Abstract
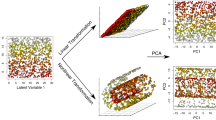
The most common means of performing ordination and classification consist in principal component, canonical variate, and between-group principal component analysis (PCA, CVA & bgPCA) for ordination, and linear and partial least squares discriminant analysis (LDA & PLSDA) for classification. Over the years, research has shown how the number of variables used in Geometric Morphometrics can be problematic for studies using small sample sizes. In the case of ordination, this implies an inflation of differences between groups, even when no differences are present. In light of this, classification tasks should also theoretically present exaggerated accuracy scores. Using a theoretically constructed geometric experiment, the present study constructs a series of imbalanced theoretical datasets containing different degrees of variation in both shape and form. Each ordination and classification task is then carried out to observe how imbalance influences the quality of results. Even when using large enough sample sizes, if one sample is considerably smaller than another, then this imbalance will have an effect on both ordination and classification results. Imbalance is thus seen to force separation among samples, and a considerable loss in classification performance. Statistical tests such as Procrustes distance calculations are not affected. The conclusions suggest that prior dimensionality reduction such as PCA are necessary for CVA, bgPCA, LDA and PLSDA. Cross-validated versions of these algorithms should also be used. An extensive discussion is also provided into alternative ordination and classification techniques that could prove useful for Geometric Morphometrics, and that are less sensitive to sample imbalance.















Similar content being viewed by others
Data Availability
No data was explicitly used for this study, however all R code developed for the simulation of data and experiments can be found at the corresponding author’s GitHub page via: https://github.com/LACourtenay/gmm_ordination_classification_experimental_toolkit.
References
Albrecht, G. H. (1992). Assessing the affinities of fossils using canonical variates and generalized distances. Journal of Human Evolution, 7(4), 49–69.
Barker, M., & Rayens, W. (2003). Partial least squares for discrimination. Journal of Chemometrics, 17, 166–173.
Bendale, A., & Boult, T. E. (2015). Towards open set deep networks. IEEE Conference on Computer Vision and Pattern Recognition. https://doi.org/10.1109/CVPR.2016.173
Bookstein, F. L. (1984). A statistical method for biological shape comparisons. Journal of Theoretical Biology, 107, 475–520.
Bookstein, F. L. (1986). Size and shape spaces for landmark data in two dimensions (with discussion). Statistical Science, 1, 181–242.
Bookstein, F. L. (1991). Morphometric tools for Landmark Data. Cambridge University Press.
Bookstein, F. L. (2017). A newly noticed formula enforces fundamental limits on geometric morphometric analyses. Evolutionary Biology, 44, 522–541. DOI: https://doi.org/10.1007/s11692-017-9424-9.
Bookstein, F. L. (2019). Pathologies of between-groups principal components analysis in geometric morphometrics. Evolutionary Biology, 46, 271–302. DOI: https://doi.org/10.1007/s11692-019-09484-8.
Boulesteix, A. L. (2004). A note on between-group pca. International Journal of Pure and Applied Mathematics, 19, 359–366.
Cardini, A., & Elton, S. (2007). Sample size and sampling error in geometric morphometric studies of size and shape. Zoomorphology, 126, 121–134. DOI: https://doi.org/10.1007/s00435-007-0036-2.
Cardini, A., Seetah, K., & Barker, G. (2015). How many specimens do I need? Sampling error in geometric morphometrics: testing the sensitivity of means and variances in simple randomized selection experiments. Zoomorphology, 134, 149–163. DOI: https://doi.org/10.1007/s00435-015-0253-z.
Cardini, A., O’Higgins, P., & Rohlf, F. J. (2019). Seeing distinct groups where there are none: spurious patterns from between-group PCA. Evolutionary Biology, 46, 303–316. DOI: https://doi.org/10.1007/s11692-019-09487-5.
Cardini, A., & Polly, P. D. (2020). Cross-validated between Group PCA scatterplots: a solution to spurious group separation? Evolutionary Biology, 47, 85–95. DOI: https://doi.org/10.1007/s11692-020-09494-x.
Chawla, N. V., Bowyer, K. W., Hall, L. O., & Kegelmeyer, W. P. (2002). SMOTE: synthetic minority over-sampling technique. Journal of Artificial Intelligence Research, 16, 321–357.
Chow, G. C. (1960). Tests of equality between sets of coefficients in two linear regressions. Econometrica, 28(3), 591–605.
