Abstract
Heat shock transcription factors (Hsfs) have important roles during plant growth and development and responses to abiotic stresses. The identification and function of Hsf genes have been thoroughly studied in various herbaceous plant species, but not woody species, especially Phoebe bournei, an endangered, unique species in China. In this study, 17 members of the Hsf gene family were identified from P. bournei using bioinformatic methods. Phylogenetic analysis indicated that PbHsf genes were grouped into three subfamilies: A, B, and C. Conserved motifs, three-dimensional structure, and physicochemical properties of the PbHsf proteins were also analyzed. The structure of the PbHsf genes varied in the number of exons and introns. Prediction of cis-acting elements in the promoter region indicated that PbHsf genes are likely involved in responses to plant hormones and stresses. A collinearity analysis demonstrated that expansions of the PbHsf gene family mainly take place via segmental duplication. The expression levels of PbHsf genes varied across different plant tissues. On the basis of the expression profiles of five representative PbHsf genes during heat, cold, salt, and drought stress, PbHsf proteins seem to have multiple functions depending on the type of abiotic stress. This systematic, genome-wide investigation of PbHsf genes in P. bournei and their expression patterns provides valuable insights and information for further functional dissection of Hsf proteins in this endangered, unique species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Adverse environmental stresses such as cold, high temperature, drought, and salt can severely affect the seed germination, growth, development, yield, and quality of plants (Kerchev et al. 2020; Kowalczewski et al. 2020; de Medeiros et al. 2023). Phoebe bournei (Hemsl.) Yang, a member of the Lauraceae family, is a well-known garden tree and one of the dominant plants in broad-leaved evergreen forests of the subtropics, with a high economic value and ecological significance (Li et al. 2018; Chen et al. 2020a; Yang et al. 2020). Its timber was used for ancient palace construction and was thus a status symbol of the nobles (Ding et al. 2015, 2019; Jiao et al. 2022). In recent years, however, its growth has been hindered by human activities and abiotic stresses (Zhang et al. 2018; Li et al. 2022), resulting in a diminished population size and fragmented distribution. Currently, the species is listed as vulnerable by the IUCN and classified as a Class II key protected plant in China. Therefore, improving the tolerance of P. bournei so that it is better able to cope with abiotic stresses is of great importance.
Hsf is a class of transcription factors (TFs) involved in the regulation of plant defense response mechanisms under abiotic stress conditions (Guo et al. 2016). Although Hsf family members of different plant species exhibit obvious differences in protein sequence length, all share five well-conserved domains (Fig. 1A), including a conservative DNA-binding domain (DBD), an oligomerization domain (OD), also known as the HR-A/B region, a nuclear localization signal domain (NLS), activator motifs (AHAs), and a nuclear export signal domain (NES) (Guo et al. 2016; Zhang et al. 2018). The DBD is located close to the N-terminal and consists of three α-helices and four β-turns (Scharf et al. 2012), characterized by the central helix-turn-helix (H2-T-H3) motif that precisely combines the HSEs (heat shock elements) (Sakurai and Enoki 2010). The HR-A/B region comprises two hydrophobic heptad repeating regions, designated as A and B. According to the variations in the number of inserted amino acid residues present in the HR-A/B regions, plant Hsfs can be classed into three subfamilies: HsfA, HsfB, and HsfC (Nover et al. 2001). NLS and NES are located at the C-terminal of the Hsf members, which influence the subcellular localization, DNA binding, and activation functions of the Hsf. Members of HsfA possess an AHA motif, which mediates transcriptional activator function. In contrast, HsfB and HsfC members lack a motif, meaning that no self-activating function is present (Doring et al. 2000; Kotak et al. 2004). Intriguingly, the C-terminal of HsfB proteins harbors a well-conserved tetrapeptide repression domain (LFGV) (Fig. 1A) (Scharf et al. 2012).
Schematic diagram of the conserved structure of Hsf protein by subfamily and their distribution across plant species. A The three Hsf subfamilies (HsfA, HsfB, and HsfC). Rounded rectangles in different colors indicate conserved functional domains of Hsf proteins (adapted from Scharf et al. 2012). B The number of reported Hsf genes by subfamily in various plant species
In plants, the Hsf gene was first cloned in tomato (Solanum Lycopersicum) (Scharf et al. 1990). Since then, the Hsf gene family has been identified and extensively analyzed in many different species; for example, 56 Hsf genes have been found in wheat (Triticum aestivum), 52 in soybean (Glycine max), 30 in maize (Zea mays), and 21 in Arabidopsis (Arabidopsis thaliana) (Scharf et al. 2012). More importantly, the construction of several Hsf gene family databases has provided comprehensive information that has further aided comparative analyses of the Hsf gene family and provided clues to exploring their expression and evolutionary mechanisms (Ratheesh Kumar et al. 2012; Yu et al. 2022). Expression of Hsf genes is affected by various biotic and abiotic stress conditions, and the genes play a significant role in plant responses to stress (Dossa et al. 2016; Guo et al. 2016). As an example, HSFA6b in Arabidopsis is not only involved in a high-temperature stress response, but also acts as a positive regulator of the ABA signaling-mediated stress pathway, significantly improving the salt and drought tolerance of these plants (Huang et al. 2016). Studies have shown that most PvHsf genes in the common bean (Phaseolus vulgaris) are differentially expressed during temperature, salt, and heavy metal stress, indicating that PvHsfs may play specific functions depending on the specific abiotic stress (Zhang et al. 2022a). Additionally, HsfA2 may play an important role in physiological adaptation to spaceflight (Zupanska et al. 2019). The Hsf gene family is also involved in regulating the growth and development processes of plants (Lin et al. 2015). In tobacco (Nicotiana tabacum) over-expressing HsfA9, the accumulation of chlorophyll increases, thus promoting photosynthesis and influencing plant growth (Prieto-Dapena et al. 2017).
