Abstract
The genus Tuber comprises ectomycorrhizal fungal species producing belowground ascomata, including the gastronomically most prominent hypogeous fungi. Since the discovery and description of new species are ongoing, the proportion of undescribed species can be considerable and the taxonomy of the genus goes often through changes. The taxonomy of the genus Tuber would therefore benefit from a periodic review. Tuber species described in Europe in recent decades include Tuber regianum, T. bernardinii and T. magentipunctatum. The common characteristics of these three species are the relatively small-sized spores with alveolate-reticulate ornamentation, a high number of spores per ascus (most frequently 6–8 spores) and small-sized ascomata with a verrucose-papillate or smooth surface. The present study aimed at examining the morphology and ecology of the three species, and providing a detailed taxonomic description of the Regianum clade using a multilocus phylogenetic analysis. In addition to this, we examined whether the apparently plesiomorphic morphological character states of this phylogenetically basal clade are the result of the conservation of ancestral character states. Our results show that in the case of certain morphological traits of the Regianum clade, the apparently plesiomorphic character states are indeed retained ancestral states, while in others, they are convergently reappeared ones. Furthermore, taking an overlook at all Tuber clades, we found that some morphological characters, such as spore length, number of spores per ascus and ascus length, changed together in the same evolutionary patterns, while others transformed in different ways within the genus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Based on our current knowledge, the family Tuberaceae comprises three lineages, Tuber, Choiromyces and Labyrinthomyces (the lineage previously called Gymnohydnotrya is a distinct family, Geomoriaceae, according to Kraisitudomsook et al. 2020). Tuber and Choiromyces are the genera of Tuberaceae distributed mostly in the Northern Hemisphere (Bonito and Smith 2016; Romero and Blumenfield 2001; Sulzbacher et al. 2017). Dingleya, Reddellomyces, Labyrinthomyces and Nothojafnea, genera of the Labyrinthomyces lineage, are found primarily in the Southern Hemisphere (Bonito and Smith 2016; Agnello 2011; Moreno et al. 2012; Malençon 1973; Pacioni 1984; Trappe et al. 1992). Although species of Choiromyces have a wide distribution, and some of the Southern Hemisphere genera have a remarkable number of species (Bonito and Smith 2016; Le Tacon 2016), their diversity is low in comparison to the genus Tuber.
Tuber species also called “true truffles” (Ascomycota, Pezizales, Tuberaceae) are ectomycorrhizal, hypogeous fungi, some of which have considerable economic and gastronomic importance. The members of the genus are intensively studied across Europe (Pacioni et al. 1991; Iotti et al. 2002; Murat et al. 2004; Leonardi et al. 2019) and North America (Gilkey 1939; Trappe and Castellano 1991; Frank et al. 2006; Guevara-Guerrero et al. 2018), but many Tuber species have been introduced in the last 30 years from previously unexplored regions, for instance the Balkan Peninsula (Polemis et al. 2019) and Asia (Wang et al. 1998; Li et al. 2014; Lin et al. 2018; Fan et al. 2022). Recently, (April 2023) 303 and 307 “Tuber” records, excluding variations, forms and subspecies, were recorded in Mycobank (http://www.mycobank.org/) and Index Fungorum (https://www.indexfungorum.org/) databases, respectively. Although taxonomic revisions and clarifications of all these taxa would probably decrease these numbers, even so, Tuber is the most species-rich genus of the family (Bonito et al. 2013). According to predictions, it embraces at least 180, but possibly up to 220–230 species (Bonito et al. 2010a; Bonito and Smith 2016). Probably due to the hidden ascoma production, the description of new species is still frequent, and the taxonomy of the genus often changes (Bonito et al. 2010a, b, 2013; Polemis et al. 2019; Fan et al. 2022). Even in Europe, where truffle science has the longest history, there are new species descriptions in the last few decades. Three examples of this are the species Tuber regianum Montecchi and Lazzari (1987), T. bernardinii Gori (2003) and T. magentipunctatum Merényi et al. (2017a).
Tuber regianum, T. bernardinii and T. magentipunctatum have a relatively narrow distribution, spanning across the Carpatho-Pannonian region, the Mediterranean region in Europe, Germany and France (Merényi et al. 2017a; Perez 2019; communication of Günther Schier, the first finder of T. bernardinii in Germany). Furthermore, they could be considered rare species, as they are only known from a few locations, and never collected in abundance (Bratek et al. 2013; Merényi 2014). For example, only ~ 0.4% of all records in our hypogeous mycotheca (> 6000 specimens) were identified as T. regianum aff. These three species constitute the Regianum clade which is one of the 13 currently known Tuber clades (Polemis et al. 2019). Most Tuber clades were well described and discussed in comprehensive phylogenetic analyses (Bonito et al. 2010b, 2013; Zambonelli et al. 2016), yet no thorough phylogenetic or taxonomic examination has been carried out on the Regianum clade. Although these three species have already been introduced using molecular data (Merényi et al. 2017a), and a basal position of the clade within the Tuber genus was anticipated (Merényi et al. 2017a; Polemis et al. 2019), a thorough phylogenetic assessment of the clade was never carried out.
The three species were described with similar morphological features, relatively small spores with alveolate-reticulate ornamentation, a dominance of 6–8-spored asci and small-sized ascomata with smooth or papillate surfaces (Montecchi and Sarasini 2000; Gori 2003; Merényi et al. 2017a). Out of these features, the small spore size and 8 spores per ascus are more typical for the sister taxa of the genus Tuber (Gamundí 1971; Zhang and Minter 1989; Trappe et al. 1992) than most Tuber species (Online Resource 1 Table S1). According to our current knowledge of Tuber evolution, alveolate-reticulate spore ornamentation is the ancestral character state of the genus, and possessing 8 spores per ascus can be also considered plesiomorphic as Tuber species evolved from epigeous cup fungi having 8 spores in a uniseriate ascus (Bonito et al. 2013). The co-occurrence of these character states in T. regianum, T. bernardinii and T. magentipunctatum presumably makes these species unique in an evolutionary aspect, and may provide the delimitation of their clade on a morphological basis.
Is the presence of the above-mentioned, plesiomorphic character states a direct consequence of the basal position of the clade, particularly the conservation of the ancestral states? In fungi, there are examples of certain morphological features appearing convergently many times, for instance in spore and fruiting body morphology (Justo et al. 2010; Bonito et al. 2013; Moreno et al. 2013; Varga et al. 2019; Virágh et al. 2022); however, basal lineages may retain several ancestral character states (Quijada et al. 2022). What can the example of this basal and morphologically prominent Tuber clade show in comparison to the other Tuber clades? How reconcilable are morphology and phylogenetics in Tuber species when a character is thought to be ancestral or derived?
The number of morphological characters useful in taxonomic characterisation is relatively low in the case of hypogeous fungal species (Reynolds 2011). As for the genus Tuber, some of its features, such as the loss of active spore discharge, are common to hypogeous pezizalean species. Nevertheless, the variable number of spores per ascus and the relatively large ascospores are unique characters among the species of Pezizales, and the majority of Tuber species have a distinctive spore morphology, which is one of the most applicable characteristics in classical morphological taxonomy supporting species recognition (Bonito and Smith 2016). In the case of Tuber species, the arrangement of ascospores in the asci is random and the asci are wider than that of the hypothesized 8-spored ancestor cup fungus. Nevertheless, the varying number of spores connotes even 8-spored asci in some Tuber species (Wang et al. 1998; Montecchi and Sarasini 2000; Gori 2003; Crous et al. 2017; Polemis et al. 2019). The morphology of the ascoma such as the size and the surface of the ascoma is also often involved in taxonomy (Montecchi and Sarasini 2000). Besides species identification, morphological features could help us to better understand fungal evolutionary trajectories. Ancestral State Reconstruction (ASR) was found to be capable of unravelling evolutionary patterns and estimating plesiomorphic character states of the genus Tuber (Bonito et al. 2013) and other taxa (Samarakoon et al. 2022; Rathnayaka et al. 2023). ASR analyses are often applied to project morphological characters on molecular phylogenetic genealogies to reveal regularities with respect to the evolutionary origin and changes in morphology (Frenzke et al. 2016; Gorin et al. 2021; Starrett et al. 2021).
Our aim was to examine the monophyletic clade of T. regianum, T. bernardinii and T. magentipunctatum to clarify its relation to other, well-defined Tuber clades based on a multilocus phylogeny. Furthermore, we aimed to give a morphological and ecological characterisation of this clade highlighting the features which provide its delimitation from all other Tuber clades. Besides, ASR analyses were performed applying six morphological characters of already described Tuber species. The purpose of our ASR was to test our hypothesis that the co-occurrence of the plesiomorphic morphological character states of the three mentioned species is due to the conservation of ancestral character states. The evolutionary changes of all involved Tuber clades’ morphological traits were also assessed to estimate their evolutionary trajectories. The currently covered aspects of Tuber evolution were inferred, which can be an informative example in terms of the evolution of all hypogeous fungi.
