Abstract
Objective
Vascular smooth muscle cell (VSMC) differentiation from stem cells is one source of the increasing number of VSMCs that are involved in vascular remodeling-related diseases such as hypertension, atherosclerosis, and restenosis. MicroRNA-146a (miR-146a) has been proven to be involved in cell proliferation, migration, and tumor metabolism. However, little is known about the functional role of miR-146a in VSMC differentiation from embryonic stem cells (ESCs). This study aimed to determine the role of miR-146a in VSMC differentiation from ESCs.
Methods
Mouse ESCs were differentiated into VSMCs, and the cell extracts were analyzed by Western blotting and RT-qPCR. In addition, luciferase reporter assays using ESCs transfected with miR-146a/mimic and plasmids were performed. Finally, C57BL/6J female mice were injected with mimic or miR-146a-overexpressing ESCs, and immunohistochemistry, Western blotting, and RT-qPCR assays were carried out on tissue samples from these mice.
Results
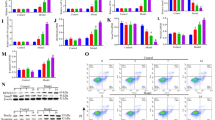
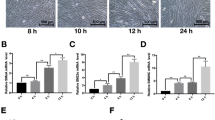
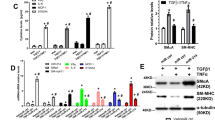
miR-146a was significantly upregulated during VSMC differentiation, accompanied with the VSMC-specific marker genes smooth muscle-alpha-actin (SMαA), smooth muscle 22 (SM22), smooth muscle myosin heavy chain (SMMHC), and h1-calponin. Furthermore, overexpression of miR-146a enhanced the differentiation process in vitro and in vivo. Concurrently, the expression of Kruppel-like factor 4 (KLF4), predicted as one of the top targets of miR-146a, was sharply decreased in miR-146a-overexpressing ESCs. Importantly, inhibiting KLF4 expression enhanced the VSMC-specific gene expression induced by miR-146a overexpression in differentiating ESCs. In addition, miR-146a upregulated the mRNA expression levels and transcriptional activity of VSMC differentiation-related transcription factors, including serum response factor (SRF) and myocyte enhancer factor 2c (MEF-2c).
Conclusion
Our data support that miR-146a promotes ESC-VSMC differentiation through regulating KLF4 and modulating the transcription factor activity of VSMCs.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Miano JM, Fisher EA, Majesky MW. Fate and State of Vascular Smooth Muscle Cells in Atherosclerosis. Circulation, 2021,143(21):2110–2116
Ho PTB, Clark IM, Le LTT. MicroRNA-Based Diagnosis and Therapy. Int J Mol Sci, 2023,13:7176
Chen T, Wu Y, Gu W, et al. Response of vascular mesenchymal stem/progenitor cells to hyperlipidemia. Cell Mol Life Sci, 2018,75(22):4079–4091
Li Z, Margariti A, Wu Y, et al. MicroRNA-199a induces differentiation of induced pluripotent stem cells into endothelial cells by targeting sirtuin 1. Mol Med Rep, 2015,12(3):3711–3717
Wang H, Ding XG, Yang JJ, et al. LncRNA MIAT facilitated BM-MSCs differentiation into endothelial cells and restored erectile dysfunction via targeting miR-200a in a rat model of erectile dysfunction. Eur J Cell Biol, 2018,97(3):180–189
Di Bernardini E, Campagnolo P, Margariti A, et al. Endothelial lineage differentiation from induced pluripotent stem cells is regulated by microRNA-21 and transforming growth factor beta2 (TGF-beta2) pathways. J Biol Chem, 2014,289(6):3383–3393
Yang D, Wang J, Xiao M, et al. Role of Mir-155 in Controlling HIF-1alpha Level and Promoting Endothelial Cell Maturation. Sci Rep, 2016,6:35316
Chen T, Margariti A, Kelaini S, et al. MicroRNA-199b Modulates Vascular Cell Fate During iPS Cell Differentiation by Targeting the Notch Ligand Jagged1 and Enhancing VEGF Signaling. Stem Cells, 2015,33(5):1405–1418
Robinson HC, Baker AH. How do microRNAs affect vascular smooth muscle cell biology?. Curr Opin Lipidol, 2012,23(5):405–411
Afzal TA, Luong LA, Chen D, et al. NCK Associated Protein 1 Modulated by miRNA-214 Determines Vascular Smooth Muscle Cell Migration, Proliferation, and Neointima Hyperplasia. J Am Heart Assoc, 2016,5(12):e004629
Chen Q, Yang F, Guo M, et al. miRNA-34a reduces neointima formation through inhibiting smooth muscle cell proliferation and migration. J Mol Cell Cardiol, 2015,89(Pt A):75–86
Yang F, Chen Q, He S, et al. miR-22 Is a Novel Mediator of Vascular Smooth Muscle Cell Phenotypic Modulation and Neointima Formation. Circulation, 2018,137(17):1824–1841
Wu Y, Li Z, Yang M, et al. MicroRNA-214 regulates smooth muscle cell differentiation from stem cells by targeting RNA-binding protein QKI. Oncotarget, 2017,8(12):19866–19878
Jin M, Wu Y, Wang Y, et al. MicroRNA-29a promotes smooth muscle cell differentiation from stem cells by targeting YY1. Stem Cell Res, 2016,17(2):277–284
Yu X, Zhang L, Wen G, et al. Upregulated sirtuin 1 by miRNA-34a is required for smooth muscle cell differentiation from pluripotent stem cells. Cell Death Differ, 2015,22(7):1170–1180
