or during the first 7 days of the index hospitalization;
-
patient was 18 years or older at the time of the ED admission;
-
patient was willing to participate in the study.
Exclusion criteria were:
-
primary or secondary immunodeficiencies (however, patients with ongoing immunosuppressive therapy were included in the study);
-
active hematological malignancy;
-
patients in whom it was not possible to clearly define the onset of symptoms.
Study examinations
Blood specimen were collected via venipuncture in a 9 ml dry test tube, after discarding the first 2 ml of blood. The variables collected for each patient were the following: age, sex, clinical symptoms, time of onset of symptoms, radiological investigations (Chest X-ray, Chest CT scan), rt-PCR for SARS-CoV-2 on oro-pharyngeal and naso-pharyngeal swab with threshold cycle for each of the three tested genes (E gene, N gene and RdRP gene as proxies for the viral load), laboratory tests (Hb, MCV, WBC, neutrophil and lymphocyte count, C-reactive protein, fibrinogen, d-dimer, LDH), need for ICU admission, outcome of the hospital stay (death or discharge), overall severity of SARS-CoV-2 infection (mild disease, interstitial pneumonia without respiratory failure, mild–moderate–severe ARDS). ARDS and ARDS severity was defined according to generally used international guidelines [26, 27].
This study was conducted in accordance with the principles of Helsinki declaration [28].
Severity of disease definition
The disease severity was defined as follows:
-
Paucisymptomatic disease: symptomatic patients without radiographic evidence of pulmonary infiltrates and nor requiring oxygen supplementation;
-
Interstitial pneumonia: patients with radiographic evidence of pulmonary infiltrates, requiring either low-flow or high-flow oxygen supplementation, or requiring non-invasive positive pressure ventilation with a PaO2/FiO2 ratio above 300;
-
Mild ARDS: patients requiring either non-invasive or invasive ventilation, with a PaO2/FiO2 ratio between 201 and 300 with the use of at least 5cmH2O positive end expiratory pressure;
-
Moderate ARDS: patients requiring either non-invasive or invasive ventilation, with a PaO2/FiO2 ratio between 101 and 200 with the use of at least 5cmH2O positive end expiratory pressure;
-
Severe ARDS: patients requiring either non-invasive or invasive ventilation, with a PaO2/FiO2 ratio below 101 with the use of at least 5cmH2O positive end expiratory pressure.
For the purpose of the analysis, paucisymptomatic disease, pneumonia and mild ARDS were considered mild disease, while moderate and severe ARDS were considered severe disease.
Assay of IgA and IgG antibodies
Serum was separated by centrifugation at 2500 g for 5 min within 12 h of collection. The specimens were analyzed with SARS-CoV-2 IgG and IgA kits (Euroimmun, Lübeck, Germany; www.euroimmun.com) on the lab workstation platform. The EUROIMMUN Anti-SARS-CoV-2 Assay is an enzyme-linked immunosorbent assay (ELISA) that provides semi-quantitative in vitro determination of human antibodies of immunoglobulin classes IgA and IgG against SARS-CoV-2 in serum or EDTA plasma. Each kit contains microplate strips with 8 break-off reagent wells coated with S1 domain of viral spike protein recombinant of SARS-CoV-2. In the first reaction step, diluted patient samples are incubated in the wells. In the case of positive samples, specific antibodies will bind to the antigens. To detect the bound antibodies, a second incubation is carried out using an enzyme-labelled antihuman IgA or IgG (enzyme conjugate) catalyzing a color reaction. Results are evaluated semi-quantitatively by calculation of a ratio of the extinction of the control or patient sample over the extinction of the calibrator. This ratio is interpreted as follows: < 0.8 negative; ≥ 0.8 to < 1.0 borderline; ≥ 1.1 positive.
Statistical analysis
We planned to perform an exploratory data analysis on levels of circulating antibodies at different time points and on the proportion of patients with positive serological assays at different time points.
First, we divided the same samples in 5-day time periods and we calculated the proportion of positive patients for each time frame. To detect the most likely time of seroconversion for each antibody class, we compared the proportion of positive patients for each time frame with the proportion of positive patients of the subsequent time frame, adjusting significant p values for multiple comparison with the Holm method.