Materials and methods
Sampling
A Microsoft Access database of hypogeous fungal materials (Merényi et al. 2010) comprises information of belowground fungal specimens. These specimens were collected by the members of the First Hungarian Truffle Society (EMSzE) in the Carpatho-Pannonian region and deposited in the mycotheca (ZB) of the society (Bratek et al. 2013). In addition to the collection data of the fruiting bodies, the database stores pedological and coenological information as well. Soil samples were collected from around the belowground fruiting bodies (layer A) and were analysed by the laboratories of the Hungarian Central Agricultural Office. Botanical data refer to the plant species and their abundances in a ~ 10 × 10 m area around the truffle nest (according to Braun-Blanquet 1927), and used for the coenosis of this area completed according to Borhidi (2003). Of the Tuber regianum aff. materials investigated in this study, 16 derived from this database (gathered in Hungary, Romania and Slovakia), while an additional 18 were collected in Greece, Spain and Italy. Hence, in total, 34 T. regianum aff. materials (including holotypes and isotypes) were included in our examinations (Table 1, Online Resource 1 Table S2).
Morphological measurements
Fresh or dried ascomata of Tuber regianum aff. materials were characterised by macromorphological characters as shape and size of ascomata, the colour and surface of the peridium, the colour of the gleba (applying the Colour Identification Chart of the Royal Botanic Garden of Edinburgh, RBGE 1969) and odour. Preparations for micromorphological measurements were mounted in polyvinyl alcohol-lactic acid-glycerol (PVLG) medium (Koske and Tessier 1983) or in 5% KOH. Micromorphological characters were determined by the structure and size of the peridium and peridial cells, size of ascospores and asci, type and height of ornamentation and the number of spores per ascus. The measurements of the quotient value (Q) and volume (Vol) of spores, the distribution of spore number per ascus and the peridial cells’ dimensions were carried out according to the methods by Merényi et al. (2017b) with the following modifications: spore width (SPW) and spore length (SPL) were assessed based on 8-spored asci, and the ten largest isodiametric peridial cells were measured. The morphological characters were considered on average per sample (herbarium material) in the morphological descriptions. Micromorphological measurements were accomplished with a Nikon Optiphot-2 research microscope at various magnifications depending on the type of structures. For scanning electron microscopy, the gleba of dried ascomata was scraped, and the pieces of the fertile tissue were fixed onto a double-sided tape. The tissue fragments were gold coated, and the adhered spores were inspected under Hitachi 2360N scanning electron microscope (SEM).
Linear discriminant analysis
A linear discriminant analysis (LDA) was performed on eleven morphological variables of 19 Tuber regianum aff. materials. Average values per sample (herbarium material) were used for each variable. The spore volume (μm3), (the spore ornamentation) mesh diameter (μm), the mesh height (μm) values and the ratio of 1–8 spored asci given as percentage (R1, R2, R3, R4, R5, R6, R7, R8) were included in the analysis after data normalisation. In terms of the spore volume, the mesh height and the mesh diameter, the spores of 8-spored asci were included. The analysis and the visualisation by a plot were done using the MASS, klaR and ggord packages of the R 4.3.2. program. The aim of the LDA was to verify the separability of the species among the T. regianum aff. samples on the basis of morphological characters.
DNA extraction, polymerase chain reaction and sequencing
Total genomic DNA was isolated from dried or fresh ascomata of the Tuber specimens using the DNeasy Plant Mini Kit (Qiagen, Courtaboeuf, France) according to the manufacturer’s instructions with some modifications: after adding buffer AP1, the samples were three times alternately frozen using liquid nitrogen and heated to 65 °C with a BIOER Mixing Block MB-102; the applied volumes of elution buffer (AE) were 30 and 50 μl. Four nuclear loci were amplified including two nuclear ribosomal DNA (rDNA) regions and two protein coding regions: (1) the loci consist of the internal transcribed spacers (ITS 1 and ITS2) and the 5.8S rDNA (for simplicity, this region is called ITS in this study), (2) the large subunit (LSU) of the nuclear ribosomal RNA, (3) the second largest subunit of RNA-polymerase II (RPB2) and (4) the variable segment of the protein kinase C (PKC) locus. ITS1F, ITS4, ITS5, ITS6 and ITS7 (White et al. 1990; Gardes and Bruns 1993; Bertini et al. 1999); LROR, LR3 and LR5 (Vilgalys and Hester 1990); and PKC1F, PKC1R (Ambra and Macino 2000) and RPB2TubF/RPB2TubR (Bonito et al. 2013) primers were used to amplify the ITS, LSU, PKC and RPB2 regions, respectively. Thermocycling was carried out under the following conditions: preliminary denaturation at 94 °C (initialization) for 5–10 min, 33–36 cycles of 94 °C (denaturation) for 30 s, annealing at 51 °C (ITS) or 54 °C (LSU) or 58 °C (PKC and RPB2) for 30–60 s, 72 °C (elongation) for 30–120 s and final synthesis 72 °C for 1 min.
In many cases, the PKC region failed to be amplified with the primer pair of Ambra and Macino (2000). To overcome this difficulty, we designed primer pairs PKCTF (5′-ATCGTATCCCGCACAGGTTC-3′) and PKCTR (5′-ACATGACTTTCCCAAAATTACC-3′) for the PKC region of the genus Tuber. We used the Primer designing tool of NCBI (https://www.ncbi.nlm.nih.gov/tools/primer-blast/index.cgi) and then checked with the OligoAnalyzer 3.1 (IDT, Coralville, IA, USA, www.idtdna.com) online tool. The annealing temperature was determined by gradient PCR, at 52, 54.7, 57.1, 63.3 and 66 °C. After optimizing the annealing temperature of PKCTF/PKCTR oligonucleotides, we applied the following thermocycle: 94 °C for 4 min (initialization), 33 cycles of 94 °C for 20 s (denaturation), 58 °C (annealing) for 30 s and 72 °C (elongation) for 50 s, and finally 72 °C (final elongation) for 7 min. The amplified PCR products of PKC region with this primer pair were around 1000 bp long.
Amplified fragments were electrophoresed on 1% agarose gel, then stained with ethidium-bromide. The amplicons were purified by GenElute™ PCR Clean-Up Kit, according to the manufacturer’s instructions. Finally, the purified PCR products were sequenced by BIOMI Ltd. Biotechnology Service Provider (Gödöllő, Hungary) with Sanger sequencing.
Electropherograms were checked and edited using FinchTV 1.4.0 (https://digitalworldbiology.com/finchtv). The resulting sequences were compared to other sequences in GenBank through BLAST search (Altschul et al. 1997), then were submitted to the GenBank database. Sequences generated in this study together with additional ones from GenBank were aligned using MAFFT (Katoh and Toh 2008) on XSEDE (7.402), on the CIPRES Science Gateway (Miller et al. 2010) (http://www.phylo.org/). The aligned DNA sequences were manually corrected by using MEGA 6.06 (Tamura et al. 2013) in cases of misalignment. File formats were converted using the online application ALTER (Glez-Peña et al. 2010).
Phylogenetic analyses
The Regianum clade
We included sequences of T. regianum aff. specimens and of taxa genetically closely related (species of Excavatum, Gennadii and Aestivum clades) in the phylogenetic reconstruction focused on the Regianum clade. The sequences of 25 herbarium specimens examined by us, additional sequences of 17 herbarium materials available in the NCBI database, in total, sequences of 12 taxa (40 ITS, 19 LSU, 11 RPB2 sequences), were incorporated (Table 1). We conducted maximum likelihood (ML) and Bayesian inference (BI) analyses on the concatenated dataset of 2169 characters. The dataset consists of ITS (1–728), LSU (729–1374) and RPB2 (1375–2169) partitions. The PKC locus was eliminated from the multilocus analysis, because just a few PKC sequences of closely related taxa are available on the GenBank database. Nevertheless, we constructed a PKC gene tree for thirteen available samples (Online Resource 1 Fig. S1, Table S3). The matrices of the alignments were deposited in TreeBASE (Piel et al. 2000) under submission 30716. The ML and BI analyses were computed through the CIPRES Science Gateway (Miller et al. 2010). The optimal substitution models for Bayesian analyses were selected by using the jModelTest 2.1.10. v20160303 (Posada 2008; Darriba et al. 2012) with Akaike information criterion (AIC), which estimated TIM3 + I + G, TIM2 + I + G and TrNef + G to be the optimal substitution models for ITS, LSU and RPB2 loci, respectively. Three independent phylogenetic reconstructions performed by MrBayes on XSEDE (3.2.7a), with four chains of 10,000,000 generations each and sampling every 500th generation after discarding the 25% of the posterior samples as a burn-in. The average standard deviation of split frequencies (ASDSF) value was 0.002019. Effective sample sizes (ESS) were checked with Tracer v1.7.1 (Rambaut et al. 2018). Phylogenetic analysis under the ML criterion was conducted with RAxML-HPC2 on XSEDE (8.2.12), using fast bootstrap (BS) with 1000 replicates; applying a GTR substitution model, the final tree was estimated and optimized under the GAMMA model. Choiromyces meandriformis served as an outgroup (Table 1). Phylogenetic trees were visualized with FigTree v1.3.1 (Rambaut 2009). Paired nucleotide divergence values of T. regianum aff. sequences for each locus were calculated with PAUP* (Swofford 2003) setting the P-distance option.