In contrast to annual herbaceous plants, trees are more susceptible to sustained damage from complex natural stresses due to their prolonged life cycle, and their genetic breeding and improvement are very challenging (Borthakur et al. 2022). Among five representative EgHsf genes in Eucalyptus grandis, three are upregulated genes and two downregulated at 4 °C and two upregulated and three downregulated genes at 40 °C (Yuan et al. 2022). When the seedlings were exposed to 200 mM NaCl, all five EgHsf genes were upregulated after 12 h and 24 h. These results suggest that Hsf genes play key roles in responding to abiotic stresses in woody plants. Thus, the functions of the Hsf gene family in P. bournei need to be determined. In 2022, the sequence of the genome of P. bournei at the chromosome level was published, accelerating functional genomic studies (Han et al. 2022).
In the present genome-wide investigation of Hsf genes in P. bournei using bioinformatic methods, we also assessed their responses to several abiotic stresses. To the best of our knowledge, this is the first report on the Hsf gene family in P. bournei, and the findings will provide valuable genetic resources for future studies of gene function and cloning to improve the stress resistance of P. bournei.
Materials and methods
Identification and analysis of Hsf proteins in P. bournei
The genomic data (cDNAs and proteins) for P. bournei were derived from the China National GeneBank DataBase (https://db.cngb.org/), and the BioProject accession number is CNP0002030 (Han et al. 2022). The PbHsf proteins were identified using two specific methods. (1) A hidden Markov model of the conserved DBD domain of the Hsf (PF00447) was obtained from the Pfam database (http://pfam.xfam.org/) (El-Gebali et al. 2018), and the Simple HMM Search module in TBtools was used to search the Hsf protein sequences of the P. bournei genome (Chen et al. 2020a). (2) We used 21 AtHsf protein sequences of Arabidopsis that were downloaded from the Plant Transcription Factor Database (PTFD) (http://planttfdb.gao-lab.org/index.php) (Jin et al. 2017) as Hsf query sequences for a BLASTp search (E-value < 1e − 5) of all P. bournei protein sequences. For the respective methods, we obtained 25 and 27 candidate sequences. These PbHsf candidates were then screened a DBD domain using the online website NCBI Conserved Domain Database (NCBI-CDD) (https://www.ncbi.nlm.nih.gov/cdd/) (Marchler-Bauer et al. 2016) and SMART (http://smart.embl-heidelberg.de/) to confirm that they are Hsf sequences.
Physicochemical properties and phylogenetic tree for PbHsf proteins
Physicochemical variables of 17 PbHsf proteins were predicted using the ExPASy ProtParam online tool (https://web.expasy.org/protparam/). Subcellular localizations were predicted using the WoLF PSORT website (https://wolfpsort.hgc.jp/) (Horton et al. 2007). We obtained 21 and 25 Hsf protein sequences in Arabidopsis and rice (Oryza sativa) from the PTFD database. Then the plant Hsf protein sequences were aligned using MUSCLE with default parameters (Edgar 2004). A phylogenetic tree of all proteins was constructed via MEGA 7 software, using the neighbor-joining (NJ) method, and the bootstrap value was set to 1000. The phylogenetic tree was visualized and post-processed with iTOL (https://itol.embl.de/itol.cgi) (Letunic and Bork 2021).
Motifs and gene structure analysis of PbHsf genes
The MEME Suite 5.4.1 online tool (http://meme-suite.org/) was used to identify the conservative motifs in PbHsf proteins, while the maximum number of motifs was set to 10 with the motif sites corresponding to a ZOOPS (Zero or One Occurrence Per Sequence) model (Bailey et al. 2009). The intron–exon distributions of PbHsf genes were obtained using GFF annotation files from the P. bournei genome. The motifs and gene structure were visualized using TBtools (Chen et al. 2020a).
Sequence alignment and three-dimensional structural of PbHsf proteins
The conserved domains of PbHsf protein sequences were edited using Jalview software (Waterhouse et al. 2009). SOPMA (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_sopma.html) was utilized for the protein secondary structure prediction, using default parameters. The SWISS-MODEL database (https://swissmodel.expasy.org/) was used to predict the protein tertiary structure using the homology modeling method.
Chromosomal location, gene duplication and synteny analysis of PbHsf genes
The distribution of PbHsf genes on the P. bournei genome chromosomes was mapped using the MG2C online site (http://mg2c.iask.in/mg2c_v2.1/). The genomes of five representative plants, namely Arabidopsis, rice, grape (Vitis vinifera), Populus trichocarpa, and E. grandis were downloaded from the NCBI database. Subsequently, gene duplication events, Ka/Ks ratios, and collinearity relationships with other plants were evaluated using TBtools (Chen et al. 2020a, 2022).
Cis-acting elements and the expression profiles of PbHsf genes
The upstream 2000-bp sequence of all PbHsf genes promoters were obtained by adding the P. bournei CDS file to the TBtools GFF3 Sequence Extractor submenu. Then, we uploaded the obtained 2000-bp sequence files to the PlantCARE online website (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) (Lescot et al. 2002) to predict cis-acting elements and visualized them using ChiPlot online tool (https://www.chiplot.online/). Transcriptome data of PbHsf genes in different tissues were downloaded from the NCBI, and the expression levels were shown in a cartoon-heatmap form (Chen et al. 2020c).
Plant materials and abiotic stress treatments
P. bournei seedlings, grown at 25 °C in a greenhouse with 75% relative humidity, with the same level of growth and free of pests and diseases were selected for abiotic stress treatment. Three biological replicates, each comprising two seedlings were used for each treatment: 10% PEG6000 for drought stress with a well-hydrated control group; 10% NaCl solution for salt stress with the control group in distilled water; heat stress in an incubator at 40 °C and cold stress at 10 °C, with the control group at room temperature (25 °C). P. bournei leaves were detached at 4, 6, 8, 12 and 24 h, immediately frozen in liquid N2, then stored at − 80 °C until used.
