Abstract
The aim of this study is to determine the genetic variability and differentiation among populations of pedunculate oak (Quercus robur) at the eastern margin of the species distribution range. We applied new set of 95 geographically informative nuclear SNP (single nucleotide polymorphism) loci developed using Double Digest Restriction Site Associated DNA (ddRAD), a new generation DNA sequencing technology. The study area is located in the basins of the Volga River and its two main tributaries, Kama and Belaya. Despite the strong reduction and fragmentation of pedunculate oak forests over the last several centuries and comparatively small sizes of present stands, a relatively similar genetic diversity of seven populations was observed (on average, allelic diversity υа = 1.472–1.603, observed heterozygosity HO = 0.305–0.358, expected heterozygosity HE = 0.272–0.343). The genetic distances between populations vary between 0.102 and 0.170 (on average, d0 = 0.128). We have identified the absence of a spatial genetic structure and the existence of genetically peculiar populations in the territory studied. The genetic distances of Gregorius between populations vary between 0.102 and 0.170 (on average, d0 = 0.128). Genetic differentiation among populations was statistically significant with a mean of the fixation index FST of 0.075. Populations from the Privolzhye Upplands and from northern parts of the study area make the most contribution to overall inter-population differentiation (d0 = 0.137). No significant differences exist among the samples at lowlands along the Volga river (d0 = 0.110). It was concluded that the multiplicity and different ages of local populations and the complexity of migration routes of the pedunculate oak from Pleistocene refugia and secondary Holocene shelters can cause this phenomenon.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The pedunculate oak (Quercus robur L.) is one of the most common European broad-leaved tree species and of considerable ecological and economic significance (Ducousso and Bordacs 2004). Population diversity and differentiation of the species, gene flow, mating system, the phylogeography and impact of postglacial re-colonization and human activities on the gene pool has been studied extensively over the western part of its range using various modern gene markers (König et al. 2002; Petit et al. 2002a; Lepoittevin et al. 2015; Leroy et al. 2020). The vast eastern part of its range, which covers a large area of the territory of the former Soviet Union up to the Ural Mountains (Fig. 1), has been little investigated and only a few studies have been published (Buschbom et al. 2011; Degen et al. 2019). There are several significant reasons for closing this gap.
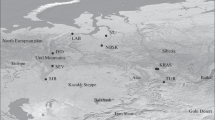
Natural distribution of Q. robur (after www.euforgen.org.); the study area is indicated by the red ellipse and the sampled stands
First, gene markers of chloroplast and mitochondrial DNA make it possible to identify colonization routes of the pedunculate oak out of glacial refugia to the territory of Central and Western Europe (Petit et al. 2002b). The distribution of some haplotypes in populations from Romania to Russia witness the eastern movement of the species, which initially located in the eastern Balkan refugium. But fossil pollen studies (Neyshtadt 1957) and morphological variation (Semerikov 1986) provide evidence of other primary and secondary refugia of Pleistocene and Holocene origins on the Russian Plain. The impact of the past gene flow from western and eastern sources on the formation of gene pools of local populations of Q. robur in this vast territory has not been studied.
Secondly, considerable reduction in the area of the species has happened in the eastern part of its range due in part to agriculture expansion (Kozharinov and Borisov 2012), and because of climate change, extreme cold winters, insect attacks, and phytopathogens (Tsaralunga et al. 2015; Zaurbekov et al. 2018). Genetic studies can be useful for estimating the effect of forest fragmentation on the genetic pool of pedunculate oak populations in changed landscapes.
Thirdly, populations on the margins of a species range are likely to be of great importance in revealing their ability to survive under extreme conditions (Buschbom et al. 2011), for example, in increasingly dry and exposed conditions in the transition zone to the Pontic steppes in Russia. But actual data on the genetic diversity of the eastern populations of the species are scarce despite the importance of this information in understanding of the species adaptation processes.
Finally, information on population genetic diversity and differentiation of forest tree species is one of the main baselines for forest management (McKenney et al. 2009; Ivetić et al. 2016).
This paper provides results on levels of genetic diversity and differentiation among populations of pedunculate oak on its eastern range using a new set of 95 SNP (single nucleotide polymorphism) loci developed using Double Digest Restriction Site Associated DNA (ddRAD), a new generation DNA sequencing technology (Peterson et al. 2012). We considered two main questions: (1) How different are the levels of genetic diversity in populations of the species on its eastern range after long periods of reduction and fragmentation? (2) How does the spatial and genetic structure of pedunculate oak populations in this area correspond to available paleogeographic data and investigations of morphological traits?
