Abstract
The potential and direction of phenotypic evolution is constrained by the distribution of genetic variation for the traits as described by the phenotypic (P) and genetic covariance matrices (G). The rank of the covariance matrix reflects the number of independent variational dimensions of the phenotype. Covariance matrices with less than full rank indicate lack of variation in some directions of the phenotype space and thus are an indication of absolute evolutionary constraints. Because selection acts upon phenotypic variation, the rank of P represents the upper limit of the dimensionality in G, relevant for selection response. The limitations of current methods to estimate matrix rank motivated us to analyze and adjust a bootstrap method and evaluate its performance by simulation. The results show that the modified bootstrap method (ABRE) gives reliable and rather conservative rank estimates when the sample size is sufficient for the number of variables studied (the sample size is at least five-fold the number of variables). Applying the method to various datasets suggests high phenotypic dimensionality in all cases. The analysis thus provides no evidence for absolute evolutionary constraints.











Similar content being viewed by others
References
Amemiya, Y., Anderson, T. W., & Lewis, P. A. W. (1990). Percentage points for a test of rank in multivariate components of variance. Biometrika, 77, 637–641.
Bennett, K. D. (1996). Determination of the number of zones in a biostratigraphical sequence. New Phytologist, 132, 155–170.
Blows, M. W. (2007). A tale of two matrices: Multivariate approaches in evolutionary biology. Journal of Evolutionary Biology, 20, 1–8.
Cattel, R. B. (1966). The scree test for the number of factors. Journal of Multivariate Behavioral Research, 1, 317–346.
Cheverud, J. M. (1988). A comparison of genetic and phenotypic correlations. Evolution, 42, 958–968.
Cheverud, J. M., Rutledge, J. J., & Atchley, W. R. (1983). Quatitative genetics of development: Genetic correlations among age-specific trait values and the evolution of ontogeny. Evolution, 37, 895–905.
Frontier, S. (1976). Etude de la decroissance des valeurs propres das une analyze en composantes principales: Comparison avec le modele de baton brise. Journal of Experimental Marine Biology and Ecology, 25, 67–75.
Guttman, L. (1954). Some necessary conditions for common factor analysis. Psychometrika, 19, 179–185.
Hansen, T. F., & Houle, D. (2008). Measuring and comparing evolutionary constraint in multivariate characters. Journal of Evolutionary Biology, 21, 1201–1219.
Hine, E., & Blows, M. W. (2006). Determining the effective dimensionality of the genetic variance–covariance matrix. Genetics, 173, 1135–1144.
Houle, D., Mezey, J., Galpern, P., & Carter, A. (2003). Automated measurement of Drosophila wings. BMC Evolutionary Biology, 3, 25.
Hutchison, D. W., & Cheverud, J. M. (1995). Fluctuating asymmetry in tamarin (Saguinus) cranial morphology: Intra- and interspecific comparisons between taxa with varying levels of genetic heterozygosity. Journal of Heredity, 86, 280–288.
Jackson, D. A. (1993). Stopping rules in principal components analysis: A comparison of heuristical and statistical approaches. Ecology, 74, 2204–2214.
Jorjani, H., Klei, L., & Emanuelson, U. (2003). A simple method for weighted bending of genetic (co)variance matrices. Journal of Dairy Science, 86, 677–679.
Keating, K. A., & Quinn, J. F. (1998). Estimating species richness: The Michaelis–Menten model revisited. Oikos, 81, 411–416.
Kenney-Hunt, J. P., Wang, B., Norgard, E. A., Fawcett, G., Falk, D., Pletscher, L. S., et al. (2008). Pleiotropic patterns of quantitative trait loci for 70 murine skeletal traits. Genetics, 178(4), 2275–2288.
Kirkpatrick, M. (2009). Patterns of quantitative genetic variation in multiple dimensions. Genetica, 136, 271–284.
Kirkpatrick, M., Lofsvold, D., & Bulmer, M. (1990). Analysis of the inheritance, selection and evolution of growth trajectories. Genetics, 124, 979–993.
Kirkpatrick, M., & Meyer, K. (2004). Direct estimation of genetic principal components: Simplified analysis of complex phenotypes. Genetics, 168, 2295–2306.