RNA extraction and RT-qPCR analysis
Total RNA was extracted from the leaf samples using a HiPure Plant RNA Mini Kit (Magen, Guangdong, China). Then, cDNA was synthesized using a PrimeScript RT Reagent Kit (Perfect Real Time) (TaKaRa, Dalian, China). The obtained cDNA was then used for RT-qPCR with specific primers for the PbHsf genes that were designed using the Primer 3 online tool (http://bioinfo.ut.ee/primer3-0.4.0/). PbEF1α gene was used as a reference control (Zhang et al. 2018). Primer sequences are given in Table S1. In quick succession, the expression of PbHsf genes in response to abiotic stresses was quantified using RT-qPCR with a CFX Real-Time PCR System (Bio-Rad, USA) and reaction solution comprising 1 μL of cDNA, 10 μL of SYBR Premix Ex TaqTM II, 2 μL of specific primers, and 7 μL of ddH2O. The thermocycling conditions were 95 °C for 30 s; 40 cycles at 95 °C for 5 s, 60 °C for 30 s, 95 °C for 5 s, 60 °C for 60 s, 50 °C for 30 s. Finally, the relative expression of each gene was calculated using the 2−ΔΔCT method (Livak and Schmittgen 2001) and visualized as a heatmap using TBtools (Chen et al. 2020a).
Result
Identification and physicochemical properties of PbHsf proteins
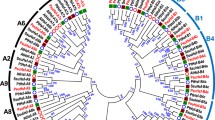
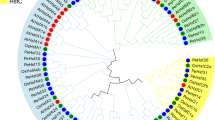
After deleting redundant sequences, we identified 17 Hsf genes from the P. bournei genome using bioinformatics methods (Fig. 2A). The PbHsf genes were renamed according to their phylogenetic relationship with the Hsf gene family in Arabidopsis and rice (Table 1 and Fig. 2C). The lengths of 17 PbHsf proteins varied from 214 aa (PbHsfA6a and PbHsfB3b) to 518 aa (PbHsfA8), and MW ranged from 24.49 kD (PbHsfB3b) to 58.18 kD (PbHsfA8). The theoretical pIs ranged from 4.78 (PbHsfA1a) to 10.05 (PbHsfB3b). The negative hydropathicity index of 17 PbHsf proteins suggested that they were hydrophilic. The aliphatic index ranged from 58.28 (PbHsfA4) to 83.63 (PbHsfA1b), and the instability index of PbHsf proteins ranged from 46.49 (PbHsfB3b) to 67.79 (PbHsfB3a). The subcellular localization predictions indicated that all of the PbHsf proteins were in the nucleus (Table 1).
Identification and evolutionary analysis of PbHsf genes in Phoebe bournei. A The two strategies employed in searching accurate PbHsf protein sequences. B Intraspecific evolutionary relationship of PbHsf genes. C Phylogenetic tree based on Hsf sequences from P. bournei (Pb), Arabidopsis (At), and rice (Os). Different colors represent different Hsf classes
Evolutionary analysis of PbHsf genes
MEGA 7 was used to analyze the protein sequences of PbHsfs through the neighbor-joining method. Based on the well-established classifications of 21 and 25 genes in Arabidopsis and rice Hsf families, respectively (Nover et al. 2001; Mittal et al. 2009; Scharf et al. 2012), the Hsf gene numbers in P. bournei were classified into subfamilies A, B, or C and classes A1, A4, A6, A8, B2, B3, B4, or C1 (Fig. 2C). Seven Hsf genes were in subfamily A and seven in B, and only three were in subfamily C (Fig. 2B). In this study, among the eight PbHsf classes, class B4 had the most P. bournei Hsf genes (four), and three Hsf genes each in classes A6 and C1. Unlike the genes in the AtHsf family, none of the PbHsf family belonged to classes A2, A3, A5, A7, A9, or B1. In addition, class C2 was exclusive to the OsHsf family (Fig. 2C).
Protein motif and gene structure analysis of PbHsf genes
Ten conserved motifs in PbHsf proteins were identified and analyzed with the MEME tool (Fig. 3B). Motifs 1, 2, and 6 constituted the DBD domain and were found in most PbHsfs. Motif 1 was present in all PbHsf proteins. Although motif 3 was observed in subfamilies A and C, it was not present in subfamily B. Motifs 5, 8, and 9 were unique to subfamily B. Motif 4 was found in subfamilies A and B (except for PbHsfA4 and PbHsfB4d), rather than subfamily C. Only subfamily A contained motif 10. The exons, introns, and UTR structure of PbHsf genes were analyzed using TBtools (Fig. 3C). The PbHsf genes had from two to seven exons, with the exception of PbHsfC1c, which only had one. PbHsfB4b had a maximum of seven exons. PbHsf genes varied containing between one and six introns, with 10 of 17 having only one intron. Notably, one gene in subfamily C, PbHsfC1c, did not have any introns. The untranslated region (UTR) sequences of PbHsf genes in P. bournei were incomplete. Specifically, nine PbHsf genes had both 5′ UTR and 3′ UTR sequences, and one gene (PbHsfA6b) only had a 5′ UTR. Conversely, seven PbHsf genes did not have any UTR sequences.
Characteristics of PbHsf genes in Phoebe bournei. A Evolutionary relationships among PbHsf genes. B The 10 motifs of PbHsf proteins, represented as rectangles with distinct colors. C Gene structure of PbHsfs. Cyan rectangle: CDS (coding sequence), orange rectangle: UTR (untranslated region); black line: intron
Multiple sequence alignment and three-dimensional structure prediction of PbHsf proteins
The protein sequences of the Hsf gene family contain five typically conserved domains: DBD, OD, NLS, AHA, and NES from the N-terminal to the C-terminal (Fig. 1A). Among them, the DBD is the most conservative domain, containing three α-helices and four β-turns (α1-β1-β2-α2-α3-β3-β4) (Fig. 4A). To further investigate protein structures of PbHsf genes, we used SOPMA and SWISS-MODEL to predict their secondary and tertiary structures. Random coils and α-helices were the most frequently occurring secondary structures (Table S2). The three-dimensional structure of PbHsf proteins was generated, resulting in similar templates obtained by comparison in the Protein Data Base (Fig. 4B). The model with the highest identity, GMQE, and QMEAN score was selected as the optimal and representative protein structure from the eight classes of PbHsfs (Table S3). Each protein from the different classes had a relatively symmetrical and highly similar structure, so we can infer that the PbHsf genes and the functions of the three subfamilies are similar at the protein structural level.