Materials and methods
Study area and sampling
The study area is located on the eastern margins of the Russian Plain and covers the basins of the Volga River and Kama and Belaya tributaries (Leummens 2016). It consists mostly of flat lowlands separated by two plateaus, Privolzhye to the west and General Syrt to the east. The eastern margin of the species range is in this territory (Fig. 1). Seven natural stands were selected in five vast regions of Russia (Table 1), representing steppe (Volgograd oblast) and forest-steppe (another four oblasts and Republics) zones. Mature trees over 100 years at least 50 m apart in each stand were sampled to minimize possible familial genetic clustering. Sampled material consisted of dried cambium of 10 trees per stand. Willing et al. (2012) showed that using a relatively large number of SNP loci makes it possible to correctly determine the levels of genetic diversity of populations and their differentiation in samples of relatively small numbers.
Laboratory procedures and data analysis
Genomic DNA was isolated according to Dumolin et al. (1995). DNA quality and quantity were estimated on a NanoDrop system (Thermo Fisher Scientific, Schwerte, Germany). A ddRAD (Peterson et al. 2012), a new generation DNA sequencing technology, was applied by the company Floragenex (Portland, OR, USA) to reveal SNPs in the nuclear genome (Blanc-Jolivet et al. 2020). Among the 26,074 detected variants, 3648 loci with a minimum flanking region of 50 bp around the SNP and a maximum of two neighbor SNPs were used for following examination. Loci with more than 10% messiness and allele frequencies less than 0.001 were removed. Discriminant analysis was carried out to reveal and select SNPs with an evident “geographical signal”. Additionally, loci with an amplification lower than 80% and markers which were monomorphic in trees from Russia were removed. Finally, 95 SNP loci were used for this study. Genotyping of 70 individuals was performed on MassARRAY®iPLEX™ (Agena Bioscience™, San Diego, CA, USA) platform using iPLEX™GOLD chemistry technology. The spectra were interpreted with the software Typer Viewer v.4.0.24.71 (Agena Bioscience™). The results were processed using the programs PAST and GDA_NT (Degen et al. 2019). We calculated allelic diversity υа (Gregorius 1987), observed heterozygosity HO, expected heterozygosity HE, parameter FST of Wright’s F-statistics (Weir and Cockerham 1984), population differentiation measures δT (Gregorius 1987) and genetic distance d0 (Gregorius 1984). The software STRUCTURE version 2.3.4 (Pritchard et al. 2000) and the ΔK method (Evanno et al. 2005) were used to estimate the number of genetic clusters. Results were also analyzed using the software CLUMPAK (Kopelman et al. 2015).
Results
All pedunculate oak populations displayed relatively similar values of allelic diversity and heterozygosity (Table 2; on average, HO = 0.341 ± 0.008, HE = 0.315 ± 0.008, υа = 1.553 ± 0.015). The Tolyatti stand trended to a lower allelic diversity and expected heterozygosity.
The population differentiation parameter δT is relatively similar in stands on lowlands along the Volga (Volgograd, Kamushin and Orenburg_1) and Belaya (Ilisch) rivers (δT = 0.085–0.088), but comparatively higher in the Tolyatti and Samara_2 stands from the Privolzhye Upplands (δT = 0.107–0.127) and the most northern stand, Udm2 (δT = 0.102).
Calculation of the genetic distances between pairs of the samples (Table 3) did not reveal a relation between genetic structures of populations and their geographic location. The results of 1000 permutation tests for gene pool distance of Gregorius (1984) for all 95 SNP loci identified three genetic groups. There were no significant genetic distances between pairs when the Volgograd, Kamushin, Orenburg from the lowlands of the Volga basin were included. The genetic distance between these samples and the Ilisch stand was significant only in one case (p < 0.05). But all values of d0 were highly significant (p < 0.001) when comparing the Tolyatti and Samara_2 samples from the Privolzhye Uplands with other populations. Similar results were obtained from calculations of pairwise F.
The results of the STRUCTURE analysis showed that the best representation of the data is gained when three genetic groups are considered (Fig. 2). They are present in all seven populations (Figs. 3 and 4) but the frequency varied, especially in the samples Tolyatti and Samara_2 samples. The analysis demonstrated a clear distinction between these two populations from the Privolzhye Uplands and differences between them and populations from the lowlands of the Volga and tributaries of the river (the other five samples). Genetic differentiation among the seven populations was relatively low but statistically significant with the mean FST measure of 0.075. It decreases by almost half when Tolyatti, Samara_2 and Udm2 samples were excluded from the analysis.
