Abstract
In recent decades, multipartite mutualisms involving microorganisms such as fungi have been discovered in associations traditionally thought of as bipartite. Ant-plant mutualisms were long thought to be bipartite despite fungi being noticed in an epiphytic ant-plant over 100 years ago. We sequenced fungal DNA from the three distinct domatium chambers of the epiphytic ant-plant Myrmecodia beccarii to establish if fungal communities differ by chamber type across five geographic locations spanning 675 km. The three chamber types serve different ant-associated functions including ‘waste’ chambers, where ant workers deposit waste; ‘nursery’ chambers, where the brood is kept; and ‘ventilation’ chambers, that allow air into the domatium. Overall, fungi from the order Chaetothyriales dominated the chambers in terms of the proportion of operational taxonomic units (OTUs; 13.4%) and sequence abundances of OTUs (28% of the total); however a large portion of OTUs (28%) were unidentified at the order level. Notably, the fungal community in the waste chambers differed consistently from the nursery and ventilation chambers across all five locations. We identified 13 fungal OTUs as ‘common’ in the waste chambers that were rare or in very low sequence abundance in the other two chambers. Fungal communities in the nursery and ventilation chambers overlapped more than either did with the waste chambers but were also distinct from each other. Differences in dominance of the common OTUs drove the observed patterns in the fungal communities for each of the chamber types. This suggests a multipartite mutualism involving fungi exists in this ant-plant and that the role of fungi differs among chamber types.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microorganisms such as fungi and bacteria engage in symbioses with other organisms that can have antagonistic (negative) or mutualistic (positive) effects on their hosts. Multipartite mutualisms consist of a prolonged association of more than two partners in which at least two of the interacting organisms receive a net positive benefit (Hussa and Goodrich-Blair 2013). Microorganisms have been found in many mutualisms previously thought of as bipartite including the coral-algae association that also involves bacteria, archaea, and viruses (Rosenberg et al. 2007); the fungus-farming attine ants and antibiotic-producing bacteria that control fungal garden parasites (Currie et al. 1999); and lichen-forming fungi that have complex associations with a green algal photobiont and cyanobacteria (Nelsen et al. 2020). In tropical regions, complex mutualisms have evolved in plants known as myrmecophytes (“ant-plants”). Ant-plant mutualisms were long thought of as bipartite—between the plant and its resident ants—but recent studies suggest that these interactions are more complex and involve hidden microorganisms such as fungi (Mayer et al. 2014; Voglmayr et al. 2011).
Ant-plants provide nesting space to ants in specialised structures called domatia, which are formed from modified plant parts such as stems, thorns, hypocotyls, or leaves (Chomicki and Renner 2015). In some ant-plants, the resident ants obtain food rewards from their host (e.g. extrafloral nectar or food bodies) (Hölldobler and Wilson 1990) and others obtain honeydew from hemipterans they rear in the domatium (Beattie 1985). The resident ants usually defend the plant against enemies such as herbivores (Janzen 1972; Rosumek et al. 2009), and some ant workers feed their host plant by depositing waste on domatium surfaces (Defossez et al. 2011; Gay 1993; Huxley 1978; Rickson 1979; Rico-Gray et al. 1989; Treseder et al. 1995).
Multipartite mutualisms involving fungi, ants, and ant-plants have been identified relatively recently. For example, domatium fungal patches are used as a source of food by ants in three independently evolved and geographically distinct ant-plant (tree) associations (Blatrix et al. 2012; Blatrix et al. 2013; Defossez et al. 2011; Defossez et al. 2009). In one of these ant-plants, the resident ants were observed defecating and depositing detritus on their fungal patch, transporting fragments of the fungus, and chewing hyphae (Defossez et al. 2009). Other ants build traps to capture insects by combining fungi with plant trichomes (hairs) in ant-carton (a combination of vegetative material and soil held together by sugary secretions) on the stems of their host myrmecophyte tree (Dejean et al. 2005). The fungi play a structural role in the trap, receive nutrients from the ants, and facilitate the transfer of nutrients to the plant (Dejean et al. 2005; Leroy et al. 2011; Leroy et al. 2017; Mayer and Voglmayr 2009; Nepel et al. 2014). The dominant fungi isolated from ant-plant systems studied so far are “black yeasts” from the orders Chaetothyriales (class Eurotiomycetes) and Capnodiales (class Dothideomycetes) of phylum Ascomycota (Voglmayr et al. 2011). A recent phylogenetic study found sufficient support for a clade of Chaetothyriales fungi obtained from ant domatia to be recognised as a separate family (Quan et al. 2020).