Chromosome location, collinearity, and Ka/Ks analysis of PbHsf genes
In the present study, 17 PbHsf genes were distributed across 10 chromosomes in P. bournei, with one to six genes present on each (Fig. 5). For instance, chromosome 1 harbored the most Hsf genes (six), with the three subfamilies represented. In contrast, only one gene was found on chromosome 2 (subfamily C); chromosomes 3, 7, and 10 (subfamily A); and chromosomes 8, 9, and 11 (subfamily B). In addition, two genes mapped chromosomes 6 (PbHsfB2/PbHsfB4b) and 12 (PbHsfB4d/PbHsfA4), respectively. Considering the results together, the distribution of PbHsf genes on chromosomes is relatively uneven.
Chromosome locations and collinearity of PbHsf genes in Phoebe bournei. chr: chromosome. A Distribution of PbHsf genes. P. bournei has 12 chromosomes; no PbHsf genes were mapped to chromosomes 4 and 5. B The green lines of the internal ring represent the density of genes per chromosome. Collinear genes are depicted by light grey lines; PbHsf gene pairs are shown as blue lines
To investigate the evolution of the Hsf genes, the tandem and fragment duplication gene pairs of PbHsf gene family were studied. As shown in Fig. 5, one tandem repeat (PbHsfB2/PbHsfB4b) was identified on chromosome 6 as described by Holub (2001), and seven fragment repeat events were found among PbHsf genes, which indicates that PbHsfs had seven pairs of collinearity (Fig. 5B and Table S4). The fragment repeat gene pairs, PbHsfA1a/PbHsfA1b, PbHsfA6a/PbHsfA6b, and PbHsfA1a/PbHsfA8 belonged to subfamily A, while PbHsfB4a/PbHsfB4a belonged to subfamily B. The three genes (PbHsfC1a, PbHsfC1b, and PbHsfC1c) in subfamily C are collinear with one another, forming three pairs with collinearity. Ka/Ks analysis indicated that PbHsf genes eliminated harmful mutations to maintain the level of purity selection (Table S4).
To analyze duplication and evolutionary processes for PbHsf genes, five plants were selected for comparative analysis of genome collinearity with P. bournei, including one monocotyledon (rice) and four dicotyledons (Arabidopsis, grape, P. trichocarpa, and E. grandis) (Fig. 6). A total of 22 PbHsf genes were collinear with those of P. trichocarpa, 16 with grape, 14 with E. grandis, nine with A. thaliana, and eight with rice, indicating a closer association between P. bournei and P. trichocarpa. These results clarified that the collinearity of the genome between P. bournei and dicotyledonous plants was greater than that between P. bournei and monocotyledonous plants.
Analysis of cis-acting elements in promoter of PbHsf genes
In the promoter region of the PbHsf genes, 32 cis-acting elements were identified (Fig. 7; Table S5). These cis-acting elements were found to be associated with hormones (nine), light (nine), plant growth and development (five), and stress (nine). The MYC element was detected in 17 PbHsf genes, and was the most frequently found cis-acting element (Fig. 7). Other cis-acting elements, found in 15 PbHsf genes, included ABRE, G-box, ARE, and WRE3. The most frequent type of element was the hormone-response type (191), including abscisic acid (ABRE), MeJA (CGTCA-motif and TGACG-motif), gibberellin (GARE-motif, MYC, P-box, and TATC-box), salicylic acid (TCA-element), auxin (TGA-element) (Table S5). The stress-response type, such as defense and stress (STRE and TC-rich repeats), low temperature (LTR), and anaerobic induction (ARE), was the second most frequent (175). A considerable number of cis-acting elements were light-response (152) or growth-and-development types (71) (Table S5).
Different tissue-specific expression analysis of PbHsf genes
To explore the functional divergence of Hsf genes, tissue-specific expression levels of PbHsf genes in root bark, root xylem, stem bark, stem xylem, and leaf were obtained from the NCBI database (Fig. 8; Table S6). In general, the expression profiles of PbHsf genes in the bark and xylem tissues of the stem were similar to those of the root, except for PbHsfA6c, which was highly expressed in stem bark. Additionally, six of 17 PbHsf genes, including PbHsfA1a, PbHsfA6c, and PbHsfC1c, were highly expressed in leaves. On the contrary, the expression of PbHsf genes of subfamily B was low in leaves. Five of the PbHsf genes were barely expressed in any of the tissues, especially PbHsf3B, with a notable lack of expression within the five tissues. Thus, the expression levels of the different genes differed significantly among the different tissues.
The expression profiles of PbHsf genes under abiotic stresses
To investigate the responses of PbHsf genes to drought, salt, or temperature stress, five representative genes (PbHsfA1a, PbHsfA6c, PbHsfB3a, PbHsfB4d, and PbHsfC1b) from three subfamilies were selected for RT-qPCR observation (Fig. 9; Table S7). Among them, PbHsfA1a, PbHsfA6c, and PbHsfC1b were highly expressed in leaves, whereas expression of PbHsfB3a and PbHsfB4d was very low (Fig. 8). Moreover, all five representative genes had a high number of stress-related cis-acting elements (Fig. 7). Under drought treatment (10% PEG6000), PbHsfA1a and PbHsfA6c were upregulated after 4, 6, and 8 h, while PbHsfB3a was upregulated after 12 and 24 h of stress. Compared with the control group (0 h), PbHsfB4d was downregulated within all sampling periods, and expression declined as the treatment duration increased (Fig. 9).
Expression of five representative PbHsf genes in seedlings of Phoebe bournei response to drought, salt or temperature stress. A Phenotypic changes in seedlings after 0, 4, 6, 8, 12, and 24 h of treatment. Samples collected at 0 h were used as the controls. Scale bars = 40 mm. B Relative expression of PbHsf genes determined by RT-qPCR. Blue: downregulated genes; red: upregulated genes
In the salt treatment (10% NaCl solution), expression of PbHsfA1a, PbHsfA6c, and PbHsfC1b was low and were downregulated throughout all sampling periods. In contrast, the expression of PbHsfB3a increased significantly after 4 h and continued to increase, suggesting a potential role for PbHsfB3a as a key gene involved in the response to salt stress. In addition, the expression of PbHsfB4d was upregulated only at 4 h and downregulated at other periods (Fig. 9). During cold stress (10 °C), the expression of all five genes was upregulated and peaked at 12 h and. However, there were slight differences in expression among PbHsf genes during the other sampling periods. Interestingly, the expression of all five PbHsf genes were significantly downregulated during the heat stress (40 °C), especially PbHsfB3a and PbHsfB4d, which was severely downregulated (Fig. 9).