Mean membership coefficients of all 70 pedunculate oak trees for the majority mode of three (K = 3) among the 10 repeated STRUCTURE runs as computed with CLUMPAK (Kopelman et al. 2015) for all seven stands
Spatial distribution of the mean membership coefficients for the majority mode of three (K = 3) among the 10 repeated STRUCTURE runs as computed with CLUMPAK (Kopelman et al. 2015) for all seven stands
Discussion
Broad-leaved forests in the study area are represented by numerous and relatively small stands, often significantly distant from each other and separated by agricultural landscapes. According to the database “Forests of Russia” of the Federal State Budgetary Institution “ROSLESINFORG”, the total area of stands dominated by the pedunculate oak is 164 km2 in the Udmurt Republic (the total area of the region is 42,100 km2), 133 km2 in the Samara oblast (53,600 km2), 1923 km2 in the Volgograd oblast (13,900 km2), 956 km2 in the Orenburg oblast (124,000 km2) and 2577 km2 in the Republic of Bashkortostan (143,600 km2). When comparing these areas and relatively vast sizes of the Russian regions studied, it is clear that the pedunculate oak stands cover very small parts of these extensive spaces. Despite this, the levels of genetic diversity (Table 2) are relatively similar. For example, the allelic diversity and heterozygosity in the sample from the Udmurt Republic are slightly higher than those from the Republic of Bashkortostan where areas of the species are larger by a hundred times. At least two main reasons may be responsible for the absence of an obvious relation of genetic diversity and pedunculate oak population sizes. The reduction in the area of populations became significant only in the last millennium due to the increasing economic development of the territory (Kozharinov and Borisov 2012; Aruova et al. 2020). A study of historical data showed that relatively abundant broad-leaved forests existed in the east section of the Russian Plain even until the end of the eighteenth century (Gorchakovsky 1968). But in subsequent centuries, these forests were seriously reduced due to a complex of factors—climate changes, agricultural development, repeating extreme cold winters accompanied by attacks of insects and phytopathogens (Tsaralunga et al. 2015). The breaks between pedunculate oak generations in Russia are 60–80 years. It is possible that the relatively small and fragmented populations in this study are not yet in a stage of serious decreasing genetic diversity due to genetic drift (Pohjanmies et al. 2016). Earlier it was noted that genetically effective long-distance pollen flow can be a factor in maintaining relatively similar genetic diversity in these populations (Buschbom et al. 2011). Heterozygosity of microsatellite loci in a geographically isolated and extremely small population (only 27 adult trees) in the Trans-Urals was even higher than for populations from the center of the distribution range. Reconstruction of possible parent pairs of alleles in the embryos revealed that at least 35% of pollination was carried out by “foreign” pollen despite the fact that other oak forests are located dozens kilometers to the West.
In general, two patterns in the genetic differentiation among populations were revealed. Despite the relatively large geographical distances between the study plots (Fig. 1) and the wide variety in climates and landscapes of the territory (Leummens 2016), results of numerical tests in 1000 permutations demonstrated that four populations from Volgograd oblast (stands Volgograd and Kamushin), Orenburg oblast (Orenburg_1 stand) and the Republic of Bashkortostan (Ilisch stand) have relatively similar gene pools. The genetic distances between them are not statistically significant. At the same time, three other populations in our study (Tolyatti, Samara_2 and Udm2_stands), located relatively closer to each other, differ genetically both from this group and among themselves on relatively high levels of statistical significance.
Recent research shows the reality of both these scenarios. The study of genetic variation at SNPs of 117 genes in 19 populations of pedunculate oak along large rivers of south-eastern Europe (Temunović et al. 2020) revealed a weak geographical structure, confirming extensive genetic connectivity or a shared ancestry. At the same time, the results of another study using 17 microsatellite loci and 10 stands of the species at the south-western range identified highly differentiated, four distinct genetic clusters (Moracho et al. 2016). It was concluded that distinct landscapes of pedunculate oaks’ refugial habitats, topography, dense vegetation, humid microclimate and high tree compatibility within a stand are likely to prevent effective pollen dispersal over long-distances.