Cobb, S. N., & O’Higgins, P. (2004). Hominins do not share a common postnatal facial ontogenetic shape trajectory. Journal of Experimental Zoology, 302(B), 302–321.
Cohen, J. (1988). Statistical power analysis for behavioural sciences. Routledge.
Colquhoun, D. (2019). The false positive risk: a proposal concerning what to do about p-values. The American Statistician, 73(Sup1), 192–201. DOI:https://doi.org/10.1080/00031305.2018.1529622.
Coombs, W. T., Aligna, J., & Oltman, D. O. (1996). Univariate and multivariate omnibus hypothesis tests to control type I error rates when population variances are not necessarily equal. Review of Educational Research, 66(2), 137–179.
Courtenay, L. A., González-Aguilera, D., Lagüela, S., del Pozo, S., Ruiz-Mendez, C., Barbero-García, I., Román-Curto, C., Cañueto, J., Santos-Durán, C., Cardeñoso-Álvarez, M. E., Roncero-Riesco, M., Hernandez-Lopez, D., Guerrero-Sevilla, D., & Rodríguez-Gonzalvez, P. (2021). Hyperspectral imaging and robust statistics in non-melanoma skin cancer analysis. Biomedical Optics Express, 12(8), 5107–5127. DOI:https://doi.org/10.1364/BOE.428143.
Courtenay, L. A., Aramendi, J., & González-Aguilera, D. (xxxx). Recruiting a skeleton crew—methods for simulating and augmenting palaeoanthropological data using Monte Carlo based algorithms. American Journal of Biological Anthropology.
Dhamija, A. R., Günther, M., & Boult, T. E. (2018). Reducing network agnostophobia. Neural Information Processing Systems, 32, 1–10.
Efron, B. (1979). Bootstrap methods: another look at the jackknife. Annals of Statistics, 7, 1–26.
Fernández, A., García, S., Galar, M., Prati, R. C., Krawczyk, B., & Herrera, F. (2018). Learning from imbalanced datasets. Springer.
Fisher, R. A. (1936). The use of multiple measurements in taxonomic problems. Annals of Eugenics, 7, 179–188.
Goodall, C. (1991). Procrustes methods in the statistical analysis of shape. Journal of the Royal Statistical Society B, 53(2), 285–339.
Gower, J. C. (1975). Generalized procrustes analysis. Pyschometrika, 40, 33–51.
Gupta, P. L., & Gupta, R. D. (1987). Sample size determination in estimating a covariance matrix. Computational Statistics and Data Analysis, 5, 185–192.
He, H., Bai, Y., Garcia, E.A. & Li, S. (2008) ADASYN: Adaptive synthetic sampling approach for imbalanced learning. Proceedings of the IEEE international joint conference on neural networks, Hong Kong, China, 1–8 June
He, H., & Ma, Y. (2013). Imbalanced Learning: foundations, algorithms and applications. Wiley.
Hinton, G. E., & Roweis, S. T. (2002). Stochastic neighbor embedding. Advances in Neural Information Processing Systems, 15, 857–864.
Kendall, D. G. (1983). The shape of poisson-delaunay triangles. In M. C. Demetrescu, & M. Iosifescu (Eds.), Studies in probabilities and related topics in Honour of Octav Onicescu (pp. 321–330). Nagard.
Kendall, D. G. (1984). Shape, manifolds, procrustean metrics, and complex projective spaces. Bulletin of the London Mathematical Society, 16, 81–121.
Kendall, M. G. (1955). Rank correlation methods. Haffner Publishing Co.
Klingenberg, C. P., & Monteiro, L. R. (2005). Distances and directions in multidimensional shape spaces: implications for morphometric applications. Systematic Biology, 54(4), 678–688. DOI: https://doi.org/10.1080/10635150590947258.
Klingenberg, C. P. (2013). Cranial integration and modularity: insights into evolution and development from morphometric data. Hystrix Italian Journal of Mammology, 24, 43–58.
Liu, X. Y., & Zhou, Z. H. (2013). Ensemble methods for class imbalance learning. In: H. He and Y. Ma (Eds.) Imbalanced Learning. 61–82
Maaten, L., & Hinton, G. (2008). Visualizing data using t-SNE. Journal of Machine Learning Research, 9, 2579–2605.
Mardia, K. V., Kent, J. T., & Bibby, J. M. (1997). Multivariate analysis. Academic Press.