Discussion
The widely distributed and specific Hsf transcription factors in plants have a conserved DBD domain, which has important roles in plant growth, development, and stress responses (von Koskull-Döring et al. 2007; Andrási et al. 2020). Extensive studies of the Hsf gene family in many plants, especially woody plants, have benefited from the increasing availability of genome sequences. For example, 21 Hsf genes have been reported for Arabidopsis (21) (Nover et al. 2001; Scharf et al. 2012), 25 for rice (Mittal et al. 2009), 25 for apple (Malus pumila) (Giorno et al. 2012), 30 for poplar (Populus) (Zhao et al. 2023), 22 for pineapple (Ananas comosus) (Wang et al. 2021), and 36 for E. grandis (Yuan et al. 2022) (Fig. 1B). However, a comprehensive investigation of the Hsf gene family in P. bournei has not yet been published.
In the present study, we identified 17 Hsf genes from the P. bournei genome (Fig. 2), which were unevenly distributed on 10 of 12 P. bournei chromosomes, with the highest gene number comprising six PbHsfs, detected on chromosome 1 (Fig. 5). In angiosperms, three subfamilies (A, B, C) of the Hsf gene family consist of seven, five, and two classes, respectively (A1–A9, B1–B5, and C1–C2) (Wang et al. 2018). As previously mentioned, the HsfC2 class exists only in monocotyledons and is absent in dicotyledons (Wang et al. 2018). The HsfC2 class was not found in P. bournei and Arabidopsis, and two members of OsHsf belonged to the HsfC2 class in rice (Fig. 2C). Therefore, the Hsf gene family in monocotyledons generally have more subfamily C members than in dicotyledons (Fig. 1B). Interestingly, class B5 is absent from all three of the aforementioned plants, suggesting that it is relatively rare. The physicochemical properties of PbHsf proteins vary greatly, but they are all located in the nucleus (Table 1) as found for poplar (Zhao et al. 2023) and E. grandis (Yuan et al. 2022).
PbHsf protein members that are closely related in the phylogenetic tree had common motif arrangements and compositions (Fig. 3). Motifs 1, 2, and 6, constituting the DBD domain, were present in almost all PbHsfs. The DBD domain was the most conserved, with three α-helices and four β-turns (α1-β1-β2-α2-α3-β3-β4) (Scharf et al. 2012; Guo et al. 2016) (Fig. 4A). Similarly, Hsf genes in three subfamilies had similar motifs in Populus (Zhang et al. 2015) and peach (Prunus persica) (Tan et al. 2021). In plants, introns mediate gene expression regulation (Le Hir et al. 2003; Shaul 2017). Therefore, it is of great significance to elucidate gene function by analyzing the intron structure. The gene structure analysis of PbHsf members revealed that all PbHsf genes but one contained at least one or more introns; PbHsfC1c had none (Fig. 3C). However, the intron length of Hsf genes varied to certain degrees in each subfamily, and the structure of each member differed significantly, which further affected their expression in plant tissues (Fig. 8).
Gene duplication events and collinearity relationships can help reveal gene expansion and evolution of the PbHsf gene family (Cannon et al. 2004). Notably, only one tandem duplication event was found in PbHsf genes, but segmental duplication had occurred in seven gene pairs, involving 10 PbHsf genes and six chromosomes. These results suggest that fragment replication events may a driving force for the expansion of the Hsf gene family in P. bournei. Subsequently, we calculated the Ka/Ks ratios of seven PbHsf gene pairs and found that they were all less than one (Table S4), indicating purified selection pressure during the evolutionary process. Comparable results have been reported for poplar (Zhao et al. 2023) and maize (Lin et al. 2011).
Cis-acting elements are critical in multiple biological signal transduction and stress responses (Ibraheem et al. 2010). In this study, many cis-acting elements were identified in the promoter regions of PbHsf genes and were associated with hormone regulation and stress response (Fig. 7). Gibberellin (MYC, P-box, etc.), which is important throughout the life of flowering plants, promotes seed germination and the division and the elongation of root cells (Sun 2011). Abscisic acid (ABRE) participates in plant stress response and other physiological processes such as bud dormancy and osmotic regulation (Chen et al. 2020b). There may be a link between hormone regulation and response to abiotic stress: for example, in PbHsfB3a, the promoter region had six AREs, six STREs, three ABREs, and two MYCs (Fig. 7), and more importantly, the gene was upregulated during drought, salt, and cold stress (Fig. 9). However, the relationship of the gene to stress responses is uncertain, and further research is required. In addition, the presence of the G-box, GATA motif, and GT1 motif indicates that the PbHsf genes might be involved in light responses (Fig. 7).
The various combinations of stresses, particularly abiotic stresses, that affect plants alter the expression of different Hsf gene members (Nishizawa et al. 2006; Sakurai and Enoki 2010). For instance, drought, salinity, and heat stresses alter the expression of CarHsf genes in chickpea (Cicer arietinum) (Chidambaranathan et al. 2018), and stresses such as powdery mildew, drought, salt, and hormone treatments distinctly change the expression of 17 FvHsf genes in strawberry (Hu et al. 2015). In our RT-qPCR assays to investigate the expression profiles of five representative PbHsf genes after drought, salt, cold, or heat treatments, we found that PbHsfA1a and PbHsfA6c respond early to drought treatment, while PbHsfB3a did not increase until 24 h. In addition, PbHsfC1b responded positively to drought treatment (Fig. 9).
