Abstract
The identity and diversity of Kordyana species on three native species of Commelinaceae in Australia were studied following surveys in 2020–2022 for Kordyana brasiliensis, which had been deliberately released as a biocontrol agent for the environmental weed Tradescantia fluminensis. Three new species of Kordyana are described from Australia based on DNA sequence analysis of the ITS and LSU rDNA regions, morphology, host associations, and geographic distributions. Two new species, Kordyana spectabilis on Aneilema acuminatum and Kordyana luteoalba on Pollia crispata, occur in shaded rainforest habitats in eastern Australia. The third new species, Kordyana occidentalis on Commelina ensifolia, occurs in forests and woodlands of the Kimberley region of Western Australia. Morphological descriptions are provided for these three new species of Kordyana as well as for the conidial stage of K. brasiliensis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Kordyana (Brachybasidaceae, Exobasidiomycetes) accommodates dimorphic biotrophic leaf pathogens that cause leaf spots with non-pigmented spores on the abaxial side, mostly on host plants in the Commelinaceae (Piepenbring et al. 2020). Kordyana species have been reported from Africa, Asia, Australia, Russia, Central America (Panama, Costa Rica), and South America (Raciborski 1900; Gäumann 1922; Sawada 1929; Petrak 1950; Gruèzo 1990; Gómez and Kisimova-Horovitz 1997; Barreto and Evans 1988; Piepenbring et al. 2010; Macedo et al. 2016; Park et al. 2021; Dudka 2023). Kordyana is typified by the type species, K. tradescantiae (Pat.) Racib. (Raciborski 1900), and accommodates seven additional valid species, K. aneilemae Sawada; K. boswelliae Thirum., Patel, G.W. Dhande & V.V. Bhatt; K. brasiliensis D.M. Macedo, O.L. Pereira & R.W. Barreto; K. celebensis Gäum.; K. commelinae Petch; K. cyphelloidis Viégas; and K. polliae Gäum. (Piepenbring et al. 2020). There are two additional species which lack a diagnosis, K. indica and K. polliae var. microspora (is considered a synonym of K. polliae) (Piepenbring et al. 2020; see Online Resource 1).
The classification of Kordyana species has traditionally been based on morphology and host range (Raciborski 1900; Gäumann 1922; Sawada 1929; Petrak 1950; Gruèzo 1990). However, the morphological approach to fungal classification is often unreliable due to the variability, or lack thereof, of phenotypic traits (including morphology and host range). In a critical revision of the Brachybasidicaeae, Piepenbring et al. (2020) used phylogenetic analysis of the ITS and LSU rDNA sequences to show that K. brasiliensis, K. celebensis, K. tradescantiae, and Dicellomyces gloeosporus formed a well-supported clade. This phylogenetic analysis was unable to resolve the relations between these taxa and contrasts the findings of Park et al. (2021), who placed K. commelinae sister to Marantokordyana species and not D. gloeosporus based on ITS rDNA. Unfortunately, the taxonomic resolution of Kordyana has been hampered as molecular barcodes are available for only four of the eight known species (Begerow et al. 2002; Macedo et al. 2016; Park et al. 2021; Dudka 2023).
Kordyana brasiliensis was deliberately released in New Zealand in 2018 and in Victoria, Australia, in 2019 as a biocontrol agent for the environmental weed Tradescantia fluminensis (Landcare Research 2020; Winston et al. 2021; Morin et al. 2022). Kordyana brasiliensis was discovered and described during exploratory surveys for potential biocontrol agents for T. fluminensis in Brazil (Macedo et al. 2016) and is reported as host-specific to T. fluminensis. Preliminary monitoring in Australia and New Zealand has shown that K. brasiliensis has become well-established in both countries since its release, with high levels of disease recorded at some locations (Morin et al. 2022; Landcare Research 2020). Kordyana brasiliensis is the only Kordyana species recorded in Australia according to the Fungi Name Index (FNI) (Australian National Species List 2024). Herbarium records show collections of unidentified specimens of Kordyana collected on Commelina sp., C. ensifolia, and Aneilema acuminatum in Australia. Seven genera within Commelinaceae occur naturally in Australia including Aneilema, Cartonema, Commelina, Cyanotis, Floscopa, Murdannia, and Pollia (PlantNET 2022; Western Australian Herbarium 2024; Queensland Government 2024). The diversity of Kordyana species on native Commelinaceae in Australia has not been previously studied.
From 2020 to 2022, surveys were made before and after the release of K. brasiliensis in New South Wales (NSW), Australia, to determine the success of the biological control release program. During these surveys, specimens of native Australian Commelinaceae, co-occurring with T. fluminensis, with white leaf lesions were found and collected. The identities of these specimens were studied using multigene phylogenetic analyses to determine whether K. brasiliensis had extended its known host range to include native Australian Commelinaceae, or whether they represented other Kordyana species. Among the specimens collected and including herbaria records, three new species of Kordyana were found, each restricted to a single host species, i.e., Aneilema acuminatum, Commelina ensifolia, and Pollia crispata.
Materials and methods
Specimens examined
Leaves of Aneilema acuminatum, Commelina spp., and Pollia crispata with white leaf lesions were collected before and after the release of K. brasiliensis in NSW in eastern Australia between 2020 and 2022 (CSIRO 2023). Additional surveys of native Australian Commelinaceae, including A. acuminatum, Commelina spp., and P. crispata, with white leaf lesions, were conducted in Queensland (QLD) in 2021. Only native Commelinaceae species co-occurring with T. fluminensis at the study sites were examined. Plant hosts were identified by morphological descriptions and known geographical distributions in Flora of NSW (PlantNET 2022). Young leaf lesions were excised and attached with petroleum jelly to the inner lids of Petri dishes and inverted over potato dextrose agar (PDA) plates. The PDA plates were incubated in the dark at 21–22 °C for 1–2 days to capture discharged basidiospores on the agar surface. Single germinated basidiospores were transferred to fresh plates with a sterile needle and incubated in the dark at 21–22 °C for several weeks.
Selected specimens (dried leaves and fungal cultures) were deposited in the Queensland Plant Pathology Herbarium (BRIP) (Table 1). Fungal cultures were permanently preserved in a metabolically inactive state at − 80 °C on agar pieces in 15% glycerol (v/v) in the Queensland Plant Pathology Herbarium (BRIP). Furthermore, three herbarium specimens of possible Kordyana on Commelinaceae collected in Australia (BRIP 53424, BRIP 65743, and PERTH 3203697) were borrowed and examined (Table 1). The identity of the fungus causing the white lesions in these previous collections had not been published.