To assess the possible association between antibodies level or antibodies development and severity of disease, we divided our blood samples according to the maximum severity of the disease. Patients who developed a moderate or severe ARDS were classified as having severe disease, while other patients were classified as having mild disease. We compared the rate of positive sample for each class of antibodies between severe and mild patients both overall and after stratifying by time frame since the onset of symptoms. For this analysis, the positive/negative result of the antibody detection assay was considered as a dichotomous variable and each comparison was performed using the Pearson’s Chi-square test or the Fisher exact test, as appropriate. Multiple comparison correction was performed only when a statistically significant result was found using the Holm procedure. Statistical significance was considered as an alpha level of 0,05, two-sided.
The degree of antibody response was also compared between patients with severe or mild disease, using the optical density of the sample as a surrogate for the antibody titer. For this analysis, severe and mild patients were compared both overall and after stratifying for the above-defined time frames. Given that optical density is a continuous variable, either the Student’s t test or the Wilcoxon–Mann–Whitney test were used, as appropriate. Multiple comparison correction was performed only when statistically significant result was found using the Holm procedure. Statistical significance was considered as an alpha level of 0,05, two-sided.
It should be noted that this is a descriptive study; thus, calculated p values are only provided to identify most pronounced differences and are not meant to quantify the risk of alpha error on hypothesis testing.
LOESS curves were added to scatterplots to facilitate visual identification of trends [29].
Data collection was performed using Microsoft Excel for Office 365 (Microsoft Corporation, Redmond, WA, USA). Statistical analysis and plot design were performed with the R statistical language version 4.0.2 [30] via the RStudio IDE (RStudio, PBC, Boston, MA, USA) [31], using the Tidyverse package [32].
Results
Patient population and serological samples
Initially, 198 patients with suspected SARS-COV-2 infection were considered for inclusion. Of those, 30 were excluded because they did not have a microbiological confirmation of SARS-CoV-2 infection; Among 168 patients with confirmed infection, 37 patients were excluded because the onset of symptoms could not be clearly identified.
Overall, serological samples included in the analysis were obtained from 131 patients (see Table 1). 55 patients had multiple samples drawn during their hospital stay. The median age of our patients was 64 years (IQR 51.5–75) (Table 1). 34 patients developed a severe form of the disease (26%), while 97 were classified as having a mild form (74%). The median time between symptoms onset and sample collection was 23 days (IQR 12–45, range 1–88).
Overall, there were a total of 237 samples processed. Each sample was processed both for IgA and for IgG antibodies. Of those, 183 resulted positive for IgA (77.2%) and 174 for IgG (73.4%).
Cross-sectional analysis
Positivity rate per time frame: IgA
The rate of positive samples for IgA divided per time frame are shown in Table 2.
There were 18 samples drawn in the first 5 days from the onset of symptoms. Of those, 3 tested positive for IgA (point estimate: 16.7%, 95% CI 4.4–42.3%). Similarly, there were 18 samples drawn between day 6 and day 10, and of those only 3 tested positive for IgA (16.7%, 95% CI 4.4–42.3%). Conversely, there were 17 samples drawn between day 11 and day 15 and, of those, 12 tested positive for IgA (70.6%, 95% CI 44.0–88.6%). The point estimate for the proportion of positive samples remained stable over 80% from day 21 onward. There were 26 samples drawn beyond 60 days after the onset of symptoms, all of which tested positive for IgA (100%, 95% CI 82.2–100%).
The highest increase in rate of proportion of positive samples was observed between the second and the third time frame, that is between the 6–10-day samples and the 11–15-day samples.
Difference between each time frame and the subsequent one showed a peak for samples drawn between days 11 and 15, suggesting that IgA response develop in the majority of patients between days 6 and 15 since the beginning of symptoms (Fig. 1).
Positive rate per time frame: IgG
The rate of positive samples for IgG divided per time frame are given in Table 2.
There were 18 samples drawn in the first 5 days since the onset of symptoms. Of those, 2 tested positive for IgG (11.1%, 95% CI 1.9–36.1%). A similar result applies to the samples drawn between days 6 and 10. A first increase for the rate of positive samples was observed for the samples drawn between 11 and 15 days from the onset of symptoms. At this time frame, there were 18 samples and of those, 6 tested positive for IgG (33.3%, 95% CI 14.4–58.8%). A second increase in the rate of positive samples was observed for samples drawn between 16 and 20 days. At this time frame, there were 17 samples and, of these, 13 tested positive for IgG (76.5%, 95% CI 49.8–92.2%). Of note, the difference between positive rates at time frame 11–15 and 16–20 was the only difference to reach statistical significance, although this significance was not maintained after correction for multiple comparison with the Holm method. There were 26 samples drawn beyond 60 days from the onset of symptoms, all of which tested positive for IgG (100%, 95% CI 82.2–100%).