Enhancing the salt tolerance of woody plants is a research hotspot as soil salinization continues to increase around the world (Polle and Chen 2015; Guo et al. 2021, 2023). As part of this goal, Guo et al. (2023) used transcriptome data of poplar under salt stress to find that a key Hsf gene, PtHsf21, is upregulated in leaves and stems (Guo et al. 2023). Among five representative PbHsf genes in P. bournei, the performance of PbHsfB3a expression was about 13-fold higher than the control group after 4 h of slat stress and continued to be upregulated (Table S7 and Fig. 9). Presumably, PbHsfB3a carries out significant function during salt stress.
Plant growth and development are also very sensitive to temperature; even small changes can cause remarkable responses in plants (Penfield 2008). Among five randomly selected PeHSF genes from passion fruit (Passiflora edulis), the expression of all PeHSF genes under heat (45 °C) and osmotic (mannitol) stress increased at 12 h of treatment (Wang et al. 2023). In the present study, PbHsf genes in P. bournei had diverse expression responses to cold and heat stresses (Fig. 9), similar to changes in the expression profiles in other plants, illustrating that they are characterized by similar cell signaling and cellular processes. Accumulating evidence thus suggests that the Hsf gene family, as a core part of the signal transduction, plays a key role in plant coordination and adaptation to numerous abiotic stresses (Fig. 10) (Scharf et al. 2012; Guo et al. 2016).
Working model for the role of Hsf transcription factor during abiotic stresses and schematic diagram of plant cell response to stresses (modified from Guo et al. 2016; Zhang et al. 2022b; Ali et al. 2023). The abiotic stress induces Ca2+ concentration increase, ROS accumulation, and protein degradation, and subsequently signal transmission to the nucleus. The Hsf then specifically binds to the heat shock element (HSE) in the promoter region of the Hsf gene and activates the expression of stress-inducible genes, thereby enhancing stress tolerance in plants. ABA, abscisic acid; SOS, salt overly sensitive; CaM, Ca2+-calmodulin; ROS, reactive oxygen species; TFs, transcription factors; HSF, heat shock transcription factor; HSP, heat shock protein; HSE, heat shock element
Conclusions
In our genome wide investigation of the Hsf gene family of P. bournei, we identified 17 PbHsf genes, which were classified among three subfamilies and had considerable evolutionary divergence. Tissue-specific expression profiles of the genes indicated potentially divergent functions in the growth and development of P. bournei. The analysis of cis-acting elements and gene expression patterns during various stresses suggest that the PbHsf genes are involved in responses to diverse abiotic stresses. Our research provides a basis for the further functional characterization of PbHsf genes and eventually increasing the yield and quality of P. bournei and other woody plants.
References
Ali A, Petrov V, Yun DJ, Gechev T (2023) Revisiting plant salt tolerance: novel components of the SOS pathway. Trends Plant Sci 28(9):1060–1069. https://doi.org/10.1016/j.tplants.2023.04.003
Andrási N, Pettkó-Szandtner A, Szabados L (2020) Diversity of plant heat shock factors: regulation, interactions, and functions. J Exp Bot 72(5):1558–1575
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME Suite: tools for motif discovery and searching. Nucleic Acids Res 37(suppl_2):W202–W208
Borthakur D, Busov V, Cao XH, Du QZ, Gailing O, Isik F, Ko JH, Li CH, Li QZ, Niu SH, Qu GZ, Vu THG, Wang XR, Wei ZG, Zhang L, Wei HR (2022) Current status and trends in forest genomics. For Res 2:11. https://doi.org/10.48130/FR-2022-0011
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4(1):10
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020a) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202
Chen K, Li GJ, Bressan RA, Song CP, Zhu JK, Zhao Y (2020b) Abscisic acid dynamics, signaling, and functions in plants. J Integr Plant Biol 62(1):25–54
Chen SP, Sun WH, Xiong YF, Jiang YT, Liu XD, Liao XY, Zhang DY, Jiang SZ, Li Y, Liu B, Ma L, Yu X, He L, Liu B, Feng JL, Feng LZ, Wang ZW, Zou SQ, Lan SR, Liu ZJ (2020c) The Phoebe genome sheds light on the evolution of magnoliids. Hortic Res 7(1):146
Chen CJ, Wu Y, Xia R (2022) A painless way to customize Circos plot: From data preparation to visualization using TBtools. iMeta 1(3):35
Chidambaranathan P, Jagannadham PTK, Satheesh V, Kohli D, Basavarajappa SH, Chellapilla B, Kumar J, Jain PK, Srinivasan R (2018) Genome-wide analysis identifies chickpea (Cicer arietinum) heat stress transcription factors (Hsfs) responsive to heat stress at the pod development stage. J Plant Res 131(3):525–542
de Medeiros RLS, de Paula RC, de Souza JVO, Pedro J (2023) Abiotic stress on seed germination and plant growth of Zeyheria tuberculosa. J For Res. https://doi.org/10.1007/s11676-023-01608-3
Ding YJ, Zhang JH, Lu YF, Ep L, Lou LH, Tong ZK (2015) Development of EST-SSR markers and analysis of genetic diversity in natural populations of endemic and endangered plant Phoebe chekiangensis. Biochem Syst Ecol 63:183–189
Ding X, Xiao JH, Li L, Conran JG, Li J (2019) Congruent species delimitation of two controversial gold-thread nanmu tree species based on morphological and restriction site-associated DNA sequencing data. J Syst Evol 57(3):234–246
Doring P, Treuter E, Kistner C, Lyck R, Chen A, Nover L (2000) The role of AHA motifs in the activator function of tomato heat stress transcription factors HsfA1 and HsfA2. Plant Cell 12(2):265–278
