Abstract
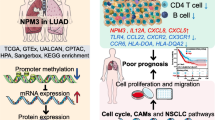
Recent studies have shown that NOP2, a nucleolar protein, is up-regulated in various cancers, suggesting a potential link to tumor aggressiveness and unfavorable outcomes. This study examines NOP2's role in lung adenocarcinoma (LUAD), a context where its implications remain unclear. Utilizing bioinformatics, we assessed 513 LUAD and 59 normal tissue samples from The Cancer Genome Atlas (TCGA) to explore NOP2's diagnostic and prognostic significance in LUAD. Additionally, in vitro experiments compared NOP2 expression between Beas-2b and A549 cells. Advanced databases and analytical tools, including LINKEDOMICS, STRING, and TISIDB, were employed to further elucidate NOP2's association with LUAD. Our findings indicate a significantly higher expression of NOP2 mRNA and protein in A549 cells compared to Beas-2b cells (P < 0.001). In LUAD, elevated NOP2 levels were linked to decreased Overall Survival (OS) and advanced clinical stages. Univariate Cox analysis revealed that high NOP2 expression correlated with poorer OS in LUAD (P < 0.01), a finding independently supported by multivariate Cox analysis (P < 0.05). The relationship between NOP2 expression and LUAD risk was presented via a Nomogram. Additionally, Gene Set Enrichment Analysis (GSEA) identified seven NOP2-related signaling pathways. A focal point of our research was the interplay between NOP2 and tumor-immune interactions. Notably, a negative correlation was observed between NOP2 expression and the immune infiltration levels of macrophages, neutrophils, mast cells, Natural Killer (NK) cells, and CD8 + T cells in LUAD. Moreover, the expression of NOP2 was related to the sensitivity of various chemotherapeutic drugs. In vitro, we found that downregulating NOP2 can decrease the proliferation, migration and invasion of A549 cells. Furthermore, NOP2 can regulate Caspase3-mediated apoptosis. Collectively, particularly regarding prognosis, immune infiltration and vitro experiments, these findings suggest NOP2's potential of serving as a poor-prognostic biomarker for LUAD and aggravating the malignancy of lung adenocarcinoma cells.










Similar content being viewed by others
Data Availability
The RNA-sequencing data and corresponding clinical information were downloaded from the Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/).
NOP2 expression data in various cancers come from TIMER database (http://timer.comp-genomics.org/).
Immunohistochemical picture of lung adenocarcinoma from HPA database (https://www.proteinatlas.org/ENSG00000111641-NOP2/tissue/lung).
The difference data of phosphorylation sites in normal tissues and primary cancer tissues comes from the UALCAN database (http://ualcan.path.uab.edu/cgi-bin/CPTAC-Result.pl?genenam=nop2&ctype=LUAD).
Data related to gene mutation is from cBioPortal database (https://www.cbioportal.org/results/cancerTypesSummary?cancer_study_list=pancan_pcawg_2020&Z_SCORE_THRESHOLD=2.0&RPPA_SCORE_THRESHOLD=2.0&profileFilter=mutations%2Ccna&case_set_id=pancan_pcawg_2020_cnaseq&gene_list=NOP2&geneset_list=%20&tab_index=tab_visualize&Action=Submit).
Protein interaction network diagram is from STRING (https://cn.string-db.org/cgi/network?taskId=bQXGyAOwacAE&sessionId=bfFT6WBHvU0K) and GeneMANIA database (http://genemania.org/search/homo-sapiens/nop2).
Tumor immune correlation analysis data is from TISDIB database (http://cis.hku.hk/TISIDB/browse.php?gene=NOP2).
Download cancer and normal cell line data from the bioGPS database (http://biogps.org/#goto=welcome).
NOP2 related genes were downloaded from LinkedOmics database (http://www.linkedomics.org/admin.php).