DNA extraction, PCR amplification, DNA sequencing, and phylogenetic analysis
Genomic DNA was isolated directly from leaf lesions of herbarium specimens, as well as from pure fungal cultures grown on PDA. Samples were homogenised in a FastPrep®-24 Classic Tissue and Cell Homogeniser (MP Biomedicals LLC, USA), and genomic DNA was extracted with the ISOLATE II Plant DNA Kit (Meridian Bioscience, Australia) and the manufacturer’s protocol. Genomic DNA was diluted 1:10 or 1:100 for PCR amplification by MyFi Mix (Meridian Bioscience, Australia) and manufacturer’s instructions with a final reaction volume of 25 µl. The primers used were ITS1-F (Gardes and Bruns 1993) and ITS4 (White et al. 1990) for ITS; LROR (Moncalvo et al. 1995) and LR5/LR6 (Vilgalys and Hester 1990) for LSU; and NS1 and NS4 (White et al. 1990) for SSU. PCR products were amplified with the following parameters: initial denaturation at 95 °C for 1 min, followed by 35 cycles of denaturation at 95 °C for 15 s, annealing at 53 °C for ITS, 62 °C for LSU and 52 °C for SSU for 15 s, and an extension at 72 °C for 30 s followed by a final extension at 72 °C for 7 min. PCR amplicons were cleaned with ISOLATE II PCR and Gel Kit (Meridian Bioscience, Australia) and then submitted to either Macrogen Inc (Seoul, Korea) or the Biomolecular Resource Facility (BRF) (Canberra, Australia) for DNA sequencing.
DNA sequence chromatograms were assembled and manually checked in Geneious Prime 2023.01.1 (Geneious Prime 2023.01.1, https://www.geneious.com). Basic Local Alignment Tool (BLAST) searches in GenBank using consensus sequences were performed to compare their identity to sequences of ex-type specimens in the Exobasidiales, and an alignment dataset for phylogenetic analysis was constructed. DNA sequences obtained in this study were deposited in GenBank (Table 1).
Sequence alignments were constructed in MEGA11 v. 11.0.13 (Tamura et al. 2021) with Muscle (Edgar 2004). DNA sequence alignments were edited manually to ensure correct alignment before the removal of poorly aligned regions with GBlocks v. 0.91.1 (Castresana 2000) through the NGPhylogeny.fr website (Lemoine et al. 2019). The parameters used for the ITS, LSU, and SSU were 18/29/11/5/with half, 22/35/8/5/with half, and 13/20/8/5/with half, respectively. The alignments were concatenated to provide a total alignment with 1983 positions (58% of the original 3442 positions).
Phylogenetic relationships were inferred by Bayesian inference (BI) and maximum likelihood (ML) based on the concatenated alignment (Online Resource 2). The BI analysis was run with MrBayes 3.2.6 (Ronquist and Huelsenbeck 2003) for two runs over four parallel chains of 1.2 million generations, with trees sampled every 400 generations after a 10% burn-in. Likelihood profiles were examined to verify that the burn-in was appropriate. Run convergence was confirmed when the average standard deviation was < 0.01 with effective sample sizes > 200. Model Finder (Kalyaanamoorthy et al. 2017) in IQ-TREE2 (Minh et al. 2020) was used to select the best-fit substitution model based on the Bayesian information criterion (BIC) (Nguyen et al. 2014): GTR + F + I + R3. Nodal support was assessed with Bayesian posterior probabilities (BPP). The ML analysis was run in RAxML (Stamatakis et al. 2008) with 1000 bootstrap replicates. Phylogenetic trees were visualised in Figtree v1.4.4, with the outgroup Rhamphospora nymphaeae (Dossansiales) as used by Piepenbring et al. (2020).
Morphology
Fresh and dried leaf specimens were examined with a LEICA DLMB microscope (Leica Microsystems, Germany) for basidia, basidiospores, conidia, and hyphae. Colony characteristics of the yeast-like cultures on PDA were recorded. Conidial measurements were taken from PDA cultures (if available) and leaf surfaces. The range of dimensions of morphological features was determined (n = 50 unless indicated otherwise). The colours of cultures were described with the colour chart of Rayner (1970).
Results
Sample collection and isolates
Forty sites in eastern Australia were surveyed for Aneilema acuminatum, Commelina spp., and Pollia crispata with white leaf lesions. Pollia crispata was the only Pollia species recorded during the surveys and was observed growing at 22 sites. At these sites, white leaf spots with white abaxial sides were found on P. crispata at 11 sites (17 detections) and remained undetected at 11 sites. Aneilema acuminatum was the only Aneilema species recorded during surveys and was observed growing at 15 sites. At these sites, white leaf spots with white abaxial sides were found on A. acuminatum at 7 sites (9 detections) and were undetected at 8 sites (Fig. 1; Table 1). Kordyana spp. were not found on Commelina spp. in eastern Australia. Cultures of Kordyana were successfully obtained from five fresh leaf specimens of P. crispata (BRIP 75725, BRIP 75719, BRIP 75718, BRIP 75717, and BRIP 75712) and one from a leaf of A. acuminatum (BRIP 75724).
Distribution of white leaf spot symptoms detected on two Commelinaceae species native in eastern Australia (NSW, QLD). a Pollia crispata observed growing at 22 sites (coloured points), white leaf spots detected n = 17 at 11 different sites, undetected n = 11; b Aneilema acuminatum observed growing at 15 sites (coloured points), white leaf spots detected n = 9 at 7 different sites, undetected n = 8. Grey points represent records of the host plants downloaded from Atlas of Living Australia (ALA) (https://www.ala.org.au/)
Phylogenetic analyses
The BI and ML trees produced similar topologies. The phylogenetic analysis based on the concatenated ITS, LSU, and SSU alignment revealed three well-supported clades that we consider to represent novel species of Kordyana (Fig. 2). Each of these three clades contained only Australian specimens, and members of each clade were restricted to A. acuminatum, C. ensifolia, or P. crispata, respectively. Sequences of Kordyana spp. formed a well-supported clade sister to Marantokordyana.