Lande, R. (1979). Quantitative genetic analysis of multivariate evolution applied to brain:body size allometry. Evolution, 33, 402–416.
Legendre, L., & Legendre, P. (1983). Numerical Ecology. Amsterdam; New York: Elsevier
Macarthur, R. H. (1957). On the relative abundance of bird species. Proceedings of the National Academy of Sciences of the United States of America, 43, 293–295.
McGuigan, K., & Blows, M. W. (2007). The phenotypic and genetic covariance structure of drosophilid wings. Evolution, 61, 902–911.
Meyer, K., & Kirkpatrick, M. (2005). Restricted maximum likelihood estimation of genetic principal components and smoothed covariance matrices. Genetics Selection Evolution, 37, 1–30.
Mezey, J. G., & Houle, D. (2005). The dimensionality of genetic variation for wing shape in Drosophila melanogaster. Evolution, 59, 1027–1038.
Pavlicev, M., Cheverud, J. M., & Wagner, G. P. (2009). Measuring morphological integration using eigenvalue variance. Evolutionary Biology, 36, 157–170.
Richtsmeier, J., Cheverud, J., & Lele, S. (1992). Advances in anthropological morphometrics. Annual Review of Anthropology, 21, 283–305.
Seguardo, P., & Araujo, M. B. (2004). An evaluation of methods for modeling species distributions. Journal of Biogeography, 31, 1555–1568.
Wagner, G. P. (1984). On the eigenvalue distribution of genetic and phenotypic dispersion matrices: Evidence for a nonrandom organization of quantitative character variation. Journal of Mathematical Biology, 21, 77–95.
Wagner, G. P., Kenney-Hunt, J. P., Pavlicev, M., Peck, J. R., Waxman, D., & Cheverud, J. M. (2008). Pleiotropic scaling of gene effects and the “Cost of complexity”. Nature, 452, 470–472.
Weber, K. E. (1990). Selection on wing allometry in Drosophila melanogaster. Genetics, 126, 975–989.
Acknowledgments
We thank D. Houle, C. Pelabon and A. Carter for the dataset on Drosophila wing, J. Kenney-Hunt for allowing the use of the dataset on mouse skeletal traits, and R. Probst for help in collecting Accipiter morphometric data. We also thank B. Walsh for constructive comments, and M. Blows and A. Carson for help in the application of SAS for the method comparison. MP was supported by Schrödinger Postdoctoral Fellowship from the Austrian Science Fund (FWF). JMC was supported by NSF grant BCS-0725068.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendix 1
Appendix 1
The Comparison of the Results of Rank Estimation by Factor Analysis (FA) and ABRE
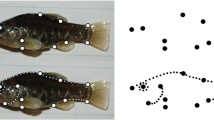
Fifteen subsets of six traits have been randomly drawn from the lateral dataset s of Drosophila wing data (A) and tamarin cranial data (B). The trait subsets 1L-15L designate the traits drawn from the left side, 1R-15R those on the right side. Both sides were drawn independently, thus, e.g., 2L doesn’t correspond to 2R trait combination. For each trait combination, the rank was estimated by FA and ABRE. Whereas FA requires multiple measurements on all individuals, ABRE was run in several versions, in which measurement error was estimated on different proportion of the total sample size. As the choice of individuals included in subsample can affect the estimate of error, we repeatedly drew 30 subsamples of the same size and the rank estimate was averaged across them (hence decimal values). ‘no conv.’ refers to the trait combinations in which FA did not reach convergence. Note that the results are consistent between FA and ABRE. The effect of estimating measurement error on low number of individuals in ABRE can be seen for 10 individuals, but is minimal for these phenotypic examples (see Table 2).
Rights and permissions
About this article
Cite this article
Pavlicev, M., Wagner, G.P. & Cheverud, J.M. Measuring Evolutionary Constraints Through the Dimensionality of the Phenotype: Adjusted Bootstrap Method to Estimate Rank of Phenotypic Covariance Matrices. Evol Biol 36, 339–353 (2009). https://doi.org/10.1007/s11692-009-9066-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11692-009-9066-7