GO and KEGG enrichment analysis data were from the metascape database (https://metascape.org/gp/index.html#/main/step1).
Drug sensitivity analysis data were from CellMiner database (https://discover.nci.nih.gov/cellminer/home.do) and GSCA online analysis platform (http://bioinfo.life.hust.edu.cn/GSCA/#/).
References
AbdulJabbar K, Raza SEA, Rosenthal R, Jamal-Hanjani M, Veeriah S, Akarca A, Lund T, Moore DA, Salgado R, Al Bakir M, Zapata L, Hiley CT, Officer L, Sereno M, Smith CR, Loi S, Hackshaw A, Marafioti T, Quezada SA, McGranahan N, Le Quesne J, Swanton C, Yuan Y (2020) Geospatial immune variability illuminates differential evolution of lung adenocarcinoma. Nat Med 26:1054–1062. https://doi.org/10.1038/s41591-020-0900-x
Bi J, Huang Y, Liu Y (2019) Effect of NOP2 knockdown on colon cancer cell proliferation, migration, and invasion. Transl Cancer Res 8:2274–2283. https://doi.org/10.21037/tcr.2019.09.46
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Carbone DP, Gandara DR, Antonia SJ, Zielinski C, Paz-Ares L (2015) Non-small-cell lung cancer: role of the immune system and potential for immunotherapy. J Thorac Oncol 10(7):974–984. https://doi.org/10.1097/JTO.0000000000000551
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, Sander C, Schultz N (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–404. https://doi.org/10.1158/2159-8290.Cd-12-0095
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, Varambally S (2017) UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 19:649–658. https://doi.org/10.1016/j.neo.2017.05.002
Chellamuthu A, Gray SG (2020) The RNA methyltransferase NSUN2 and its potential roles in cancer. Cells 9(8):1758. https://doi.org/10.3390/cells9081758
Chen F, Chandrashekar DS, Varambally S, Creighton CJ (2019) Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers. Nat Commun 10:5679. https://doi.org/10.1038/s41467-019-13528-0
Cheng JX, Chen L, Li Y, Cloe A, Yue M, Wei J, Watanabe KA, Shammo JM, Anastasi J, Shen QJ, Larson RA, He C, Le Beau MM, Vardiman JW (2018) RNA cytosine methylation and methyltransferases mediate chromatin organization and 5-azacytidine response and resistance in leukaemia. Nat Commun 9:1163. https://doi.org/10.1038/s41467-018-03513-4
Cho JK, Lee GJ, Yi KI, Cho KS, Choi N, Kim JS, Kim H, Oh D, Choi SK, Jung SH, Jeong HS, Ahn YC (2015) Development and external validation of nomograms predictive of response to radiation therapy and overall survival in nasopharyngeal cancer patients. Eur J Cancer 51:1303–1311. https://doi.org/10.1016/j.ejca.2015.04.003
DarılmazYüce G, OrtaçErsoy E (2016) Lung cancer and epigenetic modifications. Tuberk Toraks 64:163–170. https://doi.org/10.5578/tt.10231
Fang Z, Li P, Li F (2023) Combining Bulk RNA-seq and scRNA-seq data to identify RNA m5C methyltransferases NSUN1: a rising star as a biomarker for cancer diagnosis, prognosis and therapy. Transl Cancer Res 12:2336–2350. https://doi.org/10.21037/tcr-23-66
Feng J, Zhang J, Li Y, Wang J, Mo P, Lin L (2022) Upregulated expression of NOP2 predicts worse prognosis of gastric adenocarcinoma by promoting tumor growth. Saudi J Gastroenterol 28:369–377. https://doi.org/10.4103/sjg.sjg_573_21