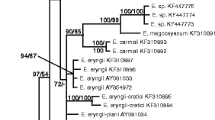
Phylogenetic tree based on BI analysis of the alignment of the combined ITS, LSU, and SSU rDNA sequences from selected species of Exobasidiales. Rhamphospora nymphaeae (CBS 172.38) was used as an outgroup. Numbers on the branches indicate Bayesian posterior probabilities (> 0.70) and ML bootstrap support values (> 70%). The scale bar represents the number of substitutions per site. Novel taxa are shown in bold. References to sequences obtained from ex-type cultures are marked by an asterisk (*)
Taxonomy
Three new species of Kordyana from Australia are described based on the phylogenetic analysis, morphology, and host species. Molecular barcodes and morphological descriptions, together with information about their known host range and distribution, are provided. The asexual life stage of K. brasiliensis is described for the first time.
Kordyana luteoalba Zeil-Rolfe, Gooden, G.C. Hunter, C.C. Linde & R.G. Shivas, sp. nov., Fig. 3.
Kordyana luteoalba (BRIP 75725a) on Pollia crispata. a Adaxial leaf surface with lesions; b abaxial leaf surface with lesion; c two sterigmata with basidiospores; d bipolar germination of two two-celled basidiospores; e ball of basidia; f conidia, one budding (arrow); g basidia, one with probasidial swelling; h colony on PDA after 8 wks. Scale bars: c–g = 10 µm; h = 1 mm
MycoBank: MB 852430.
Holotype: Australia. New South Wales, Macquarie Pass National Park, 34°34'10" S 150°40'19" E, rainforest, on leaves of Pollia crispata, 2 Aug. 2022, I. Zeil-Rolfe & J. Lester (BRIP 75725a includes a culture permanently preserved in a metabolically inactive state); ITS, LSU, and SSU rDNA sequences GenBank OR614363, OR802996, and OR616661, respectively. For data on additional specimens examined, see Table 1.
Etymology: Refers to the yellow leaf lesions surrounded by a distinctive white halo.
Leaf lesions amphigenous, circular to irregular, solitary or coalescent. Sporulation hypophyllous, leaf lesions at first pale yellow to orange with a white to cream halo around the lesion, the centre becoming brown with age, with white balls of basidia covering the lesion. Substomatal chambers in the lesions filled with dense fungal cells, predominantly thick hyphae of (2–) 3–4 (–4.5) µm width (n = 32). Basidia emerge through the stomatal openings from substomatal stromata, densely packed, pyriform to clavate to cylindrical, occasionally with probasidial swellings, (16–) 21–36.5 (–60) × (2.5–) 3.5–5 (–6) µm, bisterigmatic. Sterigmata straight or curved, splayed slightly outwards with a slight bulge, (3–) 4.5–7 (–9.5) × (1.5–) 2–3 (–3.5) µm, carrying a single basidiospore each. Basidiospores two-celled with a central septum at maturity or one-celled prior to germination, liberated in pairs or singly, hyaline, oblong to reniform, apex rounded, slightly narrowed towards the hilum, (11–) 12–14 (–16) × (3.5–) 4–5 (–6) µm, germinating from one or both poles. Germ tubes 1–2 µm wide (n = 7), often coiled, branched, hyphae forming conidia on sterigmata-like outgrowths. Conidia one-celled, fusiform or acerose, (2.5–) 3.5–6.5 (–9) × 0.5–1.5 µm, hyaline. Colonies on PDA after 14 days of growth at 22 °C in the dark reach 7 mm diam., flat, gelatinous, corrugated, central colony Sienna (8) in colour and progresses outward to Luteous (12) and a Pale Luteous (11) border, comprised of conidia and hyphae, conidia forming on sterigmata-like outgrowths from hyphae or via budding.
Notes: Kordyana luteoalba infects leaves of P. crispata in shaded rainforest habitats extending from south-east Queensland south to the Shoalhaven-Illawarra region in New South Wales. Kordyana luteoalba is phylogenetically distinct from all other sequenced Kordyana species and is restricted to P. crispata. Kordyana luteoalba is most closely related to K. tradescantiae (only 2% sequence divergence in LSU). Morphologically, K. tradescantiae differs from K. luteoalba by the germination of basidiospores from the sides of spores in addition to the poles (Begerow et al. 2002), presence of paraphyses (Raciborski 1900), shorter basidia (10–15 µm), shorter sterigmata (3.5–5 µm) and shorter conidia (3–5 µm). Piepenbring et al. (2020) note that probasidial swellings have been observed in K. tradescantiae; however, they are not reported or described in descriptions provided by Petrak (1950), Raciborski (1900), Gómez and Kisimova-Horovitz (1997), or Piepenbring et al. (2010) (see Online Resource 3).
Kordyana polliae on leaves of P. secundiflora in Indonesia (Gäumann 1922) is the only other species recorded on Pollia. Kordyana polliae differs from K. luteoalba by the presence of paraphyses, absence of conidia and probasidial swellings, presence of one-celled basidiospores only, and slightly longer and wider basidiospores (15–21 µm × 5–8 µm) (Gäumann 1922). Although K. polliae lacks DNA sequence data, we propose that the Australian specimens represent a novel species as they can be differentiated from K. polliae by morphological features, host species, and geographical distribution.
Kordyana occidentalis Zeil-Rolfe, Gooden, G.C. Hunter C.C. Linde & R.G. Shivas, sp. nov., Fig. 4.
Kordyana occidentalis (BRIP 65743) on Commelina ensifolia. a Dried leaves with lesions; b basidia emerging from a stomatal chamber; c basidia, one with sterigmata forming basidiospores, one with probasidial swelling (arrow); d bipolar germination of basidiospores (arrows); e conidium on a sterigmata-like outgrowth (arrow); f conidia. Scale bars: b–f = 10 µm
MycoBank: MB 852914.
Holotype: Australia, Western Australia, Geikie Gorge, on leaves of Commelina ensifolia, 20 Apr. 2017, B. Lemana, K. Vánky, M.J. Ryley, S.M. Thompson, M.D.E. Shivas & R.G. Shivas (BRIP 65743); ITS and LSU rDNA sequences GenBank OR791408 and OR764856, respectively. Additional specimens examined are shown in Table 1.
Etymology: Refers to the known distribution of this species in Western Australia.