Dossa K, Diouf D, Cissé N (2016) Genome-wide investigation of Hsf genes in Sesame reveals their segmental duplication expansion and their active role in drought stress response. Front Plant Sci 7:1522
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797
El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, Qureshi M, Richardson LJ, Salazar GA, Smart A, Sonnhammer ELL, Hirsh L, Paladin L, Piovesan D, Tosatto SCE, Finn RD (2018) The Pfam protein families database in 2019. Nucleic Acids Res 47(D1):D427–D432. https://doi.org/10.1093/nar/gky995
Giorno F, Guerriero G, Baric S, Mariani C (2012) Heat shock transcriptional factors in Malus domestica: identification, classification and expression analysis. BMC Genomics 13(1):639
Guo M, Liu JH, Ma X, Luo DX, Gong ZH, Lu MH (2016) The plant heat stress transcription factors (HSFs): Structure, regulation, and function in response to abiotic stresses. Front Plant Sci 7:114
Guo Q, Jiang JH, Yao WJ, Li L, Zhao K, Cheng ZH, Han LB, Wei R, Zhou BR, Jiang TB (2021) Genome-wide analysis of poplar HD-zip family and over-expression of PsnHDZ63 confers salt tolerance in transgenic Populus simonii × P. nigra. Plant Sci 311:111021. https://doi.org/10.1016/j.plantsci.2021.111021
Guo Q, Wei R, Xu M, Yao WJ, Jiang JH, Ma XJ, Qu GZ, Jiang TB (2023) Genome-wide analysis of HSF family and overexpression of PsnHSF21 confers salt tolerance in Populus simonii×P. nigra. Front Plant Sci 14:1160102
Han X, Zhang JH, Han S, Chong SL, Meng GL, Song MY, Wang Y, Zhou SC, Liu CC, Lou LH, Lou XZ, Cheng LJ, Lin E, Huang HH, Yang Q, Tong ZK (2022) The chromosome-scale genome of Phoebe bournei reveals contrasting fates of terpene synthase (TPS)-a and TPS-b subfamilies. Plant Commun 3(6):100410
Holub EB (2001) The arms race is ancient history in Arabidopsis, the wildflower. Nat Rev Genet 2:516–527
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35(suppl_2):W585–W587
Hu Y, Han YT, Wei W, Li YJ, Zhang K, Gao YR, Zhao FL, Feng JY (2015) Identification, isolation, and expression analysis of heat shock transcription factors in the diploid woodland strawberry Fragaria vesca. Front Plant Sci 6:736
Huang YC, Niu CY, Yang CR, Jinn TL (2016) The heat stress factor HSFA6b connects ABA signaling and ABA-mediated heat responses. Plant Physiol 172(2):1182–1199
Ibraheem O, Botha CEJ, Bradley G (2010) In silico analysis of cis-acting regulatory elements in 5’ regulatory regions of sucrose transporter gene families in rice (Oryza sativa Japonica) and Arabidopsis thaliana. Comput Biol Chem 34(5):268–283
Jiao LC, Lu Y, Zhang M, Chen YP, Wang ZS, Guo Y, Xu C, Guo J, He T, Ma LY, Gao WQ, Wang J, Zhou SL, Zhang YG, Jiang XM, Baas P, Yin YF (2022) Ancient plastid genomes solve the tree species mystery of the imperial wood “Nanmu” in the Forbidden City, the largest existing wooden palace complex in the world. Plants People Planet 4(6):696–709
Jin JP, Tian F, Yang DC, Meng YQ, Kong L, Luo JC, Gao G (2017) PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res 45(D1):D1040–D1045
Kerchev P, van der Meer T, Sujeeth N, Verlee A, Stevens CV, Van Breusegem F, Gechev T (2020) Molecular priming as an approach to induce tolerance against abiotic and oxidative stresses in crop plants. Biotechnol Adv 40:107503
Kotak S, Port M, Ganguli A, Bicker F, Von Koskull-Döring P (2004) Characterization of C-terminal domains of Arabidopsis heat stress transcription factors (Hsfs) and identification of a new signature combination of plant class A Hsfs with AHA and NES motifs essential for activator function and intracellular localization. Plant J 39(1):98–112
Kowalczewski PŁ, Radzikowska D, Ivanišová E, Szwengiel A, Kačániová M, Sawinska Z (2020) Influence of abiotic stress factors on the antioxidant properties and polyphenols profile composition of green barley (Hordeum vulgare L.). Int J Mol Sci 21(2):397
Le Hir H, Nott A, Moore MJ (2003) How introns influence and enhance eukaryotic gene expression. Trends Biochem Sci 28(4):215–220
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49(W1):W293–W296
Li YG, Liu XH, Ma JW, Zhang XM, Xu LA (2018) Phenotypic variation in Phoebe bournei populations preserved in the primary distribution area. J Forestry Res 29:35–44
Li X, Liu LL, Sun SX, Li YM, Jia L, Ye SL, Yu YX, Dossa K, Luan YP (2022) Leaf-transcriptome profiles of phoebe bournei provide insights into temporal drought stress responses. Front Plant Sci 13:1010314
Lin YX, Jiang HY, Chu ZX, Tang XL, Zhu SW, Cheng BJ (2011) Genome-wide identification, classification and analysis of heat shock transcription factor family in maize. BMC Genomics 12(1):76
Lin Q, Jiang Q, Lin JY, Wang DL, Li SJ, Liu CR, Sun CD, Chen KS (2015) Heat shock transcription factors expression during fruit development and under hot air stress in Ponkan (Citrus reticulata Blanco cv. Ponkan) fruit. Gene 559(2):129–136
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25(4):402–408
Marchler-Bauer A, Bo Y, Han LY, He JE, Lanczycki CJ, Lu SN, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Lu F, Marchler GH, Song JS, Thanki N, Wang ZX, Yamashita RA, Zhang DC, Zheng CJ, Geer LY, Bryant SH (2016) CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res 45(D1):D200–D203
Mittal D, Chakrabarti S, Sarkar A, Singh A, Grover A (2009) Heat shock factor gene family in rice: genomic organization and transcript expression profiling in response to high temperature, low temperature and oxidative stresses. Plant Physiol Biochem 47(9):785–795
Nishizawa A, Yabuta Y, Yoshida E, Maruta T, Yoshimura K, Shigeoka S (2006) Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J 48(4):535–547