Differences between consecutive time frames showed a more marked increase in the rate of positive values for samples drawn between days 11 and 15 and for samples drawn between days 16 and 20, suggesting that appearance of IgG may happen between days 6 and 20 (see Fig. 1).
Optical density
Overall, there were 237 samples tested for IgA. The median optical density was 4.7 unit (IQR 1.5–8.3). The same 237 samples were also tested for IgG, for which the median optical density was 5.5 (IQR 0.5–9.1). Visual inspection of the dot plot of optical density versus time showed a similar trend in time for both IgA and IgG (Fig. 2).
Serological response according to severity of disease
After stratifying per time frame and per severity of the disease, there was no statistically significant difference between severe patients and mild patients in the rate of positive result.
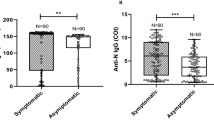
Conversely, there was a statistically significant difference for the overall optical density of IgA between severe patients and mild patients, with severe patients overall showing a higher optical density for IgA (8.3 versus 5.6, p < 0.001). A smaller difference was observed for IgG, with more severe patients showing an overall higher optical density, although for IgG statistical significance was not reached (8.4 versus 7.0, p = 0.09) (Fig. 3). Such a difference was not observed after stratifying the samples per time since the onset of symptoms.
Prospective analysis
Fifty-five patients of our sample were prospectively enrolled; for those patients, 161 blood samples were collected, with an average of 2.93 samples per patient. The average time between the onset of symptoms and the Ig test was 27.3 days. The longest interval between symptoms onset and Ig test was 88 days. Overall, of the 161 samples, 117 tested positive for IgA (72.7%, 95% CI 65.8–79.6%), while 107 tested positive for IgG (66.5%, 95% CI 59.2–73.8%).
Of those 55 patients, 19 (34.5%) developed a severe form of the disease, while 36 (65.5%) developed a mild form of the disease.
The median threshold cycle at the RT-PCR performed on the naso-pharyngeal swab at the admission were similar between patients who developed a severe form of the disease and patients with a milder form of the disease (for E gene 27.1 [IQR 24.0–29.1] versus 25.6 [IQR 22.5–31.4], for N gene 28.9 [IQR 24.8–32.1] versus 29.3 [IQR 23.0–32.3], for RdRP gene 28.9 [IQR 22.7–30.1] versus 28.3 [IQR 22.2–30.9]).
The trend of antibody response showed that both IgA and IgG became detectable between days 5 and 7 from the symptoms onset, peaked between days 21 and 27 and remained stable thereafter, with both IgA and IgG showing a similar trend before day 50. There seemed to be a declining trend for IgA past day 50, while IgG maintained high levels (Fig. 4a).
a Density versus time plot for different Ig class for prospectively enrolled patients. b Density versus time plot for different Ig class and for different severity of disease in prospectively enrolled patients. LOESS curve interpolation is shown. Shaded areas represent confidence intervals around LOESS curves
After dividing patients according to the severity of disease, peak antibody response appeared to be similar among the two groups. However, there was a more vigorous antibody response both for IgG and IgA in severe patients. Moreover, while IgG tended to stabilize in mild cases, severe cases showed a trend toward more sustained antibody response in time. IgA levels peaked at about the same time in mild and severe patients; however, in mild patients there was a relevant decline after the first month; such a trend was not observable in severe cases (Fig. 4b).
Discussion
Antibody response plays a vital role in viral infections, both for the clearing of the pathogen and for lasting immunity. Serological assays are widely used for the diagnosis of viral infections. However, there is still incomplete knowledge about the kinetic of antibody response to SARS-CoV-2.
Preliminary data reported by Wu et al. out of 175 patients with mild form of disease who did not require hospitalization in ICU highlighted the presence of neutralizing anti-SARS-CoV-2 antibodies 10–15 days after the beginning of the disease. In this paper, however, the authors highlight a rather heterogeneous humoral response in the study population, identifying approximately 30% of patients analyzed with a low antibody titer, therefore, suggesting that other immune responses may play an important role. It is not yet known whether these patients are at greater risk of reinfection [20].