Fois SS, Paliogiannis P, Zinellu A, Fois AG, Cossu A, Palmieri G (2021) Molecular epidemiology of the main druggable genetic alterations in non-small cell lung cancer. Int J Mol Sci 22(2):612. https://doi.org/10.3390/ijms22020612
Fonagy A, Swiderski C, Dunn M, Freeman JW (1992) Antisense-mediated specific inhibition of P120 protein expression prevents G1- to S-phase transition. Cancer Res 52:5250–5256
Fonagy A, Swiderski C, Ostrovsky AM, Bolton WE, Freeman JW (1994) Effect of nucleolar P120 expression level on the proliferation capacity of breast cancer cells. Cancer Res 54:1859–1864
Freeman JW, Busch RK, Gyorkey F, Gyorkey P, Ross BE, Busch H (1988) Identification and characterization of a human proliferation-associated nucleolar antigen with a molecular weight of 120,000 expressed in early G1 phase. Cancer Res 48:1244–1251
Gao Y, Wang Z, Zhu Y, Zhu Q, Yang Y, Jin Y, Zhang F, Jiang L, Ye Y, Li H, Zhang Y, Liang H, Xiang S, Miao H, Liu Y, Hao Y (2019) NOP2/Sun RNA methyltransferase 2 promotes tumor progression via its interacting partner RPL6 in gallbladder carcinoma. Cancer Sci 110:3510–3519. https://doi.org/10.1111/cas.14190
Gong Y, Liu Y, Wang T, Li Z, Gao L, Chen H, Shu Y, Li Y, Xu H, Zhou Z, Dai L (2021) Age-associated proteomic signatures and potential clinically actionable targets of colorectal cancer. Mol Cell Proteomics 20:100115. https://doi.org/10.1016/j.mcpro.2021.100115
Guo Y, Song J, Wang Y, Huang L, Sun L, Zhao J, Zhang S, Jing W, Ma J, Han C (2020) concurrent genetic alterations and other biomarkers predict treatment efficacy of EGFR-TKIs in EGFR-mutant non-small cell lung cancer: a review. Front Oncol 10:610923. https://doi.org/10.3389/fonc.2020.610923
Han J, Deng X, Sun R, Luo M, Liang M, Gu B, Zhang T, Peng Z, Lu Y, Tian C, Yan Y, Luo Z (2021) GPI is a prognostic biomarker and correlates with immune infiltrates in lung adenocarcinoma. Front Oncol 11:752642. https://doi.org/10.3389/fonc.2021.752642
Hirsch FR, Scagliotti GV, Mulshine JL, Kwon R, Curran WJ Jr, Wu YL, Paz-Ares L (2017) Lung cancer: current therapies and new targeted treatments. Lancet 389:299–311. https://doi.org/10.1016/s0140-6736(16)30958-8
Hong J, Lee JH, Chung IK (2016) Telomerase activates transcription of cyclin D1 gene through an interaction with NOL1. J Cell Sci 129:1566–1579. https://doi.org/10.1242/jcs.181040
Jin CY, Du L, Nuerlan AH, Wang XL, Yang YW, Guo R (2020) High expression of RRM2 as an independent predictive factor of poor prognosis in patients with lung adenocarcinoma. Aging (Albany NY) 13:3518–3535. https://doi.org/10.18632/aging.202292
Job B, Bernheim A, Beau-Faller M, Camilleri-Broët S, Girard P, Hofman P, Mazières J, Toujani S, Lacroix L, Laffaire J, Dessen P, Fouret P (2010) Genomic aberrations in lung adenocarcinoma in never smokers. PLoS One 5:e15145. https://doi.org/10.1371/journal.pone.0015145
Kang HW, Kim SM, Kim WT, Yun SJ, Lee SC, Kim WJ, Hwang EC, Kang SH, Hong SH, Chung J, Kwon TG, Kim HH, Kwak C, Byun SS, Kim YJ (2020) The age-adjusted Charlson comorbidity index as a predictor of overall survival of surgically treated non-metastatic clear cell renal cell carcinoma. J Cancer Res Clin Oncol 146:187–196. https://doi.org/10.1007/s00432-019-03042-7