Zhao H, Wen G, Huang Y, et al. MicroRNA-22 regulates smooth muscle cell differentiation from stem cells by targeting methyl CpG-binding protein 2. Arterioscler Thromb Vasc Biol, 2015,35(4):918–929
Afshar-Khamseh R, Javeri A, Taha MF. MiR-146a suppresses the expression of CXCR4 and alters survival, proliferation and migration rate in colorectal cancer cells. Tissue Cell, 2021,73:101654
Wang W, Zhang Y, Liu W, et al. LMP1-miR-146a-CXCR4 axis regulates cell proliferation, apoptosis and metastasis. Virus Res, 2019,270:197654
Gaytan-Pacheco N, Ibanez-Salazar A, Herrera-Van Oostdam AS, et al. miR-146a, miR-221, and miR-155 are Involved in Inflammatory Immune Response in Severe COVID-19 Patients. Diagnostics (Basel), 2022,13(1):133
Hua T, Yang M, Song H, et al. Huc-MSCs-derived exosomes attenuate inflammatory pain by regulating microglia pyroptosis and autophagy via the miR-146a-5p/TRAF6 axis. J Nanobiotechnology, 2022,20(1):324
Perry MM, Moschos SA, Williams AE, et al. Rapid changes in microRNA-146a expression negatively regulate the IL-1beta-induced inflammatory response in human lung alveolar epithelial cells. J Immunol, 2008,180(8):5689–5698
Bhaumik D, Scott GK, Schokrpur S, et al. Expression of microRNA-146 suppresses NF-kappaB activity with reduction of metastatic potential in breast cancer cells. Oncogene, 2008,27(42):5643–5647
Yuan BY, Chen YH, Wu ZF, et al. MicroRNA-146a-5p Attenuates Fibrosis-related Molecules in Irradiated and TGF-beta1-Treated Human Hepatic Stellate Cells by Regulating PTPRA-SRC Signaling. Radiat Res, 2019,192(6):621–629
Zhang T, Ma Y, Gao L, et al. MicroRNA-146a protects against myocardial ischaemia reperfusion injury by targeting Med1. Cell Mol Biol Lett, 2019,24:62
Heggermont WA, Papageorgiou AP, Quaegebeur A, et al. Inhibition of MicroRNA-146a and Overexpression of Its Target Dihydrolipoyl Succinyltransferase Protect Against Pressure Overload-Induced Cardiac Hypertrophy and Dysfunction. Circulation, 2017,136(8):747–761
Zheng X, Wu Y, Zhu L, et al. Angiotensin II promotes differentiation of mouse embryonic stem cells to smooth muscle cells through PI3-kinase signaling pathway and NF-kappaB. Differentiation, 2013,85(1–2):41–54
Zhang Q, Chen T, Zhang Y, et al. MiR-30c-5p regulates adventitial progenitor cells differentiation to vascular smooth muscle cells through targeting OPG. Stem Cell Res Ther, 2021,12(1):67
Sun SG, Zheng B, Han M, et al. miR-146a and Kruppel-like factor 4 form a feedback loop to participate in vascular smooth muscle cell proliferation. EMBO Rep, 2011,12(1):56–62
Zisman D, Safieh M, Simanovich E, et al. Tocilizumab (TCZ) Decreases Angiogenesis in Rheumatoid Arthritis Through Its Regulatory Effect on miR-146a-5p and EMMPRIN/CD147. Front Immunol, 2021,12:739592
Tebbi L, Mansoori B, Safaei S, et al. MiR-146a Restoration Suppresses Triple-Negative Breast Cancer Cell Migration: A Bioinformatic and In Vitro Study. Adv Pharm Bull, 2022,12(4):842–849
Liu GJ, Zhang QR, Gao X, et al. MiR-146a Ameliorates Hemoglobin-Induced Microglial Inflammatory Response via TLR4/IRAK1/TRAF6 Associated Pathways. Front Neurosci, 2020,14:311
Keewan E, Naser SA. MiR-146a rs2910164 G > C polymorphism modulates Notch-1/IL-6 signaling during infection: a possible risk factor for Crohn’s disease. Gut Pathog, 2020,12:48
Mei J, Bachoo R, Zhang CL. MicroRNA-146a inhibits glioma development by targeting Notch1. Mol Cell Biol, 2011,31(17):3584–3592
Zhang L, Jin M, Margariti A, et al. Sp1-dependent activation of HDAC7 is required for platelet-derived growth factor-BB-induced smooth muscle cell differentiation from stem cells. J Biol Chem, 2010,285(49):38463–38472
Wang TM, Chen KC, Hsu PY, et al. microRNA let-7g suppresses PDGF-induced conversion of vascular smooth muscle cell into the synthetic phenotype. J Cell Mol Med, 2017,21(12):3592–3601
Author information
Authors and Affiliations
Corresponding authors
Additional information
Conflict of Interest Statement
The authors declare that they have no conflicts of interest.
This work was funded by the National Natural Science Foundation of China (No. 82070376 and No. 81873491), the Natural Science Foundation of Zhejiang Province (No. LY21H020005), the Zhejiang Medical Science and Technology Project (No. 2019KY376 and No. 2018KY071), and a Ningbo Science and Technology Project (No. 202002N3173).
Supplementary data
Rights and permissions
This article is licensed under a Creative Commons Attribution 4.0 International License https://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Q., Pan, Rr., Wu, Yt. et al. MicroRNA-146a Promotes Embryonic Stem Cell Differentiation towards Vascular Smooth Muscle Cells through Regulation of Kruppel-like Factor 4. CURR MED SCI 43, 223–231 (2023). https://doi.org/10.1007/s11596-023-2736-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-023-2736-3