Moreover, Zhao et al. [18] in their study conducted among 173 confirmed patients, found out that seroconversion rate was 93,1%, 82,7% and 64,7% for total antibodies (Ab), IgM and IgG, respectively, and that median time to Ab, IgM and IgG seroconversion was 11, 12 and 14 days, separately. In this study the seroconversion of Ab was significantly quicker than that of IgM (p = 0.012) and IgG (p < 0.001). According to the authors, this is maybe because all isotypes of viral specific antibodies, including not only IgM and IgG but also IgA, can be detected by assay for Ab test.
This evidence supports the hypothesis that other responses to SARS-CoV-2 contribute to virus clearance, and that, to date, we still do not fully know all the mechanisms involved in the immune response to SARS-CoV-2, such as, for example, the role of IgA antibodies against SARS-CoV-2. A similar observation has also been made by Di Giambenedetto et al. [19].
In our study, IgA response partially resembles IgG response. However, IgA seroconversion seems to happen slightly earlier than IgG. Most patients developed IgA between days 11 and 15, with only small increases of positivity rates thereafter. Conversely, IgG seroconversion seems to occur in a longer time frame, with patients developing antibodies between days 11 and 20. Both for IgA and IgG, the positivity rate remained stable over 80% after day 21 and reached 100% for samples drawn beyond day 60. A recent large-scale seroprevalence study conducted by Gudbjartsson and colleagues showed a similar result, with more than 90% of samples testing positive after the second month since molecular diagnosis of SARS-CoV-2 infection and up to 4 months [33]. Long and colleagues found a smaller time to plateau of 6 days for IgG concentration in a sample of 249 COVID-19 patients; however, they also found that IgG positivity rate reached 100% between 17 and 19 days after symptoms onset, a finding which is similar to what we found in our sample [34].
Our data show that there seems to be a difference in the degree of antibody response between mild and severe patients. Specifically, when analyzing all the samples collected, regardless of the time point, there seemed to be a more vigorous antibody response in severe cases compared to mild cases. This difference was more evident for IgA than for IgG. However, we failed to find any difference when blood samples were stratified for time since the onset of symptoms. The finding of higher levels of IgA antibodies in severe patients compared to milder patients has been reported by other groups [35, 36], with at least one group reporting higher levels of both IgG and IgA, but not IgM, in severe patients compared to mild patients [37].
Analysis of the prospectively enrolled patients confirmed the findings of the cross-sectional part of the study, with both IgA and IgG showing a similar trend and with IgA appearing slightly earlier than IgG. Moreover, IgA showed a slight decline after the first month. Interestingly, when stratifying for the severity of disease, this decline was evident in mild cases, while no such a decline was observed in severe cases. Unlike IgA, IgG seemed to remain stable in time. In addition, in mild cases, both the IgA and IgG response seemed to be slightly less vigorous than in severe cases. Also the study conducted by Gudbjartsson and colleagues showed a decline in IgA concentration after the first month [33].
As observed in other infectious diseases, humoral response may be associated with an exaggerated immune response, which, in turn, may be associated with disease progression and with worse prognosis. This mechanism has been also hypothesized for COVID-19 [38,39,40,41]. Indeed, we observed an association between humoral response and severity of disease and such a finding has been reported also by other groups [35, 36, 42, 43]. Interestingly, also the above said-large scale study detected an association between disease severity and strength of humoral response [33].
Mechanisms of severe COVID-19 are still a subject of ongoing investigation. It is well known that higher levels of inflammatory cytokines are associated with more severe disease [44] and a link between immune response and organ damage has been proposed [45]. Both the type of immune response and the efficacy of viral clearance have been associated with disease severity [46], thus suggesting that either failure in early phase viral clearance or host immune response may be associated with excessive immune activation and organ damage. Recently, an association between efficacy of interferon response and disease severity has been proposed, thus supporting that inability of the host to achieve viral clearance may be lead to dysregulated immune response [47, 48]. Thus, higher levels of circulating antibodies may reflect either sustained antigenic stimulation due to ineffective viral clearance or a consequence of more vigorous immune response which, in turn, is responsible for end organ damage. The possible mechanism of association between immune response and disease severity in COVID-19 have been excellently reviewed elsewhere [49, 50].