Kong W, Biswas A, Zhou D, Fiches G, Fujinaga K, Santoso N, Zhu J (2020) Nucleolar protein NOP2/NSUN1 suppresses HIV-1 transcription and promotes viral latency by competing with Tat for TAR binding and methylation. PLoS Pathog 16:e1008430. https://doi.org/10.1371/journal.ppat.1008430
Lewinska A, Bednarz D, Adamczyk-Grochala J, Wnuk M (2017) Phytochemical-induced nucleolar stress results in the inhibition of breast cancer cell proliferation. Redox Biol 12:469–482. https://doi.org/10.1016/j.redox.2017.03.014
Li H, Guo L, Cai Z (2022a) TCN1 is a potential prognostic biomarker and correlates with immune infiltrates in lung adenocarcinoma. World J Surg Oncol 20:83. https://doi.org/10.1186/s12957-022-02556-8
Li M, Tao Z, Zhao Y, Li L, Zheng J, Li Z, Chen X (2022b) 5-methylcytosine RNA methyltransferases and their potential roles in cancer. J Transl Med 20:214. https://doi.org/10.1186/s12967-022-03427-2
Lin JJ, Riely GJ, Shaw AT (2017) Targeting ALK: Precision Medicine Takes on Drug Resistance. Cancer Discov 7:137–155. https://doi.org/10.1158/2159-8290.Cd-16-1123
Liu CJ, Hu FF, Xia MX, Han L, Zhang Q, Guo AY (2018) GSCALite: a web server for gene set cancer analysis. Bioinformatics 34:3771–3772. https://doi.org/10.1093/bioinformatics/bty411
Moloney JN, Cotter TG (2018) ROS signalling in the biology of cancer. Semin Cell Dev Biol 80:50–64. https://doi.org/10.1016/j.semcdb.2017.05.023
Motolani A, Martin M, Sun M, Lu T (2020) Phosphorylation of the regulators, a complex facet of NF-κB signaling in cancer. Biomolecules 11(1):15. https://doi.org/10.3390/biom11010015
Mountain CF (1997) Revisions in the International System for Staging Lung Cancer. Chest 111:1710–1717. https://doi.org/10.1378/chest.111.6.1710
Noor ZS, Cummings AL, Johnson MM, Spiegel ML, Goldman JW (2020) Targeted therapy for non-small cell lung cancer. Semin Respir Crit Care Med 41:409–434. https://doi.org/10.1055/s-0039-1700994
Pastushenko I, Blanpain C (2019) EMT transition states during tumor progression and metastasis. Trends Cell Biol 29:212–226. https://doi.org/10.1016/j.tcb.2018.12.001
Perillo B, Di Donato M, Pezone A, Di Zazzo E, Giovannelli P, Galasso G, Castoria G, Migliaccio A (2020) ROS in cancer therapy: the bright side of the moon. Exp Mol Med 52:192–203. https://doi.org/10.1038/s12276-020-0384-2
Perlaky L, Valdez BC, Busch RK, Larson RG, Jhiang SM, Zhang WW, Brattain M, Busch H (1992) Increased growth of NIH/3T3 cells by transfection with human p120 complementary DNA and inhibition by a p120 antisense construct. Cancer Res 52:428–436
Reinhold WC, Sunshine M, Liu H, Varma S, Kohn KW, Morris J, Doroshow J, Pommier Y (2012) Cell Miner: a web-based suite of genomic and pharmacologic tools to explore transcript and drug patterns in the NCI-60 cell line set. Cancer Res 72:3499–3511. https://doi.org/10.1158/0008-5472.Can-12-1370
Romaszko AM, Doboszyńska A (2018) Multiple primary lung cancer: A literature review. Adv Clin Exp Med 27:725–730. https://doi.org/10.17219/acem/68631
Sears CR, Mazzone PJ (2020) Biomarkers in lung cancer. Clin Chest Med 41:115–127. https://doi.org/10.1016/j.ccm.2019.10.004
Shah DR, Masters GA (2020) Precision medicine in lung cancer treatment. Surg Oncol Clin N Am 29:15–21. https://doi.org/10.1016/j.soc.2019.08.002