Leaf lesions amphigenous, usually circular, coalescing into larger, irregular lesions, sporulation hypophyllous. Balls of basidia suprastomatal, densely packed with basidia of various ages, (42–) 53 –84 (–95) × (33–) 44–72 (–90) µm (n = 19). Basidia holobasidia, single celled and hyaline, cylindrical with some probasidial swellings, bisterigmatic, (10.5–) 14.5–29 (–48) × (2.5–) 3.5–5.5 (–6.5) µm (n = 33). Sterigmata conical with a slight bulge in the middle, straight or slightly curved, (2.5–) 3–4.5 (–5.5) × (1–) 2–2.5 (–3) µm (n = 47), with a single basidiospore. Basidiospores liberated singly or in pairs, two-celled with a central septum at maturity or one-celled prior to germination, oblong to reniform, apex round, base narrower, (7–) 9–11.5 (–13) × (2–) 3–4 (–6) µm, germinating with thin hyphae from one or both poles ca. 1 µm wide, forming conidia on sterigmata-like outgrowths. Conidia abundant on the surface of leaf lesions, single celled, rod-shaped to fusiform, apex curved, base acute, (2.5–) 3–5 (–6.5) × 0.5–1.5 µm. Conidial budding was not observed. No cultures were obtained (isolations were not attempted from dried leaf samples).
Notes: Kordyana occidentalis infects the leaves of C. ensifolia in the Kimberley region of northern Western Australia (Table 1). Three other Kordyana species have been described from Commelina, namely K. celebensis on C. benghalensis in Indonesia (Gäumann 1922, see Begerow et al. 2002 for reference sequence data); K. commelinae on C. nudiflora in Sri Lanka (Petch 1922, see Park et al. 2021 for reference sequence data); and K. polliae var. microspora on C. maculata in India (Narendra and Rao 1977, without sequence data).
Kordyana commelinae is phylogenetically most closely related to K. occidentalis but differs morphologically from K. occidentalis by the occasional formation of three sterigmata per basidium (observed by Dudka 2023), an absence of probasidial swellings, conidial shape (described as globose by Petch 1922 and linear and acicular by Park et al. 2021), slightly narrower basidiospores (on av. 2.3 µm), and slightly shorter conidia (on av. 3.2 µm) (see Online Resource 3). Kordyana celebensis can be distinguished from K. occidentalis by phylogenetic analysis and differs by the absence of probasidial swellings and conidia, formation of one-celled basidiospores only, longer basidia (max. 60 µm), and slightly longer basidiospores (9–14 µm) (Gäumann 1922). Kordyana polliae var. microspora lacks sequence data, however, can be distinguished from K. occidentalis by host species, geographic distribution, and morphological characteristics by the presence of paraphyses, presence of one-celled basidiospores only, absence of probasidial swellings, and shorter basidiospores (6.7–7.6 µm) (Narendra and Rao 1977). Therefore, we propose that the Australian specimens on C. ensifolia represent a novel species as supported by morphological differences, unique host species, and distinct geographic range.
Kordyana spectabilis Zeil-Rolfe, Gooden, G.C. Hunter C.C. Linde & R.G. Shivas, sp. nov., Fig. 5.
Kordyana spectabilis (BRIP 75724a) on Aneilema acuminatum. a Lesions on the adaxial leaf surfaces of several leaves of A. acuminatum plant; b symptoms on the abaxial leaf surface; c basidia; d two basidiospores; e single two-celled basidiospore; f conidia, one budding (arrow); g basidiospore germinating from poles to form conidia (arrows) on small sterigmata-like outgrowths; h colony on PDA after 8 weeks. Scale bars: c–g = 10 µm; h = 1 mm
MycoBank: MB 852915.
Holotype: Australia. New South Wales, Darkwood Road, rainforest, on leaves of Aneilema acuminatum, 30°27'01" S 152°36'39" E, 5 Jun. 2022, I. Zeil-Rolfe & J. Lester (BRIP 75724a includes a culture permanently preserved in a metabolically inactive state); ITS, LSU, and SSU rDNA sequences GenBank OR614368, OR802993, and OR616657, respectively. Additional specimens examined are shown in Table 1.
Etymology: Refers to the brightly coloured leaf lesions on the host.
Leaf lesions amphigenous, circular to irregular, solitary or coalescent. Sporulation hypophyllous, leaf lesions at first yellowish orange with a cream halo, later brown or orange with a pale yellowish green halo, covered by not pigmented balls of basidia. Basidia emerge through stomata from substomatal stroma, densely packed, clavate to cylindrical, (10–) 12–21 (–26) × (1–) 2–3 (–4) µm (n = 28), lacking probasidial swellings, bisterigmatic. Sterigmata straight or curved, slightly swollen, (2–) 3.5–5 (–5.5) × (0.7–) 1–1.5 (–2) µm (n = 46), with a single basidiospore each. Basidiospores one-celled or occasionally two-celled with a central septum, liberated in pairs or singly, hyaline, oblong to reniform or slightly allantoid, apex rounded, slightly narrowed towards hilum, (10–) 11–13 (–16) × (2–) 3–4 (–5) µm, germinating from one or both poles. Germ tubes 0.5–1.5 µm wide, branched, forming conidia on sterigmata-like outgrowths. Conidia one-celled, fusiform or slightly curved, (2.5–) 4–6.5 (–9) × 0.5–1.5 µm, hyaline. Colonies on PDA after 14 days growth at 22 °C in the dark reaching 7 mm diam., flat, gelatinous, colony centre Luteous (12) becoming paler towards the colony border presenting as Pale Luteous (11), comprised of conidia and hyphae, conidia forming on sterigmata-like outgrowths from hyphae or via budding.
Notes: Kordyana spectabilis infects leaves of A. acuminatum in shaded rainforest habitats in eastern Australia from south-east QLD to the Shoalhaven-Illawarra region, NSW. Kordyana spectabilis is phylogenetically distinct from all Kordyana species for which molecular barcodes are available. There are only two other reports of Kordyana on Aneilema. One is K. aneilematis with the holotype collected on A. angustifolium in Taiwan (Sawada 1929 as K. aneilemae). The second is Kordyana sp. collected on A. umbrosum var. ovato-oblongum in Ecuador (collected by H. Sydow 1937 as K. tradescantiae according to Petrak 1950), which Petrak (1950) re-examined and noted that the lack of described morphological features meant species identification was not possible.