Two contradictory patterns revealed in our study are more understandable when referring to available paleobotanical and geobotanical data, as well as to previously studies of morphological traits of the pedunculate oak. A study of pollen content of forest tree species over the past 2500 years shows that there were seven main refugia from the Privolzhye Uplands to the western boundaries of the Russian Plain (Kozharinov and Borisov 2012). pedunculate oak was present here throughout this period. The populations of this region also differ in morphological characteristics. A study (Semerikov 1986) shows that a portion (represented in this study by the Tolyatti stand), is the eastern outpost of pedunculate oak representatives with pubescent leaves, typical for the species to the west of the Caucasus, Crimea, and Moldova. To the north of this population, there are nearby pedunculate oak forests (represented by the Samara_2 sample) with hairless leaves. These two morphological forms also differ in the time of flowering by 12–15 days and area perhaps partially isolated phenologically. Depending on the climate, these populations were a source for territorial expansion (Neyshtadt 1957) to the east in the direction of the General Syrt and the Southern Urals. The flow from this territory was mixed with oncoming movement from the east from Pleistocene and Holocene refugia of the upland plateau Common Syrt and from shelters in the western Southern Urals macroslope (Gorchakovsky 1968).
Conclusions
In general, populations of pedunculate oak of the Volga basin at the eastern margin of its range possess relatively similar levels of genetic diversity, despite fragmentation of pedunculate oak stands and reduction in their size. At the same time, populations with different genetic structure are found in the study area. It is possible that the multiplicity and different ages of refugia and the complexity of migration routes from refugia and secondary glacial shelters caused this phenomenon. Studies of larger numbers of populations from other parts of the Russian Plain can be helpful to have additional confirmation of this assumption. In addition to the scientific significance of such work, it is relevant for the scientifically based management of Russian forestry.
References
Aruova LB, Ospanova ZN, Aruov BB, Alibekova NT, Shashpan Z, Kyrgizbaev AT (2020) Cyclic tests of joints of glued wooden structures. IOP Conf Ser Mat Sci Eng 829(1):012017. https://doi.org/10.1088/1757-899X/829/1/012017
Blanc-Jolivet C, Bakhtina S, Yanbaev R, Yanbaev Y, Mader M, Guichoux E, Degen B (2020) Development of new SNPs loci on Quercus robur and Quercus petraea for genetic studies covering the whole species’ distribution range. Conserv Genet Resour 1:1–4. https://doi.org/10.1007/s12686-020-01141-z
Buschbom J, Yanbaev Y, Degen B (2011) Efficient long-distance gene flow into an isolated relict oak stand. J Hered 102(4):464–472. https://doi.org/10.1093/jhered/esr023
Degen B, Yanbaev R, Yanbaev Y (2019) Genetic differentiation of Quercus robur in the South-Ural. Silvae Genet 68(1):111–115. https://doi.org/10.2478/sg-2019-0019
Ducousso A, Bordacs S (2004) Technical guidelines for genetic conservation and use: pedunculate and sessile oaks. Bioversity International, 6 p
Dumolin S, Demesure B, Petit RJ (1995) Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theor Appl Genet 91(8):1253–1256. https://doi.org/10.1007/BF00220937
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Gorchakovsky PL (1968) Plants of European broad-leaved forests on the Eastern limit of their range. Institute of Ecology of Plants and Animals, Sverdlovsk, 207 p (In Russian)
Gregorius HR (1984) A unique genetic distance. Biom J 26(1):13–18. https://doi.org/10.1002/bimj.4710260103
Gregorius HR (1987) The relationship between the concepts of genetic diversity and differentiation. Theor Appl Genet 74(3):397–401. https://doi.org/10.1007/bf00274724
Ivetić V, Devetaković J, Nonić M, Stanković D, Šijačić-Nikolić M (2016) Genetic diversity and forest reproductive material-from seed source selection to planting. iForest 9(5):801–812. https://doi.org/10.3832/ifor1577-009
König AO, Ziegenhagen B, Van Dam BC, Csaikl UM, Coart E, Degen B, Burg K, de Vries SMG, Petit RJ (2002) Chloroplast DNA variation of oaks in western Central Europe and genetic consequences of human influences. For Ecol Manag 156(1–3):147–166. https://doi.org/10.1016/S0378-1127(01)00640-5
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15(5):1179–1191. https://doi.org/10.1111/1755-0998.12387
Kozharinov AV, Borisov PV (2012) Distribution of oak forests on the territory of the Eastern Europe for the last 13 thousand years. Lesovedenie 5:22–28 (In Russian)
