Summary
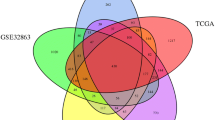
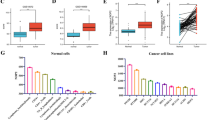
We performed a bioinformatics analysis with validation by multiple databases, aiming to evaluate the diagnostic and prognostic value of Kelch-like ECH-associated protein 1 (Keap1) mRNA for lung cancer, and to explore possible mechanisms. Diagnostic performance of Keap1 mRNA was determined by receiver operating characteristic (ROC) curve analysis. Prognostic implication of Keap1 mRNA was estimated by Kaplan-Meier survival analysis. Co-expressed genes with both Keap1 and Nfe2L2 were identified by LinkedOmics. Mechanisms of Keap1-Nfe2L2-co-expressed genes underlying the pathogenesis of lung cancer were explored by function enrichment and pathway analysis. The ROC curve analysis determined a good diagnostic performance of Keap1 mRNA for lung squamous cell carcinoma (LUSC), with an area under the ROC curve (AUC) of 0.833, sensitivity of 72.7%, and specificity of 90.6% (P<0.001). Multivariate Cox regression recognized high Keap1 mRNA to be an independent risk factor of mortality for overall lung cancer [hazard ratio (HR): 11.034, P=0.044], but an independent antagonistic factor for lung adenocarcinoma (LUAD) (HR: 0.404, P<0.001). Validation by UALCAN and GEPIA supported Oncomine findings regarding the diagnostic value of Keap1 mRNA for LUSC, but denied its prognostic value. After screening, we identified 17 co-expressed genes with both Keap1 and Nfe2L2 for LUAD, and 22 for LUSC, mainly enriched in signaling pathway of oxidative stress-induced gene expression via Nrf2. In conclusion, Keap1 mRNA has a good diagnostic performance, but controversial prognostic efficacy for LUSC. The pathogenesis of lung cancer is associated with Keap1-Nfe2L2-co-expressed genes by signaling pathway of oxidative stress-induced gene expression via Nrf2.
Similar content being viewed by others
References
Ruiz R, Hunis B, Raez LE. Immunotherapeutic agents in non-small-cell lung cancer finally coming to the front lines. Curr Oncol Rep, 2014,16(9):400
Rosell R, Skrzypski M, Jassem E, et al. BRCA1: a novel prognostic factor in resected non-small-cell lung cancer. PLoS One, 2007,2(11):e1129
Liu W, Ouyang S, Zhou Z, et al. Identification of genes associated with cancer progression and prognosis in lung adenocarcinoma: Analyses based on microarray from Oncomine and The Cancer Genome Atlas databases. Mol Genet Genomic Med, 2019,7(2):e00528
Allemani C, Weir HK, Carreira H, et al. Global surveillance of cancer survival 1995–2009: analysis of individual data for 25,676,887 patients from 279 population-based registries in 67 countries (CONCORD-2). Lancet, 2015,385(9972):977–1010
Hou GX, Liu P, Yang J, et al. Mining expression and prognosis of topoisomerase isoforms in non-small-cell lung cancer by using Oncomine and Kaplan-Meier plotter. PLoS One, 2017,12(3):e0174515
Wu B, Yang S, Sun H, et al. Keap1 Inhibits Metastatic Properties of NSCLC Cells by Stabilizing Architectures of F-Actin and Focal Adhesions. Mol Cancer Res, 2018,16(3):508–516
Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature, 2012,489(7417):519–525
Gomez DR, Byers LA, Nilsson M, et al. Integrative proteomic and transcriptomic analysis provides evidence for TrkB (NTRK2) as a therapeutic target in combination with tyrosine kinase inhibitors for non-small cell lung cancer. Oncotarget, 2018,9(18):14 268–14 284
Cloer EW, Goldfarb D, Schrank TP, et al. NRF2 Activation in Cancer: From DNA to Protein. Cancer Res, 2019,79(5):889–898
Leinonen HM, Kansanen E, Polonen P, et al. Dysregulation of the Keap1-Nrf2 pathway in cancer. Biochem Soc Trans, 2015,43(4):645–649
Huddleston HG, Wong KK, Welch WR, et al. Clinical applications of microarray technology: creatine kinase B is an up-regulated gene in epithelial ovarian cancer and shows promise as a serum marker. Gynecol Oncol, 2005,96(1):77–83
Hormigo A, Gu B, Karimi S, et al. YKL-40 and matrix metalloproteinase-9 as potential serum biomarkers for patients with high-grade gliomas. Clin Cancer Res, 2006,12(19):5698–5704
Jiang X, Barmada MM, Visweswaran S. Identifying genetic interactions in genome-wide data using Bayesian networks. Genet Epidemiol, 2010,34(6):575–581
Xie ZC, Dang YW, Wei DM, et al. Clinical significance and prospective molecular mechanism of MALAT1 in pancreatic cancer exploration: a comprehensive study based on the GeneChip, GEO, Oncomine, and TCGA databases. Onco Targets Ther, 2017,10:3991–4005
Meng N, Wang J, Sun J, et al. Using amide proton transfer to identify cervical squamous carcinoma/adenocarcinoma and evaluate its differentiation grade. Magn Reson Imaging, 2019,61:9–15
Jiang Y, Zheng X, Jiao D, et al. Peptidase inhibitor 15 as a novel blood diagnostic marker for cholangiocarcinoma. EBioMedicine, 2019,40:422–431
Heilbroner SP, Xanthopoulos EP, Buono D, et al. Impact of estrogen monotherapy on survival in women with stage III-IV non-small cell lung cancer. Lung Cancer, 2019,129: 8–15