Morphologically, K. spectabilis is most similar to K. aneilematis and K. commelinae, lacking both paraphyses and probasidial swellings (see Online Resource 3). Kordyana aneilematis differs from K. spectabilis by longer and wider basidia (34–51 µm × 4–5 µm), slightly longer and wider sterigmata (5–7 µm × 2.5 µm), shorter basidiospores (10–14 µm), and the absence of conidia (Sawada 1929). Kordyana commelinae can be distinguished from K. spectabilis phylogenetically and morphologically by longer basidia (on av. 20 µm, Petch 1922; 25.9 µm, Park et al. 2021; 20–37 µm, Dudka 2023), narrower basidiospores (on av. 2.3 µm), and shorter conidia (on av. 3.2 µm) (Park et al. 2021) (see Online Resource 3). Despite the absence of sequence data from K. aneilematis for comparison, we have proposed the Australian specimens represent a novel species based on distinct morphological features, host species, and geographic distributions.
Kordyana brasiliensis D.M. Macedo, O.L. Pereira & R.W. Barreto, Australas. Pl. Pathol. 45 (1): 51 (2016), Fig. 6.
Kordyana brasiliensis (BRIP 75726a) on Tradescantia fluminensis. a Adaxial leaf surface with lesions; b abaxial leaf surface with lesions; c basidia emerging from a stoma to form a suprastomatal ball; d basidium with probasidial swelling; e basidium with sterigmata and basidiospores; f conidia; g budding conidium; h germinating basidiospores; i colony on PDA after 12 weeks. Scale bars: c–h = 10 µm; i = 1 mm
MycoBank: MB 518070.
Holotype: Brazil. Minas Gerais, Viços, on living leaves of Tradescantia fluminensis, 10 Dec. 2009, D. M. Macedo (VIC 3136).
Leaf lesions amphigenous, circular to irregular, solitary or coalescent. Sporulation hypophyllous, leaf lesions pale white lesions becoming yellow to orange covered in white balls of basidia. Balls of basidia suprastomatal, spherical, densely packed with basidia of various developmental stages, germinated basidiospores and hyphae, (66–) 84–151 (–201) × (68–) 87–142 (–194) µm (n = 40). Basidia holobasidia, hyaline, cylindrical, some with probasidial swellings, bisterigmatic, (10–) 18–29 (–39) × (3–) 4–5 (–6) µm. Sterigmata conical in shape, straight or slightly curved, splayed slightly outwards, generally with a slight bulge, (3–) 3.5–5 (–6.5) × (1.5–) 2–3 (–3.5) µm, with a single basidiospore each. Basidiospores liberated singly or in pairs, one-celled or occasionally two-celled with a central septum after germination, oblong to reniform, somewhat allantoid, apex round, base narrowed, (12–) 14–18 (–19.5) × (3–) 3.5–3.5 (–5.5) µm, germinating by thin hyphae from one or both poles 1–2 µm wide forming conidia on sterigmata-like outgrowths. Conidia occasionally observed on the surface of leaf lesions, one-celled, fusiform or rod-shaped, often falcate, (4–) 5.5–12 (–17.5) × (0.5–) 1–1.5 (–2) µm, germinating to form hyphae. Colonies on PDA after 14 d growth at 22 ºC reaching 7.5 mm diam., flat, Pale Luteous (11) in the centre of the colony and transitioning to Straw (46) at the colony border, comprised of conidia and hyphae, conidia forming on sterigmata-like outgrowths from hyphae or via budding.
Specimen examined: On leaves of Tradescantia fluminensis. Australia. Australian Capital Territory, 35°16'301" S 149°06'49" E, cultured from material imported from Universidade Federal de Viçosa, MG, Brazil, 12 Jan. 2022, I. Zeil-Rolfe (BRIP 75726a).
Notes: Kordyana brasiliensis was introduced from its native range in Brazil into an Australian quarantine facility at CSIRO Black Mountain Science and Innovation Park, Australian Capital Territory (Morin et al. 2022). Since its field release in Australia in 2019, K. brasiliensis has spread across riparian habitats in rainforest and wet-sclerophyllous forests from north-eastern NSW southwards to the Dandenong Ranges in Victoria (ALA 2023; GBIF 2024; CSIRO 2023).
Discussion
The taxonomic resolution of Kordyana is hampered as most species lack molecular barcodes. Further specimens and DNA sequence data are needed to clarify the taxonomy of this genus (Piepenbring et al. 2020). In this study, we provide molecular barcodes for three newly recognised Australian species, K. luteoaba, K. spectabilis, and K. occidentalis, together with phenotypic data, host associations, and geographic distributions. The three Kordyana species described in this study fit within the accepted concept of Kordyana, forming bisterigmatic basidia in suprastomatal balls, each basidium bearing two basidiospores which germinate generally becoming two-celled with hyphae that form conidia.
Kordyana luteoalba, K. occidentalis, and K. spectabilis are morphologically similar and can be differentiated by the absence of probasidial swellings and shorter basidia in K. spectabilis, the rod-shaped conidia in K. occidentalis and the longer basidia, slightly longer and wider sterigmata, and longer and wider basidiospores in K. luteoalba. The species can be differentiated from all other described Kordyana species from Commelinaceae hosts by phylogenetic data (if available), morphology, unique host species, and geographical distributions. Notably, all three Australian species lack paraphyses which are described in several Kordyana species including K. tradescantiae, K. polliae, K. polliae var. microspora, and K. commelinae (Raciborksi 1900; Gäumann 1922; Narendra and Rao 1977; Dudka 2023).
Distinguishing Kordyana species, particularly those lacking DNA sequence data, is complicated by the lack of complete morphological descriptions for several species (Piepenbring et al. 2020) and variability in the intraspecific morphological descriptions of collections (e.g. K. tradescantiae and K. commelinae). Descriptions of morphological details of K. commelinae and K. tradescantiae differ depending on the specimen studied as well as the author. For example, the description of K. commelinae from Russia includes the presence of paraphyses and occasionally basidia with three sterigmata (Dudka 2023), which differs to descriptions of K. commelinae from Sri Lanka and Korea (Park et al. 2021; Petch 1922). Specimens of K. tradescantiae are described as lacking paraphyses (e.g. Raciborksi 1900, Gómez and Kisimova-Horovitz 1997; Piepenbring et al. 2010) and probasidial swellings (e.g. Raciborksi 1900, Petrak 1950; Gómez and Kisimova-Horovitz 1997) except for Petrak (1950) who observed paraphyses and drawings by F. Oberwinkler (Piepenbring et al. 2020) and Piepenbring et al. (2010) who illustrated probasidial swellings. There are issues, however, with the use of probasidial swellings to differentiate taxa as they are commonly overlooked and common among Brachybasidiaceae species (Piepenbring et al. 2020).