Nover L, Bharti K, Döring P, Mishra S, Ganguli A, Scharf KD (2001) Arabidopsis and the heat stress transcription factor world: how many heat stress transcription factors do we need? Cell Stress Chaperones 6(3):177–189
Penfield S (2008) Temperature perception and signal transduction in plants. New Phytol 179(3):615–628
Polle A, Chen S (2015) On the salty side of life: molecular, physiological and anatomical adaptation and acclimation of trees to extreme habitats. Plant Cell Environ 38:1794–1816
Prieto-Dapena P, Almoguera C, Personat JM, Merchan F, Jordano J (2017) Seed-specific transcription factor HSFA9 links late embryogenesis and early photomorphogenesis. J Exp Bot 68(5):1097–1108
Ratheesh Kumar R, Nagarajan NS, Arunraj SP, Devanjan S, Rajan VBV, Esthaki VK, D’Silva P (2012) HSPIR: a manually annotated heat shock protein information resource. Bioinformatics 28(21):2853–2855
Sakurai H, Enoki Y (2010) Novel aspects of heat shock factors: DNA recognition, chromatin modulation and gene expression. FEBS J 277(20):4140–4149
Scharf KD, Rose S, Zott W, Schöffl F, Nover L, Schöff F (1990) Three tomato genes code for heat stress transcription factors with a region of remarkable homology to the DNA-binding domain of the yeast HSF. EMBO J 9(13):4495–4501
Scharf KD, Berberich T, Ebersberger I, Nover L (2012) The plant heat stress transcription factor (Hsf) family: structure, function and evolution. Biophys Acta Gene Regul Mech 2:104–119
Shaul O (2017) How introns enhance gene expression. Int J Biochem Cell Biol 91:145–155
Sun TP (2011) The molecular mechanism and evolution of the GA-GID1-DELLA signaling module in plants. Curr Biol 21(9):R338–R345
Tan B, Yan L, Li HN, Lian XD, Cheng J, Wang W, Zheng XB, Wang XB, Li JD, Ye X, Zhang LL, Li ZQ, Feng JC (2021) Genome-wide identification of HSF family in peach and functional analysis of PpHSF5 involvement in root and aerial organ development. PeerJ 9:e10961
von Koskull-Döring P, Scharf K-D, Nover L (2007) The diversity of plant heat stress transcription factors. Trends Plant Sci 12(10):452–457
Wang XM, Shi X, Chen SY, Ma C, Xu SB (2018) Evolutionary origin, gradual accumulation and functional divergence of heat shock factor gene family with plant evolution. Front Plant Sci 9:71
Wang LL, Liu YH, Chai MN, Chen HH, Aslam M, Niu XP, Qin Y, Cai HY (2021) Genome-wide identification, classification, and expression analysis of the HSF gene family in pineapple (Ananas comosus). PeerJ 9:e11329
Wang LL, Liu YH, Chai GF, Zhang D, Fang YY, Deng K, Aslam M, Niu XP, Zhang WB, Qin Y, Wang XM (2023) Identification of passion fruit HSF gene family and the functional analysis of PeHSF-C1a in response to heat and osmotic stress. Plant Physiol Biochem 200:107800
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ (2009) Jalview Version-a multiple sequence alignment editor and analysis workbench. Bioinformatics 25(9):1189–1191
Yang ZJ, Wu XH, Grossnickle SC, Chen LH, Yu XX, El-Kassaby YA, Feng JL (2020) Formula fertilization promotes Phoebe bournei robust seedling cultivation. Forests 11(7):781
Yu T, Bai Y, Liu Z, Wang ZY, Yang QH, Wu T, Feng SY, Zhang Y, Shen SQ, Li Q, Gu LQ, Song XM (2022) Large-scale analyses of heat shock transcription factors and database construction based on whole-genome genes in horticultural and representative plants. Hortic Res. https://doi.org/10.1093/hr/uhac035
Yuan T, Liang JX, Dai JH, Zhou XR, Liao WH, Guo ML, Aslam M, Li SB, Cao GQ, Cao SJ (2022) Genome-wide identification of Eucalyptus heat shock transcription factor family and their transcriptional analysis under salt and temperature stresses. Int J Mol Sci 23(14):8044
Zhang J, Liu BB, Li JB, Zhang L, Wang Y, Zheng HQ, Lu MZ, Chen J (2015) Hsf and Hsp gene families in Populus: genome-wide identification, organization and correlated expression during development and in stress responses. BMC Genomics 16(1):181
Zhang JH, Zhu YJ, Pan Y, Huang HH, Li CL, Li GZ, Tong ZK (2018) Transcriptomic profiling and identification of candidate genes in two Phoebe bournei ecotypes with contrasting cold stress responses. Trees 32(5):1315–1333
Zhang HM, Zhu JH, Gong ZZ, Zhu JK (2022a) Abiotic stress responses in plants. Nat Rev Genet 23(2):104–119
Zhang Q, Geng J, Du YL, Zhao Q, Zhang WJ, Fang QX, Yin ZG, Li JH, Yuan XK, Fan YR, Cheng X, Du JD (2022b) Heat shock transcription factor (Hsf) gene family in common bean (Phaseolus vulgaris): genome-wide identification, phylogeny, evolutionary expansion and expression analyses at the sprout stage under abiotic stress. BMC Plant Biol 22(1):33
Zhao K, Dang H, Zhou LD, Hu J, Jin X, Han YZ, Wang SJ (2023) Genome-wide identification and expression analysis of the HSF gene family in Poplar. Forests 14(3):510
Zupanska AK, LeFrois C, Ferl RJ, Paul AL (2019) HSFA2 functions in the physiological adaptation of undifferentiated plant cells to spaceflight. Int J Mol Sci 20(2):390
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Project funding: This study was supported by the Fujian Province Seed Industry Innovation and Industrialization Project “Innovation and Industrialization Development of Precious Tree Seed Industries (Phoebe bornei)” (ZYCX-LY-202102), and the Sub-project of National Key R&D Program “Phoebe bornei Efficient Cultivation Technology” (2016YFD0600603-2).
The online version is available at http://www.springerlink.com
Corresponding editor: Tao Xu.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liao, W., Tang, X., Li, J. et al. Genome wide investigation of Hsf gene family in Phoebe bournei: identification, evolution, and expression after abiotic stresses. J. For. Res. 35, 11 (2024). https://doi.org/10.1007/s11676-023-01661-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11676-023-01661-y