Genus-wise species tree for the ASR analyses
We assembled a four-loci (EF1α, ITS, LSU and RPB2) dataset of Tuber species to perform ASR analyses. We used the sequences of T. regianum aff. and T. rufum aggr. materials generated in this study, in addition to the sequences from GenBank selected based on the literature (Online Resource 1 Table S1). For most species, the ITS region was included, and the EF1α, LSU and RPB2 regions were included if they were available from GenBank or newly generated for this study (the PKC locus was eliminated from the multilocus species tree as well). Finally, in total, 148 species (OTUs currently registered as species) were incorporated, including outgroup sequences and species described since the comprehensive study of Bonito et al. (2013). Information of sequences and specimens were retrieved from GenBank flatfiles using gbk2fas (Göker et al. 2009) program (available on http://www.goeker.org/mg/clustering/). Before concatenation, we performed BI analysis for the four loci individually to ensure there are no conflicts between highly supported nodes. The concatenated datasets were aligned using MAFFT multiple sequence alignment program. We conducted ML analysis on the combined multilocus datasets. The complete dataset contains 3225 characters, EF1α (1–844 characters), ITS (845–1479), LSU (1480–2363) and RPB2 (2364–3225) loci. ML phylogenetic analysis was constructed using RAxML with 1000 bootstrap replicates, applying a GTR + GAMMA substitution model. We included 141 Tuber taxa, one taxon of Choiromyces lineage (Choiromyces meandriformis) and four taxa of Labyrinthomyces lineage (Labyrinthomyces sp., Nothojafnea thaxteri, Reddellomyces sp., Dingleya sp.). Two members of the Geomoriaceae family, Geomorium australianum and Geomorium echinulatum, were included as outgroup. Taxa and clade names were managed and depicted in this work according to previous studies (Bonito et al. 2010b, 2013; Alvarado et al. 2012a; Merényi et al. 2014; Healy et al. 2016a; Kinoshita et al. 2016; Wan et al. 2016, 2017; Guevara-Guerrero et al. 2018; Páez et al. 2018; Leonardi et al. 2019; Polemis et al. 2019). Information related to the sequences of included Tuber species, sister taxa and outgroup are available in Online Resource 1 Table S1.
Ancestral State Reconstruction
We performed ASR analyses using the above-described genus-wise species tree. Morphological data was derived from our measurements (T. regianum aff. and T. rufum aggr. materials) and from the literature (references in Online Resource 1 Table S1). The six morphological characters (two discrete and four continuous) involved in the ASR analyses are shown in Table 2. To meet the assumption of normal distribution, a logarithmic (log10) transformation was performed and applied for the values of average spore length. The ancestral character states were reconstructed, using the phytools (Revell 2012) R package (R Core Team 2014). For discretely valued traits, two methods, the continuous-time Markov chain (MK) model and stochastic character mapping were applied. For stochastic character mapping, Simmap command of phytools was used under the all rates different (ARD) model and simulated 100 times. For continuous characters, the ancestral character states were reconstructed using the fastAnc function of phytools package.
Results
Molecular phylogeny
Multilocus phylogenetic analyses were performed to examine and reassess the clade of Tuber regianum, T. bernardinii and T. magentipunctatum. At least one locus of 25 specimens was sequenced (Table 1, Online Resource 1 Table S3). We could not obtain sequences in five cases including an isotype of T. regianum and the holotype of T. bernardinii, probably due to the poor condition of the materials (old specimens (~ 30 years) or nonoptimal preservation methods (e.g. camphor)).
BLAST search in GenBank resulted in matches with more than 99% sequence similarities for some of our T. magentipunctatum sequences (ITS: MT374044, MT374045, MT374046, MT374047; LSU: MT350483, MT350484), which are sequences of materials originating in France (Perez 2019). No hits over 96% similarity were found for RPB2 or PKC sequences of T. magentipunctatum and for any loci of T. regianum and T. bernardinii.
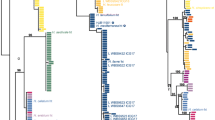
In the multilocus phylogenetic analysis, 565 of the 2169 characters were parsimony informative. The constructed concatenated phylogenetic BI and ML trees resulted in the same tree topology (Fig. 1). The topology and the support values (PP = 1, BS = 100% for all three species) of the three loci (ITS, LSU and RPB2) phylogenetic tree strongly support the existence of three independent lineages (T. regianum, T. magentipunctatum and T. bernardinii) among the T. regianum aff. specimens. The PKC-based phylogeny also supports the split of T. magentipunctatum and T. bernardinii (BS = 100% for both species, Online Resource 1 Fig. S1). P-distance-based nucleotide distance values were considerably low within each group for all loci suggesting intraspecific variation (e.g. < 3% for ITS, according to Nilsson et al. 2008), while the three species show high nucleotide distances from each other based on all but RPB2 locus, implying interspecific relation (Online Resource 1 Table S4). Nevertheless, our main goal was to examine the common phylogenetic unit of the three species to clarify its relevance and position within the genus Tuber. The topology of the phylogenetic trees and the support values of their common branch (PP = 0.98, BS = 85% in the ITS-LSU-RPB2 tree; BS = 100% in the PKC tree) indicates an individual phylogroup (Fig. 1). Therefore, in concordance with Polemis et al. (2019), we refer to this phylogroup as the Regianum clade according to its earliest described species, Tuber regianum.
BI phylogenetic tree of Regianum clade and closely related taxa derived from concatenated internal transcribed spacer (ITS), 28S rRNA (LSU) and second largest subunit of RNA polymerase II (RPB2) regions. Choiromyces meandriformis, a species of Tuberaceae family, was involved as outgroup. The species of Regianum clade are in boldface; holotype specimen is labelled with *, herbarium vouchers in brackets. Bayesian posterior probabilities (PP > 0.95) and maximum likelihood (ML > 70%) bootstrap values are indicated at the nodes of branches, respectively, divided by /. The scale bar represents 0.03 expected nucleotide changes per site
Morphology
Fresh or dried ascomata of 33 T. regianum aff. specimens (Online Resource 1 Table S2) were morphologically identified as T. regianum, T. bernardinii or T. magentipunctatum. All three species can be distinguished from other members of the genus by the dominance of 6–8 spored asci (Online Resource 1 Fig. S2), conspicuously small ascospores (14.9–23.1 × 11.1–16.6 µm on average), with alveolate-reticulate ornamentation (Fig. 2) and relatively small ascomata. The microstructure of the peridium also shares common characteristics between these three species: 50–230-μm-thick external pseudoparenchyma with roundish cells (average size of largest cells 14–25 μm) and a 50–400-μm internal plectenchyma. The external 30–350-μm layer of the peridium of T. regianum and T. magentipunctatum is highly pigmented. We present a morphological key below providing a delimitation of the Regianum clade from other Tuber clades (relying on the currently known three species) as well as the delimitation of the three species from each other.
Morphological characteristics of the species of Regianum clade. Ascomata: a T. regianum (JMV980925-1). b T. magentipunctatum (ZB5307). c T. bernardinii (collection of Günther Schier). Asci: d T. regianum (M22). e T. magentipunctatum (ZB4293). f T. bernardinii (M34). SEM micrographs of spores: g T. regianum (ZB3081). h T. magentipunctatum (ZB4559). i T. bernardinii (M45)
Linear discriminant analysis
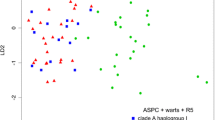
According to the results of the LDA, 100% of the included samples predicted to be correctly identified as Tuber regianum, T. bernardinii or T. magentipunctatum based on the eleven applied morphological characters (the visualisation by plot and the decision rules of the LDA model are in Online Resource 1 Fig. S3).
Taxonomy
Regianum clade
Hypogeous ascomata 0.5–3 cm in diameter, globose or slightly lobed, reddish or blackish-brownish in colour with smooth or minutely papillate peridia. Highest number of spores per ascus is 8, asci most frequently 6–8-spored. Size of ascospores between 14.7–25 × 10–18.4 µm, which is the lowest range for all Tuber clades. Ascospores have a regular alveolate-reticulate ornamentation.
Key to species of Regianum clade
1 Average size of meshes larger than 6 μm, the connection of ridges appear like an awl-like spicule under light microscope, but form complete meshes higher than 2 μm (3.4 μm on average). Ascoma surface smooth with small golden yellow hair patches (septate moniliform cystidia). The dominant number of spores per ascus around 6–8, but the ratio of 8-spored asci is under 30%. Spore size (15)16.6–22.1(25) × (11.7)12.1–15.8(17.7) µm..............................................................................................................Tuber bernardinii
1* Average size of meshes much smaller (max 4.6 μm) as well as the height of them (max 3.3 μm). Ascomata papillate or smooth, but without any hair patches ..........................................................................................2
2 Average spore volume less than 1500 μm3. Average size of meshes 3.0–4.6 μm (total average 3.7 μm) for 8-spored asci, the ridges of meshes well-developed (average height 2.4 μm), thickened on the top, hence appear clubbed under light microscope. Ascoma surface is minutely warted. Dominantly 8 spores per ascus, with the ratio of 8-spored asci above 35%. Spore size (14.7)15.9–17.5(20) × (10)11.1–12.9(13.7) µm……Tuber regianum
2* Average spore volume higher than 1500 μm3. Average size of meshes 2.4–3.1 μm (total average 2.7 μm), the connection of poorly developed ridges forms tiny spicule shorter than 2.3 μm (1.5 μm on average) for 8-spored asci. Ascoma surface smooth (rarely with minutely warted patches), always multicoloured with reddish-purplish patches. Dominant spore number per ascus around 6–8, but the ratio of 8-spored asci is under 35%. Spore size (15)16.4–20.1(24.2) × (11.7)12.6–16.3(18.4) µm…………………………………………Tuber magentipunctatum
Ecology
Regarding the ecological characters of the three species, some similarities and differences are worth mentioning. The ascomata of T. magentipunctatum were found usually at altitudes of 100–300 m (sometimes at 600–700 m), T. regianum and T. bernardinii at 800–1300 m above sea level. Due to the lack of studies on ectomycorrhizae regarding the three Tuber species, their host plant species could only be assumed on the basis of their proximity to the truffle nests. Based on the coenological data, the putative host plant of T. regianum is mostly Fagus sylvatica (in two cases Corylus avellana is also present), which is concordant with the altitude where the fungus has been found. According to the data, T. regianum may have a plant host specificity, while the other two species have a wide range of potential host plants (Quercus, Ostrya, Carpinus, Betulus, Picea, Fagus, Castanea, Tilia, Populus) (Online Resource Table S2). For both T. regianum and T. magentipunctatum, coenosis of only one habitat could be completed: the plant association of a T. regianum habitat is Acereto (pseudoplatani)-Fagetum; in the case of T. magentipunctatum, it is Quercetum petraeae-cerris. Our data on soil pH is 5.43 (5.17–5.81) on average for T. regianum, and 7.08 (6.71–7.67) for T. magentipunctatum. Soil pH values of T. bernardinii are not available in our database.