Measurements of morphological features have been traditionally used to differentiate Kordyana species (e.g. Macedo et al. 2016; Park et al. 2021); however, comparisons of collections demonstrate that these also vary within a species. For example, Piepenbring et al. (2010) record significantly longer and wider basidiospores of K. tradescantiae compared to Raciborski (1900), Petrak (1950), and Gómez and Kisimova-Horovitz (1997). Basidia and basidiospore lengths vary between different descriptions of collections of K. commelinae (Petch 1922; Park et al. 2021; Dudka 2023). These issues highlight that the morphological approach to fungal classification of Kordyana species is unreliable and requires DNA barcodes to reliably distinguish species.
The discovery of K. luteoalba, K. occidentalis, and K. spectabilis, each apparently restricted to one of three native Australian species of Commelinaceae (P. crispata, C. ensifolia, and A. acuminatum, respectively), was a consequence of surveys before and after the release of K. brasiliensis. Kordyana spectabilis and K. luteoalba were found to be broadly distributed across eastern Australia, and K. occidentalis was only found in the Kimberly region in Western Australia. In 2019, K. brasiliensis was introduced into New South Wales and Victoria in eastern Australia as a classical biocontrol agent for T. fluminensis. The co-occurrence of white leaf lesions on T. fluminensis as well as on A. acuminatum and P. crispata in eastern Australia raised the concern that K. brasiliensis may have extended its host range to native Australian Commelinaceae. This study provides strong evidence that K. brasiliensis remains highly host-specific to T. fluminensis.
The inclusion of additional sequence data for K. brasiliensis, K. spectabilis, K. luteoalba, and K. occidentalis has clarified the taxonomic relationship between Kordyana and Dicellomyces. We found that Kordyana formed a well-supported clade sister to Marantokordyana and not a sister to D. gloesporus. Further, M. nicotianae was sister to D. gloesporus in our analysis rendering Meira polyphyletic.
Dimorphic life cycles that include an asexual yeast-like stage (conidia) are known for several taxa in the Brachybasidiaceae, including K. spectabilis, K. luteoalba, and K. occidentalis (in our study), as well as Exobasidium, Marantokordyana, and Meira (Ingram et al. 2019; Limtong et al. 2017; Park et al. 2021; Piepenbring et al. 2020; Tanaka et al. 2008). Meira spp. and one unknown species of Kordyana have been isolated from leaf phylloplanes (Albu 2012; Limtong et al. 2017; Tanaka et al. 2008). In a study of the diversity of phylloplane basidiomycetous yeasts on fern leaves in the USA, Albu (2012) used the spore-fall method and isolated Kordyana sp. from a senescent fertile frond of Pelazoneuron kunthii (Polypodiales). This may indicate that Kordyana spp. can survive epiphytically on the leaf phylloplane, either as basidiospores or conidia. Ungerminated conidia were often seen on the surface of leaf lesions around the suprastomatal balls of K. spectabilis, K. luteoalba, and K. occidentalis, in contrast to K. brasiliensis, which produced few conidia. The growth of these pathogens in a yeast-like state is thought to be advantageous for survival, dispersal, and multiplication (Bauer et al. 2001; Ingram et al. 2019).
This study demonstrates that knowledge of local fungal biodiversity can be increased by ecological monitoring of plant pathogens. We describe three previously unknown species of Kordyana on native Australian Commelinaceae, two of which co-occurred with the exotic and weedy T. fluminensis. Kordyana spectabilis, K. luteoalba, and K. occidentalis were discovered as a direct consequence of surveys before and after the release of K. brasiliensis. These three novel species are the first published records of Kordyana spp. on native Commelinaceae in Australia. Several other native Australian Commelinaceae species remain to be studied to determine if they also are hosts of Kordyana species. The collection of fresh specimens of Kordyana is required to further clarify the taxonomy, the intraspecific morphological variability, and distribution of these pathogens both in Australia and globally.
Data Availability
All sequence data generated in this study was deposited on GenBank (https://www.ncbi.nlm.nih.gov/genbank/). Alignment of phylogeny is available in Supplementary Information S1.
References
Albu S (2012) A survey of ballistosporic phylloplane yeasts in Baton Rouge, Louisiana. LSU Master’s Thesis, Louisiana State University
Atlas of Living Australia (2023) Kordyana brasiliensis D.M.Macedo, O.L.Pereira & R.W.Barreto. https://bie.ala.org.au/species/https://id.biodiversity.org.au/taxon/fungi/60081342. Accessed 12 November 2023
Australian National Species List (2024) Fungi Name Index (FNI). https://fungi.biodiversity.org.au/nsl/services/search/names. Accessed 26 April 2024
Barreto RW, Evans HC (1988) Taxonomy of a fungus introduced into Hawaii for biological control of Ageratina riparia (Eupatorieae; Compositae), with observations on related weed pathogens. Trans Br Mycol Soc 91:81–97
Bauer R, Begerow D, Oberwinkler F, Piepenbring M, Berbee ML (2001) Ustilaginomycetes. In: McLaughlin DJ, McLaughlin EG, Lemke PA (eds) Systematics and evolution. The Mycota, Springer Berlin, Heidelberg, pp 57–83
Begerow D, Bauer R, Oberwinkler F (2002) The Exobasidiales: an evolutionary hypothesis. Mycol Prog 1:187–199. https://doi.org/10.1007/s11557-006-0018-7
Boekhout T, Theelen B, Housbraken J, Robert V, Scorzetti G, Gafni A, Gerson U, Sztejnberg A (2003) Novel anamorphic mite-associated fungi belonging to the Ustilaginomycetes: Meira geulakonigii gen. nov., sp. nov., Meira argovae sp. nov. and Acaromyces ingoldii gen. nov., sp. nov. Int J Syst Evol Microbiol 53:1655–1664. https://doi.org/10.1099/ijs.0.02434-0