Limitations
Our study has several limitations and our results should be interpreted with caution. First, its stated goal is exploratory in nature. Thus, the result of our study should be used only for hypothesis generation, and not to be intended as confirmatory. Where statistical significance has been formally calculated, p values below 0,05 should be interpreted as a signal of a possible association, not as a confirmatory result.
Our study included only hospitalized patients or patients in supervised health care setting who were unable to maintain home isolation, thus generalization to other patients’ populations, especially to asymptomatic patients or outpatients, should be done with caution. This is particularly true in the case of association between antibody response and severity of the disease. While there seemed to be an association between intensity of IgA response and the severity of the disease when the overall sample was evaluated, we could not find such an association at the single evaluated time frames. This could be due to lack of association or to the fact that the sample size for each time frame was too small to detect such an association. Furthermore, patients who were enrolled in this study were either hospitalized for active disease or had positive swabs for SARS-CoV-2 infection. Thus, they had active antigen stimulation. Our study does not allow us to draw conclusions on patients who have recovered from their disease or have achieved complete viral clearance. Moreover, another major limitation is that we did not collect data on the time to complete viral clearance nor we quantified the viral load. Thus, we could not evaluate the role of viral load or time to viral clearance as a possible explanation of our findings.
Furthermore, due to the small cohort of patients enrolled it was not possible to evaluate the role of other characteristics such as age or gender on the humoral response.
Moreover, in the prospective part of the study, we only collected blood samples while the patients were hospitalized. We were not able to collect blood samples at the prespecified time points after patient’s discharge.
Finally, our institution is a large university hospital that is serving as referral center for the COVID-19 epidemics and our study has been conducted during the initial peak of incidence of COVID-19 in Italy; thus, generalization of our results to other settings or to other time period should be made with caution.
Conclusion
In summary, IgA and IgG antibody response seem to be closely related. Seroconversion for IgA may occur between days 6 and 15 in symptomatic patients. IgG antibody response may occur slightly later, peaking at day 20 after symptoms onset. Both IgA and IgG are maintained beyond 2 months since the beginning of symptoms. There seems to be no correlation between time of antibody response and severity of the disease while such an association is possible for intensity of antibody response. IgA levels seem to decline after 1 month since the onset of symptoms. The small sample size, the lack of information about viral load and the stated exploratory nature of our study constitute major limitations of our study and caution should be used in interpreting our results. Moreover, our results are obtained in hospitalized symptomatic patients during the first peak of COVID-19 epidemics in Italy and, thus, they are not generalizable to other time frame, patients’ populations or clinical setting. Finally, our study was not designed to assess relationship between antibody response and time to viral clearance nor to evaluate relationship between other clinical variables and antibody response.
References
Zhu N, Zhang D, Wang W et al (2020) A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 382:727–733. https://doi.org/10.1056/NEJMoa2001017
Lu R, Zhao X, Li J et al (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet (London, England) 395:565–574. https://doi.org/10.1016/S0140-6736(20)30251-8
WHO (2020) WHO Director-General’s opening remarks at the media briefing on COVID-19—11 March 2020. https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020. Accessed 15 May 2020
Wu Z, McGoogan JM (2020) Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China. JAMA 323:1239. https://doi.org/10.1001/jama.2020.2648