Genus-wise species tree of the ASR analyses
Between the single-locus phylogenies, there were no conflicts with high support. However, T. magnatum separated from the Aestivum clade in the ITS phylogeny (similarly as in Bonito et al. (2013) for the LSU locus). In the LSU phylogeny, due to the poor resolution of three neighboring lineages (with low branch lengths and Bayesian PP values < 0.88), the Puberulum, the Latisporum and the Maculatum clades, five species were positioned differently between the three clades. Tuber californicum was located in the Maculatum clade (instead of the Puberulum clade), T. sphaerosporum basally to the Latisporum clade (instead of being inside the Puberulum clade), T. pseudosphaerosporum in the Puberulum clade, T. elevatireticulatum basally to all three clades (both instead of being inside the Latisporum clade) and T. dryophilum basally to the Puberulum clade (instead of being inside of it). Nevertheless, these cases were the outcomes of the poor resolution of certain lineages for a given locus. In our concatenated dataset used for the ASR analyses, out of the 3225 characters of the combined EF1α-ITS-LSU-RPB2 matrix, 1199 were parsimony informative. Based on the multilocus ML species tree, the topology of Tuber clades was largely congruent with previous studies (Fig. 3) (Bonito et al. 2010b, 2013; Alvarado et al. 2012b; Merényi et al. 2014; Healy et al. 2016a; Kinoshita et al. 2016; Wan et al. 2016, 2017; Guevara-Guerrero et al. 2018; Páez et al. 2018; Leonardi et al. 2019; Polemis et al. 2019; Fan et al. 2022). The ML bootstrap values of the branches of all thirteen clades were over 90%. T. regianum aff. specimens unambiguously constitute a distinct clade within genus Tuber with high support (BS = 100, Online Resource 1 Fig. S4).
Results of Ancestral State Reconstruction of genus Tuber, by performing the fastAnc function of phytools package in R. Maximum number of spores per ascus (1–8) (left) and average length of spores (16.5–89 μm) on a logarithmic (log10) scale (right) of the currently examined Tuber species and estimated states of ancestral taxa. Schematic representation of a characterful spore of each clade is presented as well. The scale bar (upper rigth corner) for illustrations of spores is 10 µm. AES, Aestivum; EXC, Excavatum; GEN, Gennadii; GIB, Gibbosum; LAT, Latisporum; MAC, Maculatum; MACR, Macrosporum; MEL, Melanosporum; MUL, Multimaculatum; PUB, Puberulum; REG, Regianum; RUF, Rufum; TUR, Turmericum; lab, Labyrinthomyces lineage; cho, Choiromyces sp.; geo, Geomorium spp. Dingleyasp1, Dingleya sp.; Labsp3, Labyrinthomyces sp.; Redsp5, Reddellomyces sp.; Nottha, Nothojafnea thaxteri; Chome, Choiromyces meandriformis; Geoaus, Geomorium australianum; Geoech, Geomorium echinulatum. The species names of spores and the references whose microscopic photos were used to create the schematic spore illustrations: a T. tequilanum (Guevara-Guerrero et al. 2015). b T. panzhihuanense (Deng et al. 2013). c T. arnoldianum (Healy et al. 2016b). d T. oregonense (Bonito et al. 2010b). e T. glabrum (Fan et al. 2014). f T. crassitunicatum (Yan et al. 2018). g T. longispinosum (Kinoshita et al. 2018). h T. multimaculatum (Alvarez et al. 1992). i T. flavidosporum (Kinoshita et al. 2016). j T. sinoaestivum (Zhang et al. 2013). k T. regianum (M22, this study). l T. lacunosum (Alvarado et al. 2012a). m T. depressum (Wan et al. 2017)
Ancestral State Reconstruction
The three currently known species of the Regianum clade possess four morphological traits with hypothetically plesiomorphic character states. To estimate the evolutionary path of the six included morphological traits and to expose evolutionary patterns in the morphological changes of all Tuber clades, ASR analyses were performed. The two methods (MK model and stochastic character mapping) applied for discrete characters provided the same results.
Estimated character states of the Most Recent Common Ancestor of Tuber species
According to our ASR results, the Most Recent Common Ancestor (MRCA) of Tuber species had 5 spores per ascus, a medium–low average spore length (around 34 µm) (Fig. 3), alveolate-reticulate spore ornamentation (Online Resource 1 Fig. S5), medium–low average ascus length (around 115 µm) (Online Resource 1 Fig. S6) and a medium-sized ascoma (up to about 4 cm in diameter) (Online Resource 1 Fig. S7) with verrucose-papillate surface (Online Resource 1 Fig. S8). Some character states, such as 8-spored asci and a low average spore length (below 30 µm), are typical of the sister taxa of genus Tuber, therefore can be considered as plesiomorphic character states in the evolution of Tuberaceae. Nevertheless, our current estimates suggest that the Tuber MRCA probably did not possess these character states.
Morphological character states of the Regianum clade and their evolutionary emergence
Based on our current knowledge of Tuberaceae evolution, the state of four morphological characters (out of the examined six) of the Regianum clade appeared to be plesiomorphic. The hypothesized ancestral states are (i) the relatively low spore length (16.5–20.5 µm), (ii) up to 8 spores per ascus, (iii) the alveolate-reticulate spore ornamentation and (iv) the papillate or smooth surface of the ascomata. Interestingly, two out of the four character states, the low spore length and the 8-spored asci, are estimated as not characteristic of the Tuber MRCA and are uncharacteristic of the Excavatum and Gennadii clades which were basal to the Regianum clade in our analyses. However, these two states are typical for the sister taxa of the genus (Fig. 3). These two character states are estimated as ancestral within Tuberaceae, but they were lost in the MRCA of the Tuber genus based on our analysis above. We also found that these states are estimated to be plesiomorphic to the Regianum clade which suggests the convergent evolution of smaller spore size and 8-spored asci within Tuberaceae. In contrast, alveolate-reticulate spore ornamentation and verrucose-papillate ascoma surface were estimated to be ancient character states of the genus Tuber, but not of the sister lineages of the genus (of which verrucate or spiny spores and smooth ascoma surface are the typical character states). Alveolate-reticulate ornamentation and verrucose-papillate ascoma surface are suggested by our analyses to be characteristic of the Tuber MRCA; therefore, these two features are conserved plesiomorphic character states within the genus Tuber (i.e., in the Regianum clade as well) (Online Resource 1 Fig. S5 and S8). Average length of asci shows a continuous decline from the Tuber MRCA towards the Regianum clade. This feature represents a unidirectional evolutionary change towards the Regianum clade (Online Resource 1 Fig. S6). The maximum size of the ascomata varies in the sister taxa of the genus Tuber (although medium and small sizes are more common), and this feature also presents a decrease from the Tuber MRCA towards the Regianum clade.
In comparison, the sister lineages of the Regianum clade, the Gennadii and the Excavatum clades, are characterised by considerably larger, and the Aestivum clade by slightly larger average spore lengths than the Regianum clade. The maximum number of spores per ascus is low (4 or 5) in the Gennadii and Excavatum clades. The species of the Aestivum clade possess up to 4–8 spores per ascus representing another lineage besides the Regianum clade with even 8-spored asci, but the dominance of 8-spored asci is not typical of the Aestivum clade. The species of the Excavatum, the Gennadii and (with the exception of Tuber panniferum) the Aestivum clade, as well as the Regianum clade, have alveolate-reticulate spore ornamentation. These four basal Tuber clades are uniform in terms of this trait. The characteristic ascoma surface is smooth in the Gennadii clade, primarily verrucose-papillate (or with transitions to smooth or warted type) in the Excavatum clade, and mainly warted (or smooth or papillate) in the Aestivum clade. The distinctive robust flat warts of most species in the Aestivum clade are conspicuously different from the ascoma surface types in the Regianum clade, while the surface types in the Gennadii and Excavatum clades are more similar to the types in the Regianum clade. The average ascus length values of all three sister lineages are higher than that of the species in the Regianum clade. The species of the Aestivum clade have the smallest ascus length values among the three clades (closest to the ascus length of the species in the Regianum clade), and the species of the Gennadii clade have the highest length values. The maximum size of the ascoma in all three sister lineages are also higher than in the Regianum clade. However, the size range in the Aestivum clade is wide, while that of the Excavatum and the Gennadii clades are uniformly slightly higher than the maximum ascoma sizes in the Regianum clade.