Marčenko, V. A., & Pastur, L. A. (1967). Distribution of eigenvalues for some sets of random matrices. Mathematics of the USSR Sbornik, 1, 457–483.
McInnes, L., Healy, J., & Melville, J. (2018). UMAP: Uniform Manifold approximation and projection. Journal of Open Source Software, 3(29), 861. DOI: https://doi.org/10.22105/joss.00861.
Mitteroecker, P., & Bookstein, F. (2011). Linear discrimination, ordination, and the visualization of selection gradients in modern morphometrics. Evolutionary Biology, 38, 100–114. doi:https://doi.org/10.1007/s11692-011-9109-8.
Nguyen, A., Yosinski, J., & Clune, J. (2015). Deep neural networks are easily fooled: high confidence predictions for unrecognizable images. Computer Vision and Pattern Recognition, 15, 427–436.
O’Higgins, P., & Dryden, I. L. (1993). Sexual dimorphism in hominoids: further studies of craniofacial shape differences in Pan, Gorilla and Pongo. Journal of Human Evolution, 24, 182–205.
Pearson, K. (1895). Note on regression and inheritance in the case of two parents. Proceedings of the Royal Society of London. 58:347–352
Rao, C. R. (1951). An asymptotic expansion of the distribution of Wilk’s criterion. Bulletin of the International Statistical Institute, 33(2), 177–180.
Rokach, L. (2010). Pattern classification using ensemble methods. World Scientific.
Rohlf, J. F. (1996). Morphometric spaces, shape components, and the effects of linear transformations. In L. F. Marcus, M. Corti, A. Loy, G. P. Naylor; and, & D. E. Slice (Eds.), Advances in Morphometrics (pp. 117–129). Plenum.
Rohlf, J. F. (1999). Shape statistics: Procrustes superimposition and tangent spaces. Journal of Classification, 16, 197–223.
Rohlf, J. F. (2000a). On the use of shape spaces to compare morphometric methods. Hystrix Italian Journal of Mammology, 11, 8–24.
Rohlf, J. F. (2000b). Statistical power comparisons among alternative morphometric methods. American Journal of Physical Anthropology, 111, 463–478.
Rohlf, J. F. (2021). Why clusters and other patterns can seem to be found in analyses of high-dimensional data. Evolutionary Biology, 48, 1–16. DOI: https://doi.org/10.1007/s11692-020-09518-6.
Shannon, C. E. (1948a). A mathematical theory of communication. The Bell System Technical Journal, 27(3), 379–423. DOI: https://doi.org/10.1002/j.1538-7305.1948.tb01338.x.
Shannon, C. E. (1948b). A mathematical theory of communication. The Bell System Technical Journal, 27(4), 623–656. DOI: https://doi.org/10.1002/j.1538-7305.1948.tb00917.x.
Slice, D. E. (2001). Landmark coordinates aligned by procrustes analysis do not lie in Kendall’s shape space. Systematic Biology, 50(1), 141–149.
Szegedy, C., Zaremba, W., Sutskever, I., Bruna, J., Erhan, D., Goodfellow, I., & Fergus, R. (2014). Intriguing properties of neural networks. International Conference on Learning Representations. https://arxiv.org/abs/1312.6199. Accessed 17 June 2022
Takeshita, T., Nozawa, S., & Kimura, F. (1993). On the bias of Mahalanobis distance due to limited sample size effect. Proceedings of the International Conference on Document Analysis and Recognition, 2, 171–174. https://doi.org/10.1109/ICDAR.1993.395756
Zhou, Z. H. (2012). Ensemble methods: foundations and algorithms. Taylor & Francis.
Acknowledgements
The corresponding author would like to thank Julia Aramendi for her support and suggestions when carrying out his research.
Funding
L.A.C. is funded by the Spanish Ministry of Science, Innovation and Universities with an FPI Predoctoral Grant (Ref. PRE2019-089411), associated with the project RTI2018-099850-B-I00 and the University of Salamanca.
Author information
Authors and Affiliations
Contributions
L.A.C. Conceptualization, Formal Analysis, Investigation, Methodology, Software, Visualization, Writing – Original Draft, Review & Editing.
Corresponding author
Ethics declarations
Conflict of interest
The corresponding author has no competing interests to declare.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Courtenay, L.A. Can we Restore Balance to Geometric Morphometrics? A Theoretical Evaluation of how Sample Imbalance Conditions Ordination and Classification. Evol Biol 50, 90–110 (2023). https://doi.org/10.1007/s11692-022-09590-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11692-022-09590-0