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Sun F, Wu K, Yao Z, Mu X, Zheng Z, Sun M, Wang Y, Liu Z, Zhu Y (2020) Long noncoding RNA LINC00963 induces NOP2 expression by sponging tumor suppressor miR-542–3p to promote metastasis in prostate cancer. Aging (Albany NY) 12:11500–11516. https://doi.org/10.18632/aging.103236
Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork P, Jensen LJ, von Mering C (2011) The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39:D561-568. https://doi.org/10.1093/nar/gkq973
Tsimberidou AM, Fountzilas E, Nikanjam M, Kurzrock R (2020) Review of precision cancer medicine: Evolution of the treatment paradigm. Cancer Treat Rev 86:102019. https://doi.org/10.1016/j.ctrv.2020.102019
Wang F, Yuan JH, Wang SB, Yang F, Yuan SX, Ye C, Yang N, Zhou WP, Li WL, Li W, Sun SH (2014) Oncofetal long noncoding RNA PVT1 promotes proliferation and stem cell-like property of hepatocellular carcinoma cells by stabilizing NOP2. Hepatology 60:1278–1290. https://doi.org/10.1002/hep.27239
Wang N, Tang H, Wang X, Wang W, Feng J (2017) Homocysteine upregulates interleukin-17A expression via NSun2-mediated RNA methylation in T lymphocytes. Biochem Biophys Res Commun 493:94–99. https://doi.org/10.1016/j.bbrc.2017.09.069
Wang H, Wang L, Wang Z, Dang Y, Shi Y, Zhao P, Zhang K (2020) The nucleolar protein NOP2 is required for nucleolar maturation and ribosome biogenesis during preimplantation development in mammals. Faseb J 34:2715–2729. https://doi.org/10.1096/fj.201902623R
Xing J, Yi J, Cai X, Tang H, Liu Z, Zhang X, Martindale JL, Yang X, Jiang B, Gorospe M, Wang W (2015) NSun2 promotes cell growth via elevating cyclin-dependent kinase 1 translation. Mol Cell Biol 35:4043–4052. https://doi.org/10.1128/mcb.00742-15
Yang S, Zhou D, Zhang C, Xiang J, Xi X (2023) Function of m(5)C RNA methyltransferase NOP2 in high-grade serous ovarian cancer. Cancer Biol Ther 24:2263921. https://doi.org/10.1080/15384047.2023.2263921
Zou J, Wang E (2019) Cancer biomarker discovery for precision medicine: new progress. Curr Med Chem 26:7655–7671. https://doi.org/10.2174/0929867325666180718164712
Acknowledgements
Our gratitude is extended to the TCGA, bioGPS, HPA, UALCAN, LINKEDOMICS, Metascape, TIMER, STRING, GeneMANIA, cBioPortal, and TISIDB databases for their availability.
Funding
This study was financially supported by China Postdoctoral Science Foundation (2022M721682).
Author information
Authors and Affiliations
Contributions
WZQ, QZ, and GQF contributed to data curation, analysis, investigation, methodology, and manuscript writing; PMW, ZJL, and WL provided guidance and reviewed the thesis. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for Publication
Informed consent for the publication of identifiable information or images in an open-access journal was obtained from all study participants.
Institutional Review Board Statement
Not applicable.
Informed Consent
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qin, W., Fei, G., Zhou, Q. et al. Nuclear protein NOP2 serves as a poor-prognosis predictor of LUAD and aggravates the malignancy of lung adenocarcinoma cells. Funct Integr Genomics 24, 58 (2024). https://doi.org/10.1007/s10142-024-01337-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-024-01337-8