Morphological character states of all involved Tuber clades and evolutionary patterns of their emergence
In addition to the Regianum clade, observations regarding all included Tuber clades have also noteworthy aspects. The alveolate-reticulate spore ornamentation is retained in most clades (in basal ones as well as in late branching clades). The spiny ornamentation occurs in species of the Melanosporum and Rufum clades, and emerges at the MRCA of these two clades. In the lineage formed by the Melanosporum and Rufum clades, the spiny ornamentation is retained in some species, while the spino-reticulate and smooth spores evolved nine times and once, respectively. The spiny spores of T. panniferum (Aestivum clade) suggest an independent appearance of this spore ornamentation (Online Resource 1 Fig. S5). The average spore length is the highest in the Multimaculatum, Macrosporum clades and a few species of the Puberulum clade (over 45 µm), which are the results of three recurring transformations. Medium to medium–high spore length (around 35–45 µm) are typical for the Excavatum, Gennadii and Turmericum clades as well as for several species of the Puberulum clade, and a few species of the Melanosporum, Rufum and Maculatum clades. This character state appeared at least seven independent times. Nevertheless, most Tuber species have medium–low average spore length (about 24–34 µm), while low spore length (below 22 µm) is rare. The decrease of spore length appeared convergently at least eight independent times within the genus Tuber, and the extent of this decline in spore length is the greatest in the Regianum clade (Fig. 3). As for the maximum number of spores per ascus, it has a narrow range in the case of some clades (e.g. Regianum 8, Excavatum 4–5, Gibbosum 5–6, Latisporum 3–4), while in others, it is very diverse, therefore not distinctive for the given clade. The emergence of the 8-spored asci in the Regianum-Aestivum lineage and the Melanosporum lineage indicates at least two independent reappearances of this character state, a state more typical for the sister taxa of the genus Tuber. For most Tuber clades, this morphological character is represented by a spore number close to the estimated character state of the Tuber MRCA (a maximum of about 5 spores per ascus). In a few cases, such in Turmericum, Multimaculatum and Macrosporum clades, a lower number (maximum 1–4) of spores per ascus was observed, which are the outcomes of three distinct convergent transitions (Fig. 3). In terms of the average length of the asci, a decrease from the Tuber MRCA was shown towards most of the currently examined Tuber species, and the extent of this decline is the greatest in the Regianum and Aestivum clades. A few exceptions for this are the higher ascus length that reappeared in T. multimaculatum, a few species of the Gennadii, Macrosporum, Maculatum and Puberulum clades (at least six independent cases) (Online Resource 1 Fig. S6). (In contrast to ascus length, ascus width is relatively invariant across the genus (therefore not included in our ASR analyses), around 50–70 µm on average in clades with both long and short asci, with the exception of the Macrosporum clade with a 95 µm on average.) With regard to the maximum size of ascoma, basal Tuber clades are characterised by a more uniform size (Excavatum: medium-small, Gennadii: medium, Regianum: small, Aestivum: large sizes), while multiple convergent reappearance of both smaller and larger size is common in the later branching clades (Online Resource 1 Fig. S7). Regarding the ascoma surface, the possession of verrucose-papillate type occurs in basal and late branching clades as well, retaining the plesiomorphic character state of the genus Tuber in each case. On the other hand, smooth surface (characteristic of the sister taxa of the genus) re-emerged several times convergently within Tuber. The smooth surface appeared at least twenty-one times in the genus, while warted surface is rare, converged in three independent cases (Online Resource 1 Fig. S8).
Discussion
Similarly to all fungal taxa, the taxonomy of the genus Tuber has frequently been modified and new species have been introduced even today (Bonito et al. 2010a, b, 2013; Polemis et al. 2019; Guevara-Guerrero et al. 2015; Kinoshita et al. 2016; Páez et al. 2018; Yan et al. 2018; Leonardi et al. 2019; Fan et al. 2022). T. regianum, T. bernardinii and T. magentipunctatum, the species forming the Regianum clade, are not frequently and not abundantly found species (Bratek et al. 2013), and have a narrow European distribution (Merényi et al. 2017a; Perez 2019). Although this clade’s phylogenetically basal position was presumed (Merényi et al. 2017a; Polemis et al. 2019), it has not been examined in a multilocus phylogenetic framework and no detailed description has been given so far. In addition, the seemingly plesiomorphic morphological character states of the clade also deserve attention from an evolutionary point of view.
According to our multilocus phylogeny, the Regianum clade is highly supported as a distinct phylogroup of the genus Tuber. In concordance with this, the three species can be clearly delimited from the rest of the Tuber clades based on conventional morphological features. The most prominent distinguishing features characteristic to all species in the Regianum clade are the smallest spore size among each Tuber clade (average of all Tuber clades 26.6–37.9 × 20.9–30.6 µm, average of the Regianum clade 14.9–23.1 × 11.1–16.6 µm, average of the Macrosporum clade (the largest sizes) 66.3–82.0 × 46.8–61.6 µm) and the dominance of 6–8-spored asci. In contrast to the morphology, the ecological data do not show clear patterns distinctive to the clade. T. magentipunctatum can be found at lower while the other two species at a higher altitude, and T. regianum grows in acidic soil but T. magentipunctatum was found in mainly neutral soil. Therefore, morphology but not ecology can be used for taxonomic delimitation.
The clear morphological pattern within the Tuber genus also points to the evolutionary relevance of fruiting body and spore traits. Accordingly, the character states of four out of the six examined morphological traits are likely plesiomorphic within the Regianum clade. Based on the ASR analyses, the alveolate-reticulate spore ornamentation and verrucose-papillate ascoma surface are retained ancestral characters, but the other two (up to 8 spores per ascus and small-sized spores) are features which emerged convergently within the genus.
In terms of all examined Tuber clades, our results indicate that the morphological characters analysed for this study evolved different ways. In accordance with the conclusions of Bonito et al. (2013), we found that alveolate-reticulate spore ornamentation is the ancestral character state of genus Tuber, and spiny ornamentation evolved two times in the genus (Melanosporum-Rufum lineage and T. panniferum in the Aestivum clade). Alveolate-reticulate ornamentation is a retained plesiomorphic character not only in basal but in the latest branching Tuber clades as well. In the clades where spiny ornamentation emerged, alveolate-reticulate type had never reappeared. However, spino-reticulate type transformed several times independently from spiny ornamentation. Except for the latter, the type of spore ornamentation could not be seen as a frequently converging character; because it is uniform within lineages and clades the change from one ornamentation type to another occurred rarely based on our analyses (Online Resource 1 Fig. S5). Nevertheless, it is necessary to mention that within these types of spore ornamentations, there is a wide variety of transitions (e.g. mesh size of alveolate-reticulate ornamentation, with or without spines), for example, the ornamentation of T. magentipunctatum could even be considered spinoreticulate on the basis that it also has spicules. With regard to the possible factors in the background of the transformations, presumably some ecological changes could cause the spore morphology to be driven through evolutionary changes for effective spore dispersal (Calhim et al. 2018). Another aspect of spore morphology, the spore length changed convergently within the genus several times (both to lower and higher length), as the low spore length in the Regianum clade is actually a convergently reappeared state (Fig. 3). The maximum number of spores per ascus correlates with spore length, as higher spore length co-occurs with lower number of spores per ascus and vice versa (Fig. 3) (a phenomenon already reported for certain Tuber species in Halász et al. 2005 and Mang et al. 2017). In most cases, ascus length also changed parallel with the spore length and the number of spores per ascus: the higher the length of asci, the higher the spore length and the lower the number of spores per ascus. In contrast to ascus width, which was invariant within the genus, ascus length appears to vary in correlation mainly with spore size, which may be the consequence of the space requirement of many small spores that is lower than that of a few large spores. Nevertheless, the extent of the change of ascus length is not always shown to be as proportionate as the change of spore length and spore number per ascus together. Furthermore, the joint change of the three characters stands only for the genus Tuber (Fig. 3, Online Resource 1 Fig. S6), since Geomorium and Labyrinthomyces lineages are characterised by uniseriate asci. Out of the three characters, presumably spore length may be under selection pressure the most (as spore production is the goal of the sexual stage of the fungal life cycle, and spores must be dispersible and resistant), a shift in the number of spores per ascus and ascus length might primarily be the consequence of the changes in spore size. The evolutionary changes of spore size could be driven by the selection towards beneficial larger size as it enables the accumulation of more nutrients, hence longer persistence (Bässler et al. 2015; Calhim et al. 2018; Aguilar-Trigueros et al. 2023), although larger spore size might be associated with a lower number of spores, which can be a drawback. Although multiple convergent transitions of spore length within the genus Tuber are perhaps the most interesting message of the current study, speaking of the sexual propagules, it is worth also considering the morphology of the structure producing and protecting them. As for the size of the ascoma, basal Tuber clades are characterised by uniformity, while late branching ones are characterised by diversity and with several convergent transitions. Unlike ascoma size, the surface of the ascomata can be both uniform and varied, regardless of whether it is a basal or late branching Tuber clade. The estimated ancestral state of the genus (verrucose-papillate surface) is present in both basal and late branching clades, retaining the plesiomorphic state in all cases. On the other hand, the smooth and warted type appeared convergently several times. Since the peridium serves as a protection for the inner parts of the ascoma (Antony-Babu et al. 2014), environmental changes (particularly that of the soil environment) can presumably be a factor in the evolution of peridial surface. Besides, based on the importance of bacterial species in truffle aroma production (Vahdatzadeh et al. 2015) and the differences in possible microbial intrusions through different types of peridia (Pacioni 1990; Pacioni and Leonardi 2016), the differences in odour and spore dispersal vectors (Trappe et al. 2009) might be in correlation with the emergence of different peridial surfaces. Whatever effects would play a role in the evolution of the ascoma surface, they seem to be independent of the changes of ascoma size. On the basis of the locations and number of observed convergences, the evolutionary changes of the two currently examined features of the ascoma represent different patterns (Online Resource 1 Fig. S7 and S8). Multiple convergent reappearance of morphological character states within the genus Tuber, shown by our ASR results, is an evolutionary phenomenon already observed in other fungi, animals and plants as well (Schmitt et al. 2009; Frenzke et al. 2016; Gorin et al. 2021; Samarakoon et al. 2022; Virágh et al. 2022).