Cao Y, Li P, Zhao J, Wang H, Jeewon R, Bhoyroo V, Aruna B, Lin F, Wang Q (2018) Morph-molecular characterization of Meira nicotianae sp. nov., a novel basidiomycetous, anamorphic yeast-like fungus associated with growth improvement in tobacco plant. Phytotaxa 365:169–181. https://doi.org/10.11646/phytotaxa.365.2.4
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552. https://doi.org/10.1093/oxfordjournals.molbev.a026334
Commonwealth Scientific and Research Organisation (CSIRO) (2023) Wandering trad biological control. https://research.csiro.au/wandering-trad/our-research/. Accessed 14 December 2023
Crous PW, Groenewald JZ, Carroll G (2003) Muribasidiospora indica causing a prominent leaf spot disease on Rhus lancea in South Africa. Australas Plant Pathol 32:313–316. https://doi.org/10.1071/AP03007
Crous PW, Wingfield MJ, Schumacher RK, Akulov A, Bulgakov TS, Carnegie AJ, Jurjević Ž, Decock C, Denman S, Lombard L, Lawrence DP, Stack AJ, Gordon TR, Bostock RM, Burgess T, Summerell BA, Taylor PWJ, Edwards J, Hou LW, Cai L, Rossman AY, Wöhner T, Allen WC, Castlebury LA, Visagie CM, Groenewald JZ (2020) New and interesting fungi. Fungal Syst Evol 6:157–231. https://doi.org/10.3114/fuse.2020.06.09
Dudka VA (2023) The first record of Kordyana commelinae (Brachybasidiaceae, Exobasidiomycetes) on Commelina communis in the Russian Far East (Primorsky Krai). Nova Hedwigia 116:349–362. https://doi.org/10.1127/nova_hedwigia/2023/0875
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Gäumann E (1922) Uber die Gattung Kordyana Rac. Ann Mycol 20:257–271
Global Biodiversity Information Facility (2024) Kordyana brasiliensis D.M. Macedo, O.L. Pereira & R.W. Barreto. https://www.gbif.org/species/9801103. Accessed 12 Nov 2023
Gómez LD, Kisimova-Horovitz L (1997) Basidiomicetos de Costa Rica. Exobasidiales, Cryptobasidiales. Notas históricas, taxonómicas y fitogeográficas. Rev Biol Tro 45:1293–1310
Gruèzo SW (1990) The genus Kordyana Rac. (Exobasidiaceae) in the Philippines. Nat Hist Bull Siam Soc 28:89–92
Ingram RJ, Ludwig HD, Scherm H (2019) Epidemiology of Exobasidium leaf and fruit spot of rabbiteye blueberry: pathogen overwintering, primary infection, and disease progression on leaves and fruit. Plant Dis 103:1293–1301. https://doi.org/10.1094/PDIS-09-18-1534-RE
Kalyaanamoorthy S, Minh BQ, Wong TKF, Von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Kottke I, Suárez JP, Herrera P, Cruz D, Bauer R, Haug I, Garnica S (2010) Atractiellomycetes belonging to the ‘rust’ lineage (Pucciniomycotina) form mycorrhizae with terrestrial and epiphytic neotropical orchids. Proc R Soc B 277:1289–1298. https://doi.org/10.1098/rspb.2009.1884
Landcare Research (2020) Biocontrol success in the Waingaro Valley. Weed Biocontrol: what’s New? 91, 6. https://www.landcareresearch.co.nz/uploads/public/Publications/Weed-biocontrol/weed_biocontrol_issue91.pdf. Accessed 17 November 2021
Lemoine F, Correia D, Lefort V, Doppelt-Azeroual O, Mareuil F, Cohen-Boulakia S, Gascuel O (2019) NGPhylogeny.fr: new generation phylogenetic services for non-specialists. Nucleic Acids Res 47:260–265. https://doi.org/10.1093/nar/gkz303
Limtong S, Polburee P, Chamnanpa T, Khunnamwong P, Limtong P (2017) Meira siamensis sp. nov., a novel anamorphic ustilaginomycetous yeast species isolated from the vetiver grass phylloplane. Int J Syst Evol Microbiol 67:2418–2422. https://doi.org/10.1099/ijsem.0.001969
Macedo DM, Pereira OL, Hora Júnior BT, Weir BS, Barreto RW (2016) Mycobiota of the weed Tradescantia fluminensis in its native range in Brazil with particular reference to classical biological control. Australas Plant Pathol 45:45–56. https://doi.org/10.1007/s13313-015-0388-x
Maier W, Khoza T, Harmse N, Wingfield BD, Wingfield MJ (2006) A disease epidemic on Zizyphus mucronata in the Kruger National Park caused by Coniodictyum chevalieri. Stud Mycol 55:279–88. https://doi.org/10.3114/sim.55.1.279
Matheny PB, Gossmann JA, Polona Z, Arun Kumar TK, Hibbett DS (2006) Resolving the phylogenetic position of the Wallemiomycetes: an enigmatic major lineage of Basidiomycota. Can J Bot 84:1794–1805. https://doi.org/10.1139/b06-128
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, Von Haeseler A, Lanfear R, Teeling E (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534. https://doi.org/10.1093/molbev/msaa015
Moncalvo JM, Wang HH, Hseu RS (1995) Phylogenetic relationships in Ganoderma inferred from the internal transcribed spacers and 25S ribosomal DNA sequences. Mycologia 87:223–238. https://doi.org/10.1080/00275514.1995.12026524
Morin L, Incoll B, Lester J, Zeil-Rolfe I, Gooden B (2022) Biological control of the invasive plant Tradescantia fluminensis with the fungus Kordyana brasiliensis in Australia: host range and initial releases. Biol Control 172:104978. https://doi.org/10.1016/j.biocontrol.2022.104978
Narendra DV, Rao VG (1977) A new record of Kordyana rac. (Exobasidiaceae) from Maharashtra. Curr Sci 46:677–678
Nasr S, Lutz M, Amoozegar MA, Eparvier V, Stein D, Fazeli SAS, Yurkov A (2019) Graphiola fimbriata: the first species of Graphiolaceae (Exobasidiales, Basidiomycota) described only based on its yeast stage. Mycol Progress 18:359–368. https://doi.org/10.1007/s11557-018-1450-1
Nguyen V, Phung D, Nguyen X, Edu X, Venkatesh S, Bui HH (2014) Bayesian nonparametric multilevel clustering with group-level contexts. Proceedings of the 31st International Conference on Machine Learning 32:288–296
Park JH, Jung BN, Choi IY, Shin HD (2021) Kordyana commelinae associated with white smut-like disease on Commelina communis and C. minor in Korea. Mycobiology 49:275–279. https://doi.org/10.1080/12298093.2021.1923617
Petch T (1922) Additions to Ceylon fungi (II). Ann Royal Botanical Gardens Peradeniya 7:279–320
Petrak F (1950) Beiträge zur Pilzflora von Ekuador. Sydowia 4:451–452