Li Z, Yi Y, Luo X et al (2020) Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J Med Virol 92:1518–1524. https://doi.org/10.1002/jmv.25727
Deeks JJ, Dinnes J, Takwoingi Y et al (2020) Antibody tests for identification of current and past infection with SARS-CoV-2. Cochrane database Syst Rev 6:CD013652. https://doi.org/10.1002/14651858.CD013652
Lee N, Chan PKS, Ip M et al (2006) Anti-SARS-CoV IgG response in relation to disease severity of severe acute respiratory syndrome. J Clin Virol 35:179–184. https://doi.org/10.1016/j.jcv.2005.07.005
Ip M, Chan PKS, Lee N et al (2004) Seroprevalence of antibody to severe acute respiratory syndrome (SARS)-associated coronavirus among health care workers in SARS and non-SARS medical wards. Clin Infect Dis 38:e116–e118. https://doi.org/10.1086/421019
Lee HKK, Tso EYK, Chau TN et al (2003) Asymptomatic severe acute respiratory syndrome–associated coronavirus infection. Emerg Infect Dis 9:1491–1492. https://doi.org/10.3201/eid0911.030401
Tso EYK, Tsang OTY, Lam B et al (2004) Natural course of severe acute respiratory syndrome-associated coronavirus immunoglobulin after infection. J Infect Dis 190:1706–1707. https://doi.org/10.1086/424573
Wilder-Smith A, Teleman MD, Heng BH et al (2005) Asymptomatic SARS coronavirus infection among healthcare workers, Singapore. Emerg Infect Dis 11:1142–1145. https://doi.org/10.3201/eid1107.041165
Hsueh P-R, Huang L-M, Chen P-J et al (2004) Chronological evolution of IgM, IgA, IgG and neutralisation antibodies after infection with SARS-associated coronavirus. Clin Microbiol Infect 10:1062–1066. https://doi.org/10.1111/j.1469-0691.2004.01009.x
Li G, Chen X, Xu A (2003) Profile of specific antibodies to the SARS-associated coronavirus. N Engl J Med 349:508–509. https://doi.org/10.1056/NEJM200307313490520
He Z, Dong Q, Zhuang H et al (2004) Kinetics of severe acute respiratory syndrome (SARS) coronavirus-specific antibodies in 271 laboratory-confirmed cases of SARS. Clin Diagnostic Lab Immunol 11:792–794. https://doi.org/10.1128/CDLI.11.4.792-794.2004
Haveri A, Smura T, Kuivanen S et al (2020) Serological and molecular findings during SARS-CoV-2 infection: the first case study in Finland, January to February 2020. Eurosurveillance. https://doi.org/10.2807/1560-7917.ES.2020.25.11.2000266
Okba NMA, Müller MA, Li W et al (2020) Severe acute respiratory syndrome coronavirus 2−specific antibody responses in coronavirus disease patients. Emerg Infect Dis 26:1478–1488. https://doi.org/10.3201/eid2607.200841
Zhao J, Yuan Q, Wang H et al (2020) Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin Infect Dis. https://doi.org/10.1093/cid/ciaa344
Di Giambenedetto S, Ciccullo A, Posteraro B et al (2020) Still much to learn about the diagnostic role of SARS-CoV-2 antibody detection. Clin Infect Dis. https://doi.org/10.1093/cid/ciaa532
Wu F, Wang A, Liu M et al (2020) Neutralizing antibody responses to SARS-CoV-2 in a COVID-19 recovered patient cohort and their implications. medRxiv. https://doi.org/10.1101/2020.03.30.20047365
Terauchi Y, Sano K, Ainai A et al (2018) IgA polymerization contributes to efficient virus neutralization on human upper respiratory mucosa after intranasal inactivated influenza vaccine administration. Hum Vaccin Immunother 14:1351–1361. https://doi.org/10.1080/21645515.2018.1438791
Woo PCY, Lau SKP, Wong BHL et al (2004) Longitudinal profile of immunoglobulin G (IgG), IgM, and IgA antibodies against the severe acute respiratory syndrome (SARS) coronavirus nucleocapsid protein in patients with pneumonia due to the SARS coronavirus. Clin Diagn Lab Immunol 11:665–668. https://doi.org/10.1128/CDLI.11.4.665-668.2004
Lippi G, Mattiuzzi C (2020) Clinical value anti-SARS-COV-2 serum IgA titration in patients with COVID-19. J Med Virol. https://doi.org/10.1002/jmv.26539
Xue M, Zhang T, Hu H et al (2020) Predictive effects of IgA and IgG combination to assess pulmonary exudation progression in COVID-19 patients. J Med Virol. https://doi.org/10.1002/jmv.26437
Chao YX, Rötzschke O, Tan E-K (2020) The role of IgA in COVID-19. Brain Behav Immun 87:182–183. https://doi.org/10.1016/j.bbi.2020.05.057
Bernard GR, Artigas A, Brigham KL et al (1994) The American-European Consensus on ARDS. Definitions, mechanisms, relevant outcomes, and clinical trial coordination. Am J Respir Crit Care Med 149:818–824. https://doi.org/10.1164/ajrccm.149.3.7509706