In conclusion, our study suggests that some morphological characters in species of the genus Tuber appear to change together in similar evolutionary patterns presumably due to their functionally and structurally close relationship, as of spore length, maximum number of spores per ascus and ascus length, while other features evolved independently. The Regianum clade, this phylogenetically and morphologically clearly separate Tuber clade, is a prominent example of that even seemingly plesiomorphic character states of a basal clade do not undoubtedly all emerge on the same path. Two of the six studied morphological traits of Regianum clade have retained ancestral character states, two have convergently reappeared plesiomorphic states, while two are characters that have evolved unidirectionally from the ancestors. However, the lack of fossil data on Tuberaceae and the fact that our constantly expanding knowledge of Tuber taxonomy may affect our future view of Tuber phylogeny imply the limits of the present ASR study. In order to refine our current view, a possible topic for future studies could be that how certain environmental factors and their combinations may affect and correlate with the combinations of character states in Tuber clades. This would require a more thorough understanding of the effects of environmental factors on fungal morphology, and the inclusion of ecological variables in ASR studies beside morphology.
Data availability
The matrices of the alignments used for the phylogenetic trees are available on the TreeBASE database under the submission 30716.
References
Agnello C (2011) Raccolte mediterranee di Reddellomyces donkii. Ascomycete.org 3(1):9–13
Aguilar-Trigueros CA, Krah FS, Cornwell WK, Zanne AE, Abrego N, Anderson IC, Andrew CJ, Baldrian P, Bässler C, Bissett A, Chaudhary VB (2023) Symbiotic status alters fungal eco-evolutionary offspring trajectories. Ecology Letters 26:1523–1534. https://doi.org/10.1111/ele.14271
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402. https://doi.org/10.1093/nar/25.17.3389
Alvarado P, Moreno G, Manjón JL (2012a) Comparison between Tuber gennadii and T. oligospermum lineages reveals the existence of the new species T. cistophilum (Tuberaceae, Pezizales). Mycologia 104(4):894–910. https://doi.org/10.3852/11-254
Alvarado P, Moreno G, Manjón JL (2012b) A new Tuber without spore ornamentation. Tuber melosporum comb. nov. Boletín de la Sociedad Micológica de Madrid 36:194
Alvarez IF, Parladé J, Trappe JM (1992) Loculotuber gennadii gen. et comb. nov. and Tuber multimaculatum sp. nov. Mycologia 84(6):926–929. https://doi.org/10.1080/00275514.1992.12026228
Ambra R, Macino G (2000) Cloning and characterization of PKC-homologous genes in the truffle species Tuber borchii and Tuber magnatum. FEMS Microbiol Lett 189(1):45–53. https://doi.org/10.1111/j.1574-6968.2000.tb09204.x
Antony-Babu S, Deveau A, Van Nostrand JD, Zhou J, Le Tacon F, Robin C, Frey-Klett P, Uroz S (2014) Black truffle-associated bacterial communities during the development and maturation of Tuber melanosporum ascocarps and putative functional roles. Environ Microbiol 16(9):2831–2847. https://doi.org/10.1111/1462-2920.12294
Bässler C, Heilmann-Clausen J, Karasch P, Brandl R, Halbwachs H (2015) Ectomycorrhizal fungi have larger fruit bodies than saprotrophic fungi. Fungal Ecol 17:205–212. https://doi.org/10.1016/j.funeco.2014.06.005
Bertini L, Amicucci A, Agostini D, Polidori E, Potenza L, Guidi C, Stocchi V (1999) A new pair of primers designed for amplification of the ITS region in Tuber species. FEMS Microbiol Lett 173:239–245. https://doi.org/10.1111/j.1574-6968.1999.tb13508.x
Bonito GM, Gryganskyi AP, Trappe JM, Vilgalys R (2010a) A global meta-analysis of Tuber ITS rDNA sequences: species diversity, host associations and long-distance dispersal. Mol Ecol 19(22):4994–5008. https://doi.org/10.1111/j.1365-294X.2010.04855.x
Bonito G, Trappe JM, Rawlinson P, Vilgalys R (2010b) Improved resolution of major clades within Tuber and taxonomy of species within the Tuber gibbosum complex. Mycologia 102(5):1042–1057. https://doi.org/10.3852/09-213
Bonito G, Trappe JM, Donovan S, Vilgalys R (2011) The Asian black truffle Tuber indicum can form ectomycorrhizas with North American host plants and complete its life cycle in non-native soils. Fungal Ecol 4(1):83–93. https://doi.org/10.1016/j.funeco.2010.08.003
Bonito G, Smith ME, Nowak M, Healy RA, Guevara G, Cázares E, Kinoshita A, Nouhra RE, Dominguez SL, Tedersoo L, Murat C, Wang Y, Moreno BA, Pfister DH, Nara K, Zambonelli A, Trappe JM, Vilgalys R (2013) Historical biogeography and diversification of truffles in the Tuberaceae and their newly identified southern hemisphere sister lineage. PLoS ONE 8(1):e52765. https://doi.org/10.1371/journal.pone.0052765
Bonito GM, Smith ME (2016) General systematic position of the truffles: evolutionary theories. In: Zambonelli A, Iotti M, Murat C (Eds.) True Truffle (Tuber spp.) in the World: Soil Ecology, Systematics and Biochemistry, vol. 47. Springer, Cham. pp 69–86
Borhidi A (2003) Plant associations of Hungary. Akadémiai Kiadó, Budapest
Bratek Z, Merényi Z, Varga T (2013) Changes of hypogeous funga in the Carpathian-Pannonian region in the past centuries. Acta Mycol 48(1):33–39. https://doi.org/10.5586/am.2013.005
Braun-Blanquet J (1927) Pflanzensoziologie. 1932. Plant Sociology McGraw-Hill, New York
Calhim S, Halme P, Petersen JH, Læssøe T, Bässler C, Heilmann-Clausen J (2018) Fungal spore diversity reflects substrate-specific deposition challenges. Sci Rep 8(1):1–9. https://doi.org/10.1038/s41598-018-23292-8
Crous PW, Wingfield MJ, Burgess TI, Hardy GSJ et al (2017) Fungal Planet description sheets: 558–624. Persoonia 38:380–381. https://doi.org/10.3767/003158517X698941
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772–772. https://doi.org/10.1038/nmeth.2109
Deng XJ, Liu PG, Liu CY, Wang Y (2013) A new white truffle species, Tuber panzhihuanense from China. Mycol Prog 12:557–561. https://doi.org/10.1007/s11557-012-0862-6
Fan L, Feng S, Cao JZ (2014) The phylogenetic position of Tuber glabrum sp. nov. and T. sinomonosporum nom. nov., two Paradoxa-like truffle species from China. Mycol Prog 13:241–246. https://doi.org/10.1007/s11557-013-0908-4
Fan L, Li T, Xu YY, Yan XY (2022) Species diversity, phylogeny, endemism and geography of the truffle genus Tuber in China based on morphological and molecular data. Persoonia 48(1):175–202. https://doi.org/10.3767/persoonia.2022.48.05
Frank JL, Southworth D, Trappe JM (2006) NATS truffle and truffle-like fungi 13: Tuber quercicola and T. whetstonense, new species from Oregon, and T. candidum redescribed. Mycotaxon 95:229–240
Frenzke L, Goetghebeur P, Neinhuis C, Samain MS, Wanke S (2016) Evolution of epiphytism and fruit traits act unevenly on the diversification of the species-rich genus Peperomia (Piperaceae). Front Plant Sci 7:1145. https://doi.org/10.3389/fpls.2016.01145
Gamundí IJ (1971) Algunos discomycetes de Chile. Bol Soc Argent Bot 13:260–289
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Gilkey HM (1939) Tuberales of North America (Vol. 1). Oregon State College
Glez-Peña D, Gómez-Blanco D, Reboiro-Jato M, Fdez-Riverola F, Posada D (2010) ALTER: program-oriented format conversion of DNA and protein alignments. Nucleic Acids Research 38:W14–W18. Web Server issue ISSN: 0305–1048. https://doi.org/10.1093/nar/gkq321
Göker M, García-Blázquez G, Voglmayr H, Tellería MT, Martín MP (2009) Molecular taxonomy of phytopathogenic fungi: a case study in Peronospora. PLoS ONE 4(7):e6319. https://doi.org/10.1371/journal.pone.0006319
Gori L (2003) Tuber Bernardinii. Rivista Di Micologia 46(3):232
Gorin VA, Scherz MD, Korost DV, Poyarkov NA (2021) Consequences of parallel miniaturisation in Microhylinae (Anura, Microhylidae), with the description of a new genus of diminutive South East Asian frogs. Zoosystematics and Evolution 97(1):21–54. https://doi.org/10.3897/zse.97.57968