Piepenbring M, Espinoza J, Saldaña L, Cáceres O (2010) New records, host plants, morphological and molecular data of Exboasidiales (Basidiomycota) from Panama. Nova Hedwigia 91:231–242. https://doi.org/10.1127/0029-5035/2010/0091-0231
Piepenbring M, Nold F, Trampe T, Kirschner R (2012) Revision of the genus Graphiola (Exobasidiales, Basidiomycota). Nova Hedwigia 94:67–96. https://doi.org/10.1127/0029-5035/2012/0094-0067
Piepenbring M, Hartmann M, Hofmann TA, Lutz M (2020) Two new species in a new genus and a critical revision of Brachybasidiaceae (Exobasidiales, Basidiomycota) in honor of Franz Oberwinkler. Mycol Prog 19:351–365. https://doi.org/10.1007/s11557-020-01564-w
PlantNET (2022) New South Wales Flora Online. https://plantnet.rbgsyd.nsw.gov.au/. Accessed 6 May 2024
Queensland Government (2024) WildNet – Species Lists. https://www.qld.gov.au/environment/plants-animals/species-information/species-list. Accessed 6 May 2024
Raciborski M (1900) Parasitishe Algen und Pilze Java’ s. II Teil Bibl Mycol 37:1–46
Rayner RW (1970) A mycological colour chart. Kew and British Mycological Society, Commonwealth Mycological Institute
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Rush TA, Aime MC (2013) The genus Meira: phylogenetic placement and description of a new species. Antonie Van Leeuwenhoek 103:1097–1106. https://doi.org/10.1007/s10482-013-9889-1
Sawada K (1929) Fungi from Taiwan (no. 27). Transactions of the Natural History Society of Formosa 19:31–39
Sepúlveda G, Arismendi M, Huanca-Mamani W, Cárdenas-Ninasivincha S, Salvatierra R, Latorre B (2017) Presence of false smut (Graphiola phoenicis (Moug. ex Fr.) Poit.) on Canary date palm (Phoenix canariensis) on Easter Island. Chile Cienc Inv Agr 44:307–311. https://doi.org/10.7764/rcia.v44i3.1787
Singh A, Nautiyal MC, Gautam AK, Singh PN, Singh SK (2020) Taxonomic and phylogenetic analysis of Clinoconidium lauracearum (Cryptobasidiaceae) producing galls on fruits of Cinnamomum tamala (Lauraceae) in India. Phytotaxa 450:73–84. https://doi.org/10.11646/phytotaxa.450.1.5
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. System Biol 57:758–771. https://doi.org/10.1080/10635150802429642
Tamura K, Stecher G, Kumar S (2021) MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol and Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Tanaka E, Shimizu K, Imanishi Y, Yasuda F, Tanaka C (2008) Isolation of basidiomycetous anamorphic yeast-like fungus Meira argovae found on Japanese bamboo. Mycoscience 49:329–333. https://doi.org/10.1007/S10267-008-0429-1
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
Wang QM, Begerow D, Groenewald M, Liu XZ, Theelen B, Bai FY, Boekhout T (2015) Multigene phylogeny and taxonomic revision of yeasts and related fungi in the Ustilaginomycotina. Stud Mycol 81:55–83. https://doi.org/10.1016/j.simyco.2015.10.004
Western Australian Herbarium (2024) Florabase—the Western Australian Flora. Department of Biodiversity, Conservation and Attractions. https://florabase.dbca.wa.gov.au/. Accessed 6 May 2024
White TJ, Bruns TD, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA Genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR Protocols: a guide to methods and applications. Academic Press, London, pp 315–322
Winston RL, Schwarzländer M, Hinz HL, Day MD, Cock MJW, Julien MH (2021) Biological control of weeds: a world catalogue of agents and their target weeds. https:// www.ibiocontrol.org/catalog/. Accessed 12 November 2023
Yasuda F, Yamagishi D, Akamatsu H, Izawa H, Kodama M, Otani H (2006) Meira nashicola sp. nov., a novel basidiomycetous, anamorphic yeastlike fungus isolated from Japanese pear fruit with reddish stain. Mycoscience 37:36–40. https://doi.org/10.1007/s10267-005-0266-4
Acknowledgements
Vegetation surveys were undertaken with approval of site managers under NPWS Scientific Licence number SL102231.We would like to acknowledge and thank all natural resource managers (local government area biosecurity officers, Landcare, members of local ‘friends of’ groups, private landholders, and national parks rangers) for their assistance in monitoring site selection and access. We would like to acknowledge and thank both the Western Australian Herbarium and the Queensland Plant Pathology Herbarium (BRIP) for the loan of herbarium specimens. We also wish to thank Yu Pei Tan (BRIP) for her assistance with the submission and lodgement of samples, John Lester for assisting with fieldwork and sample collection, Chris Ormond for the detection of samples in the Bellingen region, and Brian Patterson for the collection of samples in NSW.
Funding
Open access funding provided by CSIRO Library Services. This project was funded by the NSW Government’s Environmental Trust (Project title: Biocontrol Research for Weed Management—Stage 3; Grant Number: 2019/MG/0012) as part of a project focused on the release and evaluation of the approved biological control agent (leaf-smut fungus Kordyana brasiliensis) for Tradescantia fluminensis in New South Wales (July 2020 to June 2023) (https://research.csiro.au/wandering-trad/background/).
Author information
Authors and Affiliations
Contributions
Isabel Zeil-Rolfe, Ben Gooden, Gavin Hunter, and Celeste Linde contributed to study conception and design. Data collection was undertaken by Isabel Zeil-Rolfe and Ben Gooden and data analysis by Isabel Zeil-Rolfe and Celeste Linde. The first draft of the manuscript was written by Isabel Zeil-Rolfe and Roger Shivas and all authors commented and edited the previous versions. The final manuscript was read and approved by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Meike Piepenbring
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zeil-Rolfe, I., Gooden, B., Hunter, G.C. et al. Diversity of Kordyana species (Brachybasidaceae) on Commelinaceae in Australia. Mycol Progress 23, 41 (2024). https://doi.org/10.1007/s11557-024-01981-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-024-01981-1