Definition Task Force ARDS, Ranieri VM, Rubenfeld GD et al (2012) Acute respiratory distress syndrome: the Berlin Definition. JAMA 307:2526–2533. https://doi.org/10.1001/jama.2012.5669
World Medical Association (2013) World Medical Association Declaration of Helsinki. JAMA 310:2191. https://doi.org/10.1001/jama.2013.281053
Cleveland WS (1979) Robust locally weighted regression and smoothing scatterplots. J Am Stat Assoc 74:829. https://doi.org/10.2307/2286407
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
RStudio Team (2019) RStudio: Integrated Development Environment for R. R. RStudio Inc, Boston
Wickham H, Averick M, Bryan J et al (2019) Welcome to the Tidyverse. J Open Source Softw 4:1686. https://doi.org/10.21105/joss.01686
Gudbjartsson DF, Norddahl GL, Melsted P et al (2020) Humoral Immune Response to SARS-CoV-2 in Iceland. N Engl J Med. https://doi.org/10.1056/NEJMoa2026116
Long Q-X, Liu B-Z, Deng H-J et al (2020) Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat Med 26:845–848. https://doi.org/10.1038/s41591-020-0897-1
Huang Z, Chen H, Xue M et al (2020) Characteristics and roles of severe acute respiratory syndrome coronavirus 2-specific antibodies in patients with different severities of coronavirus 19. Clin Exp Immunol. https://doi.org/10.1111/cei.13500
Hasan Ali O, Bomze D, Risch L et al (2020) Severe COVID-19 is associated with elevated serum IgA and antiphospholipid IgA-antibodies. Clin Infect Dis. https://doi.org/10.1093/cid/ciaa1496
Yu H-Q, Sun B-Q, Fang Z-F et al (2020) Distinct features of SARS-CoV-2-specific IgA response in COVID-19 patients. Eur Respir J 56:7–10. https://doi.org/10.1183/13993003.01526-2020
Tay MZ, Poh CM, Rénia L et al (2020) The trinity of COVID-19: immunity, inflammation and intervention. Nat Rev Immunol 20:363–374. https://doi.org/10.1038/s41577-020-0311-8
Jose RJ, Manuel A (2020) COVID-19 cytokine storm: the interplay between inflammation and coagulation. Lancet Respir Med 8:e46–e47. https://doi.org/10.1016/S2213-2600(20)30216-2
Mehta P, McAuley DF, Brown M et al (2020) COVID-19: consider cytokine storm syndromes and immunosuppression. Lancet (London, England) 395:1033–1034. https://doi.org/10.1016/S0140-6736(20)30628-0
García LF (2020) Immune Response, Inflammation, and the Clinical Spectrum of COVID-19. Front Immunol 11:1441. https://doi.org/10.3389/fimmu.2020.01441
Yang W, Sirajuddin A, Zhang X et al (2020) The role of imaging in 2019 novel coronavirus pneumonia (COVID-19). Eur Radiol. https://doi.org/10.1007/s00330-020-06827-4
Ma H, Zeng W, He H et al (2020) Serum IgA, IgM, and IgG responses in COVID-19. Cell Mol Immunol 17:773–775. https://doi.org/10.1038/s41423-020-0474-z
Zhu Z, Cai T, Fan L et al (2020) Clinical value of immune-inflammatory parameters to assess the severity of coronavirus disease 2019. Int J Infect Dis 95:332–339. https://doi.org/10.1016/j.ijid.2020.04.041
Sinha P, Matthay MA, Calfee CS (2020) Is a “cytokine storm” relevant to COVID-19? JAMA Intern Med 180:1152. https://doi.org/10.1001/jamainternmed.2020.3313
Lucas C, Wong P, Klein J et al (2020) Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584:463–469. https://doi.org/10.1038/s41586-020-2588-y
Zhang Q, Bastard P, Liu Z et al (2020) Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 370:eabd4570. https://doi.org/10.1126/science.abd4570
Bastard P, Rosen LB, Zhang Q et al (2020) Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science. https://doi.org/10.1126/science.abd4585
Moore JB, June CH (2020) Cytokine release syndrome in severe COVID-19. Science 368:473–474. https://doi.org/10.1126/science.abb8925
Fajgenbaum DC, June CH (2020) Cytokine Storm. N Engl J Med 383:2255–2273. https://doi.org/10.1056/NEJMra2026131
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no funding or conflicts of interest to disclose.
Ethical approval
The study was approved by the Ethical Committee of our Institution (ID 3256). Informed consent was obtained from each patient.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Carnicelli, A., Fiori, B., Ricci, R. et al. Characteristic of IgA and IgG antibody response to SARS-CoV-2 infection in an Italian referral COVID-19 Hospital. Intern Emerg Med 17, 53–64 (2022). https://doi.org/10.1007/s11739-021-02750-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11739-021-02750-8