Guevara-Guerrero G, Bonito G, Cázares-González E, Healy R, Vilgalys R, Trappe J (2015) Novel Tuber spp. (Tuberaceae, Pezizales) in the Puberulum Group from Mexico. Ascomycete.org 7:367–374
Guevara-Guerrero G, Bonito G, Smith ME, Healy R, Grupe AC II, Cázares E, Castellano MA, Trappe JM (2018) Tuber aztecorum sp. nov., a truffle species from Mexico belonging to the Maculatum clade (Tuberaceae, Pezizales). MycoKeys 30:61–72. https://doi.org/10.3897/mycokeys.30.22887
Halász K, Bratek Z, Szegő D, Rudnóy S, Rácz I, Lásztity D, Trappe JM (2005) Tests of species concepts of the small, white, European group of Tuber spp. based on morphology and rDNA ITS sequences with special reference to Tuber rapaeodorum. Mycol Prog 4:281–290. https://doi.org/10.1007/s11557-006-0132-6
Healy R, Bonito GM, Smith ME (2016a) A brief overview of the systematics, taxonomy, and ecology of the Tuber rufum clade. In: Zambonelli A, Iotti M, Murat C (Eds.) True Truffle (Tuber spp.) in the World: Soil Ecology, Systematics and Biochemistry, vol. 47. Springer, Cham. Springer, Cham. pp 125–136
Healy RA, Zurier H, Bonito G, Smith ME, Pfister DH (2016b) Mycorrhizal detection of native and non-native truffles in a historic arboretum and the discovery of a new North American species and: Tuber Arnoldianum sp. nov. Mycorrhiza 26:781–792. https://doi.org/10.1007/s00572-016-0713-4
Iotti M, Amicucci A, Stocchi V, Zambonelli A (2002) Morphological and molecular characterization of mycelia of some Tuber species in pure culture. New Phytol 155(3):499–505. https://doi.org/10.1046/j.1469-8137.2002.00486.x
Justo A, Morgenstern I, Hallen-Adams HE, Hibbett DS (2010) Convergent evolution of sequestrate forms in Amanita under Mediterranean climate conditions. Mycologia 102(3):675–688. https://doi.org/10.3852/09-191
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9(4):286–298
Kinoshita A, Sasaki H, Nara K (2016) Two new truffle species, Tuber japonicum and Tuber flavidosporum spp. nov. found from Japan. Mycoscience 57(5):366–373. https://doi.org/10.1016/j.myc.2016.06.006
Kinoshita A, Nara K, Sasaki H, Feng B, Obase K, Yang ZL, Yamanaka T (2018) Using mating-type loci to improve taxonomy of the Tuber indicum complex, and discovery of a new species, T. longispinosum. PLoS One 13(3):e0193745. https://doi.org/10.1371/journal.pone.0193745
Koske RE, Tessier B (1983) A convenient, permanent slide mounting medium. Newsletter Mycol Soc Amer 34:133–142
Kraisitudomsook N, Healy RA, Pfister DH, Truong C, Nouhra E, Kuhar F, Mujic AB, Trappe JM, Smith ME (2020) Resurrecting the genus Geomorium: systematic study of fungi in the genera Underwoodia and Gymnohydnotrya (Pezizales) with the description of three new South American species. Persoonia 44(1):98–112. https://doi.org/10.3767/persoonia.2020.44.04
Leonardi M, Paz-Conde A, Guevara G, Salvi D, Pacioni G (2019) Two new species of Tuber previously reported as Tuber malacodermum. Mycologia 111(4):676–689. https://doi.org/10.1080/00275514.2019.1603777
Li SH, Zheng LY, Liu CY, Wang Y, Li L, Zhao YC, Zhang XL, Yang M, Xiong HK, Qing Y, Wang L (2014) Two new truffles species, Tuber alboumbilicum and Tuber pseudobrumale from China. Mycol Prog 13:1157–1163. https://doi.org/10.1007/s11557-014-1004-0
Lin CL, Tsai MJ, Fu CH, Chang TT, Li HT, Wong KF (2018) Tuber elevatireticulatum sp. nov., a new species of whitish truffle from Taiwan. Bot Stud 59:1–10. https://doi.org/10.1186/s40529-018-0241-y
Malençon G (1973) Champignons hypogés du nord de l’Afrique—I. Ascomycetes. Persoonia 7(2):261–279
Mang SM, Zotta T, Camele I, Racioppi R, D’Auria M, Rana GL (2017) Morphological, physico-chemical and molecular investigations on Tuber bellonae from Basilicata-Italy. J Anim Plant Sci 27:528–541
Merényi Z, Illyés Z, Völcz G, Bratek Z (2010) A database and its application for the development of truffle cultivation methods. Österreichische Zeitschrift Für Pilzkunde 19:239–244
Merényi Z, Varga T, Geml J, Orczán ÁK, Chevalier G, Bratek Z (2014) Phylogeny and phylogeography of the Tuber brumale aggr. Mycorrhiza 24:101–113. https://doi.org/10.1007/s00572-014-0566-7
Merényi Z, Nagy I, Bratek Z, Stielow JB (2017a) Tuber magentipunctatum Merényi, Nagy, Stielow & Bratek, sp. nov. Fungal Planet description sheets: 558–624. Persoonia 38:380–381. https://doi.org/10.3767/003158517X698941
Merényi Z, Varga T, Hubai AG, Pitlik P, Erős Á, Trappe JM, Bratek Z (2017b) Challenges in the delimitation of morphologically similar species: a case study of Tuber brumale agg. (Ascomycota, Pezizales). Mycol Prog 16:613–624. https://doi.org/10.1007/s11557-017-1296-y
Merényi Z (2014) Molekuláris biológiai és ökológiai vizsgálatok hipogeikus gombákon. Doctoral dissertation. Eötvös Loránd University
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In 2010 gateway computing environments workshop (GCE) (New Orleans, LA) pp 1–8. https://doi.org/10.1109/GCE.2010.5676129
Montecchi A, Sarasini M (2000) Funghi Ipogei d'Europa. Associaazione Micologi ca Bresadola – Fondacione centro Studi Micologici, Trento-Vicenza
Montecchi A, Lazzari G (1987) Un nuovo tartufo di montagna: Tuber regianum sp. nov. Rivista di Micologia 30(1–2):3–11
Moreno G, Alvarado P, Manjón JL (2012) Phylogenetic affiliation of Choiromyces magnusii and C. venosus (Tuberaceae, Ascomycota) from Spain. Mycol Prog 11:463–471. https://doi.org/10.1007/s11557-011-0762-1
Moreno G, Alvarado P, Manjón JL (2013) Hypogeous desert fungi. In: Desert Truffles: Phylogeny, Physiology, Distribution and Domestication Berlin, Heidelberg: Springer Berlin Heidelberg pp 3–20
Murat C, Díez J, Luis P, Delaruelle C, Dupré C, Chevalier G, Bonfante P, Martin F (2004) Polymorphism at the ribosomal DNA ITS and its relation to postglacial re-colonization routes of the Perigord truffle Tuber melanosporum. New Phytol 164(2):401–411. https://doi.org/10.1111/j.1469-8137.2004.01189.x
Nilsson RH, Kristiansson E, Ryberg M, Hallenberg N, Larsson KH (2008) Intraspecific ITS variability in the kingdom Fungi as expressed in the international sequence databases and its implications for molecular species identification. Evol Bioinforma 4:193–201
Pacioni G (1984) Champignons hypogés nouveaux pour l’Afrique du Nord. Bull Trimest Soc Mycol Fr 100(1):111–125
Pacioni G (1990) Scanning electron microscopy of Tuber sporocarps and associated bacteria. Mycol Res 94(8):1086–1089. https://doi.org/10.1016/S0953-7562(09)81338-5
Pacioni G, Leonardi M (2016) Truffle-inhabiting fungi. In: Zambonelli A, Iotti M, Murat C (eds) True Truffle (Tuber spp.) in the World: Soil Ecology, Systematics and Biochemistry, vol 47. Springer, Cham, pp 283–299
Pacioni G, Bologna MA, Laurenzi M (1991) Insect attraction by Tuber: a chemical explanation. Mycol Res 95(12):1359–1363. https://doi.org/10.1016/S0953-7562(09)80385-7
Páez CP, Bonito GM, Guevara-Guerrero G, Castellano MA, Garibay-Orijel R, Trappe JM, Rámirez RP (2018) Description and distribution of Tuber incognitum sp. nov. and Tuber anniae in the Transmexican Volcanic Belt. MycoKeys 41:17–27. https://doi.org/10.3897/mycokeys.41.28130
Perez JB (2019) Tuber magentipunctatum, récolté en Lorraine, une espèce nouvelle pour la France. Ascomycete.org 11(6):210–212. https://doi.org/10.25664/art-0276
Piel WH, Donoghue MJ, Sanderson MJ (2000) TreeBASE: A Database of Phylogenetic Knowledge. To the interoperable “Catalog of Life” with partners Species. 2000:41–47
Polemis E, Konstantinidis G, Fryssouli V, Slavova M, Tsampazis T, Nakkas V, Assyov B, Kaounas V, Zervakis GI (2019) Tuber pulchrosporum sp. nov., a black truffle of the Aestivum clade (Tuberaceae, Pezizales) from the Balkan peninsula. MycoKeys 47:35–51. https://doi.org/10.3897/mycokeys.47.32085
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25(7):1253–1256. https://doi.org/10.1093/molbev/msn083
Quijada L, Matočec N, Kušan I, Tanney JB, Johnston PR, Mešić A, Pfister DH (2022) Apothecial ancestry, evolution, and re-evolution in Thelebolales (Leotiomycetes, Fungi). Biology 11(4):583
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67(5):901–904. https://doi.org/10.1093/sysbio/syy032
Rambaut A (2009) FigTree v1. 3.1: Tree figure drawing tool