Huang X, Zhao J, Bai J, et al. Risk and clinicopathological features of osteosarcoma metastasis to the lung: A population-based study. J Bone Oncol, 2019,16:100230
Khorrami M, Jain P, Bera K, et al. Predicting pathologic response to neoadjuvant chemoradiation in resectable stage III non-small cell lung cancer patients using computed tomography radiomic features. Lung Cancer, 2019,135:1–9
Yamamoto M, Kurokawa Y, Kobayashi N, et al. Prognostic Value of the Combined Index of Plasma Fibrinogen and the Neutrophil-Lymphocyte Ratio in Gastric Cancer. World J Surg, 2020,44(1):207–212
Wang S, Xiong Y, Zhao Q, et al. Columnar cell papillary thyroid carcinoma prognosis: findings from the SEER database using propensity score matching analysis. Am J Transl Res, 2019,11(9):6262–6270
Xu M, Jin T, Chen L, et al. Mortalin is a distinct bio-marker and prognostic factor in serous ovarian carcinoma. Gene, 2019,696:63–71
Li F, Guo P, Dong K, et al. Identification of Key Biomarkers and Potential Molecular Mechanisms in Renal Cell Carcinoma by Bioinformatics Analysis. J Comput Biol, 2019,26(11):1278–1295
Bhattacharjee A, Richards WG, Staunton J, et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci USA, 2001,98(24): 13 790–13 795
Garber ME, Troyanskaya OG, Schluens K, et al. Diversity of gene expression in adenocarcinoma of the lung. Proc Natl Acad Sci USA, 2001,98(24):13784–13789
Beer DG, Kardia SL, Huang CC, et al. Gene-expression profiles predict survival of patients with lung adenocarcmoma. Nat Med, 2002,8(8):816–824
Wachi S, Yoneda K, Wu R. Interactome-transcriptome analysis reveals the high centrality of genes differentially expressed in lung cancer tissues. Bioinformatics, 2005,21(23):4205–4208
Talbot SG, Estilo C, Maghami E, et al. Gene expression profiling allows distinction between primary and metastatic squamous cell carcinomas in the lung. Cancer Res, 2005,65(8):3063–3071
Selamat SA, Chung BS, Girard L, et al. Genome-scale analysis of DNA methylation in lung adenocarcinoma and integration with mRNA expression. Genome Res, 2012,22(7):1197–1211
Stearman RS, Dwyer-Nield L, Zerbe L, et al. Analysis of orthologous gene expression between human pulmonary adenocarcinoma and a carcinogen-induced murine model. Am J Pathol, 2005,167(6):1763–1775
Hou J, Aerts J, den Hamer B, et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS One, 2010,5(4):e10312
Su LJ, Chang CW, Wu YC, et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics, 2007,8:140
Okayama H, Kohno T, Ishii Y, et al. Identification of genes upregulated in ALK-positive and EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res, 2012,72(1):100–111
Landi MT, Dracheva T, Rotunno M, et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS One, 2008,3(2):e1651
Chen C, Guo Q, Song Y, et al. SKA1/2/3 serves as a biomarker for poor prognosis in human lung adenocarcinoma. Transl Lung Cancer Res, 2020,9(2): 218–231
Zhao Y, Xue C, Xie Z, et al. Comprehensive analysis of ubiquitin-specific protease 1 reveals its importance in hepatocellular carcinoma. Cell Prolif, 2020,53(10):e12908
Wong KF, Xu Z, Chen J, et al. Circulating markers for prognosis of hepatocellular carcinoma. Expert Opin Med Diagn, 2013,7(4):319–329
Frank R, Scheffler M, Merkelbach-Bruse S, et al. Clinical and Pathological Characteristics of KEAP1- and NFE2L2-Mutated Non-Small Cell Lung Carcinoma (NSCLC). Clin Cancer Res, 2018,24(13):3087–3096
Zhao J, Lin X, Meng D, et al. Nrf2 Mediates Metabolic Reprogramming in Non-Small Cell Lung Cancer. Front Oncol, 2020,10:578315
Liu Q, Gao Y, Ci X. Role of Nrf2 and Its Activators in Respiratory Diseases. Oxid Med Cell Longev, 2019, 2019:7090534
Mizumura K, Maruoka S, Shimizu T, et al. Role of Nrf2 in the pathogenesis of respiratory diseases. Respir Investig, 2020,58(1):28–35
Pan Y, Li W, Feng Y, et al. Edaravone attenuates experimental asthma in mice through induction of HO-1 and the Keap1/Nrf2 pathway. Exp Ther Med, 2020,19(2):1407–1416
Chapple SJ, Cheng X, Mann GE. Effects of 4-hydroxynonenal on vascular endothelial and smooth muscle cell redox signaling and function in health and disease. Redox Biol, 2013,1(1):319–331
Morimitsu Y, Nakagawa Y, Hayashi K, et al. A sulforaphane analogue that potently activates the Nrf2-dependent detoxification pathway. J Biol Chem, 2002,277(5):3456–3463
Conaway CC, Wang CX, Pittman B, et al. Phenethyl isothiocyanate and sulforaphane and their N-acetylcysteine conjugates inhibit malignant progression of lung adenomas induced by tobacco carcinogens in A/J mice. Cancer Res, 2005,65(18):8548–8557
Hellyer JA, Padda SK, Diehn M, et al. Clinical implications of KEAP1-NFE2L2 mutations in non-small cell lung cancer. J Thorac Oncol, 2021,16(3):395–403
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Liu, Gy., Zhang, W., Chen, Xc. et al. Diagnostic and Prognostic Significance of Keap1 mRNA Expression for Lung Cancer Based on Microarray and Clinical Information from Oncomine Database. CURR MED SCI 41, 597–609 (2021). https://doi.org/10.1007/s11596-021-2378-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-021-2378-2