Lepoittevin C, Bodénès C, Chancerel E, Villate L, Lang T, Lesur I, Boury C, Ehrenmann F, Zelenica D, Boland A, Besse C, Garnier-Géré P, Plomion C, Kremer A (2015) Single-nucleotide polymorphism discovery and validation in high-density SNP array for genetic analysis in European white oaks. Mol Ecol Resour 15(6):1446–1459. https://doi.org/10.1111/1755-0998.12407
Leroy T, Rougemont Q, Dupouey JL, Bodénès C, Lalanne C, Belser C, Labadie K, Le Provost G, Aury JM, Kremer A, Plomion C (2020) Massive postglacial gene flow between European white oaks uncovered genes underlying species barriers. New Phytol 226(4):1183–1197. https://doi.org/10.1111/nph.16039
Leummens HJL (2016) Volga river basin (Russia). In: Finlayson C, Milton G, Prentice R, Davidson N (eds) The wetland book. Springer, Dordrecht, pp 1–11. https://doi.org/10.1007/978-94-007-6173-5_88-7
McKenney D, Pedlar J, O’Neill G (2009) Climate change and forest seed zones: past trends, future prospects and challenges to ponder. Forest Chron 85(2):258–266. https://doi.org/10.5558/tfc85258-2
Moracho E, Moreno G, Jordano P, Hampe A (2016) Unusually limited pollen dispersal and connectivity of pedunculate oak (Quercus robur) refugial populations at the species’ southern range margin. Mol Ecol 25(14):3319–3331. https://doi.org/10.1111/mec.13692
Neyshtadt MI (1957) History of forests and paleogeography of USSR in Holocene. Izd-vo AN SSSR, Moscow, p 402 (In Russian)
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS One 7(5):e37135. https://doi.org/10.1371/journal.pone.0037135
Petit RJ, Brewer S, Bordács S, Burg K, Cheddadi R, Coart E, Cottrell J, Csaikl UM, van Dam B, Deans JD, Espinel S, Fineschi S, Finkeldey R, Glaz I, Goicoechea PG, Jensen JS, König AO, Lowe AJ, Kremer A (2002a) Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. For Ecol Manag 156(1–3):49–74. https://doi.org/10.1016/S0378-1127(01)00634-X
Petit RJ, Csaikl UM, Bordács S, Burg K, Coart E, Cottrell J, van Dam B, Deans JD, Dumolin-Lapègue S, Fineschi S, Finkeldey R, Gillies A, Glaz I, Goicoechea PG, Jensen JS, König AO, Lowe AJ, Madsen SF, Mátyás G, Munro RC, Olalde M, Pemonge MH, Popescu F, Slade D, Tabbener H, Taurchini D, de Vries SGM, Ziegenhagen B, Kremer A (2002b) Chloroplast DNA variation in European white oaks: phylogeography and patterns of diversity based on data from over 2600 populations. For Ecol Manag 156(1–3):5–26. https://doi.org/10.1016/S0378-1127(01)00645-4
Pohjanmies T, Elshibli S, Pulkkinen P, Rusanen M, Vakkari P, Korpelainen H, Roslin T (2016) Fragmentation-related patterns of genetic differentiation in pedunculate oak (Quercus robur) at two hierarchical scales. Silva Fenn 50(2):1510. https://doi.org/10.14214/sf.1510
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Semerikov LF (1986) Population structure of woody plants (on the example of oak in the European part of the USSR and the Caucasus). Nauka, Moscow, 140 p (In Russian)
Temunović M, Garnier-Géré P, Morić M, Franjić J, Ivanković M, Bogdan S, Hampe A (2020) Candidate gene SNP variation in floodplain populations of pedunculate oak (Quercus robur L.) near the species’ southern range margin: weak differentiation yet distinct associations with water availability. Mol Ecol 29(13):2359–2378. https://doi.org/10.1111/mec.15492
Tsaralunga VV, Furmenkova ES, Kryukova AA (2015) External signs of pathology of the pedunculate oak. Voronezh State Technical University, Voronezh, 231 p (In Russian)
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370. https://doi.org/10.2307/2408641
Willing EM, Dreyer C, Van Oosterhout C (2012) Estimates of genetic differentiation measured by F ST do not necessarily require large sample sizes when using many SNP markers. PLoS One 7(8):e42649. https://doi.org/10.1371/journal.pone.0042649
Zaurbekov N, Aidosov A, Zaurbekova N, Aidosov G, Zaurbekova G, Zaurbekov I (2018) Emission spread from mass and energy exchange in the atmospheric surface layer: two-dimensional simulation. Energ Source A 40(23):2832–2841. https://doi.org/10.1080/15567036.2018.1511656
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Tao Xu
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Project funding: The work was supported by the Russian Science Foundation No. 19-16-00084.
The online version is available at http://www.springerlink.com.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Degen, B., Yanbaev, Y., Ianbaev, R. et al. Genetic diversity and differentiation among populations of the pedunculate oak (Quercus robur) at the eastern margin of its range based on a new set of 95 SNP loci. J. For. Res. 32, 2237–2243 (2021). https://doi.org/10.1007/s11676-020-01265-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11676-020-01265-w