Rathnayaka AR, Chethana KT, Phillips AJ, Liu JK, Samarakoon MC, Jones EG, Karunarathna SC, Zhao CL (2023) Re-evaluating Botryosphaeriales: ancestral state reconstructions of selected characters and evolution of nutritional modes. J Fungi 9(2):184. https://doi.org/10.3390/jof9020184
Revell LJ (2012) phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol Evol 2:217–223. https://doi.org/10.1111/j.2041-210X.2011.00169.x
Reynolds HT (2011) Systematics, phylogeography and ecology of Elaphomycetaceae (Doctoral dissertation)
Romero AI, Blumenfield S (2001) Tuber rufum from Rio Negro, Argentina, with notes on Spegazzini’s Tuberales. Mycologist 15:173–175. https://doi.org/10.1016/S0269-915X(01)80013-6
Roux C, Séjalon-Delmas N, Martins M, Parguey-Leduc A, Dargent R, Bécard G (1999) Phylogenetic relationships between European and Chinese truffles based on parsimony and distance analysis of ITS sequences. FEMS Microbiol Lett 180(2):147–155. https://doi.org/10.1111/j.1574-6968.1999.tb08789.x
Royal Botanic Garden E. (1969) Flora of British fungi: colour identification chart. HM Stationery Office. Edinburgh, UK
Samarakoon MC, Hyde KD, Maharachchikumbura SS, Stadler M, Gareth Jones EB, Promputtha I, Suwannarach N, Camporesi E, Bulgakov TS, Liu JK (2022) Taxonomy, phylogeny, molecular dating and ancestral state reconstruction of Xylariomycetidae (Sordariomycetes). Fungal Diversity 112(1):1–88. https://doi.org/10.1007/s13225-021-00495-5
Schmitt I, Del Prado R, Grube M, Lumbsch HT (2009) Repeated evolution of closed fruiting bodies is linked to ascoma development in the largest group of lichenized fungi (Lecanoromycetes, Ascomycota). Mol Phylogenet Evol 52(1):34–44. https://doi.org/10.1016/j.ympev.2009.03.017
Starrett J, Bui A, McGinley R, Hebets EA, Bond JE (2021) Phylogenomic variation at the population-species interface and assessment of gigantism in a model wolf spider genus (Lycosidae, Schizocosa). Insect Syst Divers 5(5):5. https://doi.org/10.1093/isd/ixab016
Sulzbacher MA, Grebenc T, Giachini AJ, Baseia IG, Nouhra ER (2017) Hypogeous sequestrate fungi in South America—how well do we know them? Symbiosis 71:9–17. https://doi.org/10.1007/s13199-016-0461-4
Swofford DL (2003) PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer Associates, Sunderland, Massachusetts
Le Tacon F (2016) Influence of climate on natural distribution of Tuber species and truffle production. In: Zambonelli A, Iotti M, Murat C (Eds.) True Truffle (Tuber spp.) in the World: Soil Ecology, Systematics and Biochemistry, vol. 47. Springer, Cham. pp 153–167
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729. https://doi.org/10.1093/molbev/mst197
Trappe JM, Castellano MA (1991) Keys to the genera of truffles (Ascomycetes). McIlvanea 10:47–65
Trappe JM, Castellano MA, Malajczuk N (1992) Australasian truffle-like Fungi, II.* Labyrinthomyces, Dingleya and Reddellomyces gen. nov. (Ascomycotina). Aust Syst Bot 5(5):597–611
Trappe JM, Molina R, Luoma DL, Cázares E, Pilz, D, Smith JE, Castellano MA, Miller SL, Trappe MJ (2009) Diversity, ecology, and conservation of truffle fungi in forests of the Pacific Northwest. General Technical Report PNW-GTR-772. USDA, Pacific Northwest Research Station, Oregon, pp 126–132
Vahdatzadeh M, Deveau A, Splivallo R (2015) The role of the microbiome of truffles in aroma formation: a meta-analysis approach. Appl Environ Microbiol 81(20):6946–6952. https://doi.org/10.1128/AEM.01098-15
Varga T, Krizsán K, Földi C, Dima B, Sánchez-García M, Sánchez-Ramírez S, Szöllősi GJ, Szarkándi JG, Papp V, Albert L, Andreopoulos W (2019) Megaphylogeny resolves global patterns of mushroom evolution. Nat Ecol Evol 3(4):668–678. https://doi.org/10.1038/s41559-019-0834-1
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172(8):4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
Virágh M, Merényi Z, Csernetics Á, Földi C, Sahu N, Liu XB, Hibbett DS, Nagy LG (2022) Evolutionary morphogenesis of sexual fruiting bodies in basidiomycota: toward a new evo-devo synthesis. Microbiol Mol Biol Rev 86(1):e00019-21. https://doi.org/10.1128/MMBR.00019-21
Wan SP, Wang XH, Zheng Y, Yu FQ (2016) Tuber shidianense and T. calosporum, two new truffle species from southwest China. Mycoscience 57(6):393–399. https://doi.org/10.1016/j.myc.2016.06.007
Wan S, Xu W, Tan N, Wang Y, Zheng Y, Yu F (2017) Three Excavatum species Tuber badium, T. depressum and T verrucosivolvum from Sichuan Province China. Phytotaxa 296(3):228–238. https://doi.org/10.11646/phytotaxa.296.3.2
Wang Y, Moreno G, Riousset LJ, Manjón JL, Riousset G, Fourre G, Di Massimo G, García-Montero LG, Díez J (1998) Tuber pseudoexcavatum sp. nov. A new species from China commercialised in Spain, France and Italy with additional comments on Chinese truffles. Cryptogam Mycol 19(1–2):113–120
Wedén C (2004) Black truffles of Sweden: systematics, population studies, ecology and cultivation of Tuber aestivum syn. T. uncinatum Doctoral dissertation, Acta Universitatis Upsaliensis
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA et al (eds) PCR Protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Yan X, Cao J, Fan L (2018) Four new Tuber species added to the Rufum group from China based on morphological and molecular evidence. Mycologia 110(4):771–779. https://doi.org/10.1080/00275514.2018.1490120
Zambonelli A, Iotti M, Murat C (Eds.) (2016) True Truffle (Tuber spp.) in the World. Soil ecology, systematics and biochemistry, 1st ed. Vol. 47. Springer International Publishing
Zhang BC, Minter DW (1989) Gymnohydnotrya: a new hypogeous ascomycete genus from Australia. Mycol Res 92(2):192–198
Zhang JP, Liu PG, Chen J (2013) Tuber sinoaestivum sp. nov., an edible truffle from southwestern China. Mycotax 122:73–82. https://doi.org/10.5248/122.73
Acknowledgements
In Memoriam of Lamberto Gori, who provided us materials from his mycotheca. We are grateful to Amer Montecchi and Pablo Alvarado for making their herbarial specimens available to us. We want to express our gratitude to Zsófia Gáti for her technical assistance. We also want to thank Sára Pólya for help in designing the PKCTF/PKCTR primer pair. We are grateful to Günther Schier for his photos of Tuber bernardinii from his collection. Many thanks to all associates of Biomi Ltd. for sequencing. We are very grateful to all members of EMSzE and other collectors, who provided the materials used in this work.
Funding
Open access funding provided by Eötvös Loránd University. This work has benefited from the MIKOQUAL project under the Ányos Jedlik Programme and the QUTAOMEL project under the National Technology Programme.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and preparation. Fungal materials, data on the natural habitats, explanations and refinements on the related data were provided by István Nagy, Vasileios Kaounas, Josep Maria Vidal and Aurelia Paz. Morphological measurements were performed by Zsolt Merényi and Torda Varga. SEM microphotographs were made by Károly Bóka and Torda Varga. DNA extraction, primer design and PCR were carried out by Zsolt Merényi and Torda Varga. The phylogenetic trees were constructed by Lilla Bóna. ASR analyses were performed by Zsolt Merényi and Lilla Bóna. Graphical design of figures was performed by Lilla Bóna and Péter Cseh; schematic illustrations of spores and the LDA were prepared by Péter Cseh. The draft of the manuscript was written by Péter Cseh, the contribution of Lilla Bóna to the concept of the first draft version was essential, and all authors commented on previous versions of the manuscript. This work was supervised by Zoltán Bratek and Zsolt Merényi.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Yasmina Marin-Felix
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cseh, P., Merényi, Z., Bóna, L. et al. Taxonomic characterisation of the Regianum clade (genus Tuber) and the trait evolution of spore size among true truffles. Mycol Progress 23, 11 (2024). https://doi.org/10.1007/s11557-024-01949-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-024-01949-1