Epiphytic ant-plants usually grow on trees for support and are typically nutrient-limited, because, like other epiphytes, they do not obtain nutrients or water directly from soil or from their host tree. For this reason, waste deposition by ant workers in the domatium is believed to be particularly important for epiphytic ant-plants (Janzen 1974). In the Australasian region, 47% of ant-plants are epiphytic, whereas most ant-plants are trees or shrubs in Africa (no epiphytes) and the Americas (15% epiphytes) (Chomicki and Renner 2015). Fungi were first noticed in the epiphytic ant-plant Myrmecodia tuberosa Jack (Gentianales: Rubiaceae) (Miehe 1911), but have since been mentioned rarely in the literature (Huxley 1978; Bailey 1920; Janzen 1974) or dismissed as opportunistic (Bailey 1920; Miehe 1911).
Our study investigated fungi in the epiphytic ant-plant Myrmecodia beccarii Hook.f. (Gentianales: Rubiaceae), endemic to northern Queensland, Australia. Myrmecodia beccarii is listed as vulnerable under the Environment Protection and Biodiversity Conservation Act 1999 (Commonwealth of Australia) and the Nature Conservation Act 1992 (State of Queensland), with the main threat being habitat loss due to the destruction of forests containing its host trees (Kemp et al. 2007). The domatium of M. beccarii contains a network of multiple chambers commonly occupied by the native ant Philidris cordata Smith F. (Hymenoptera: Formicidae) (Huxley 1982; Volp and Lach 2019). The chambers include smooth-walled (‘nursery’) chambers where P. cordata keeps its brood (eggs, larvae, and pupae), warted (‘waste’) chambers that absorb nutrients from faeces and waste deposited by ant workers (Huxley 1978; Volp 2016), and superficial (‘ventilation’) chambers that allow air-flow into the system (Huxley 1978, 1982; Jebb 1991). While Huxley (1982) noted two fungal taxa in the waste and nursery chambers of M. beccarii, they were not investigated in detail.
In this work, we hypothesised that epiphytic ant-plants have consistent associations with fungi that correspond with chamber type. To test this, we sampled the fungal communities in the three domatium chambers of M. beccarii across five geographic locations to answer the questions: (a) which fungi dominate the domatium chambers of M. beccarii? (b) are fungi unique to each of the three chamber types or shared among the chambers? (c) do fungal communities differ among the three chamber types, and if so, is this consistent across geographic sites? If fungal communities are consistently distinct in each of the three chamber types across the five locations, it would suggest fungi play different roles in the different chambers because the three chamber types serve different ant-associated functions.
Materials and methods
Study sites and sampling
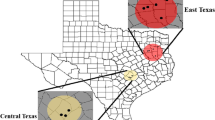
We collected 46 whole Myrmecodia beccarii plants from Melaleuca trees at five locations spanning 675 km in northern Queensland, Australia (Fig. 1) from March 2016 until March 2017. We collected from two sites at Cardwell (10 plants total), one site at Cowley Beach (10 plants), residential Melaleuca trees in Port Douglas (10 plants), one site at Annan River National Park (9 plants), and two sites at Kutini-Payamu National Park (7 plants total). We aimed to analyse the fungal communities from 10 M. beccarii plants from each location, but the plants were rare (and often too high in the canopy to be accessible) in Kutini-Payamu National Park, and one plant collected from Annan River National Park was decomposing inside its domatium at time of dissection and was excluded. The study area represents a large part of the known distribution of M. beccarii from Cardwell in the south (18° 19′ 35.5″ S, 146° 02′ 47.6″ E) to Kutini-Payamu National Park in the north (12° 43′ 19.7″ S, 143° 09′ 38.05″ E) (Fig. 1).
a Map of northern Queensland, Australia, showing the five locations where Myrmecodia beccarii ant-plants were collected during the study. b Known distribution of M. beccarii - red dots indicate where M. beccarii has been found. https://www.openstreetmap.org/copyright/en
We selected M. beccarii ant-plants for collection based on size (circumference of domatium approximately 300 mm around the base), presence of ant workers, and accessibility. All ant-plants collected were from Melaleuca trees at least 20 m distant from any other ant-plant collected. Each M. beccarii was removed from the host tree and placed immediately into a 27-L plastic box that had been lined with Fluon® (Livefoods Unlimited, Tinbeerwah, Qld) on its sides and Tangle-Trap® (Australian Entomological Supplies Pty. Ltd, Murwillumbah, NSW, Australia) in a 25-mm strip around the upper edge of the containers to prevent the resident ant colonies from escaping. The collected M. beccarii ant-plants were kept in their plastic boxes in a greenhouse at James Cook University, Cairns Campus (16° 48′ 58.83′′ S, 145° 41′ 16.73′′ E) until dissection (approximately 1 to 21 days after collection). During this time, we sprayed the roots of the ant-plants with tap water three to four times per week with a garden pressure sprayer to the point of run-off. Each ant colony was provided with two meal worms once per week and approximately 15 mL of 25% sucrose solution divided into two plastic 8 mL vials (plugged with a small ball of cotton wool) twice per week.
Dissection of Myrmecodia beccarii ant-plants and collection of chambers
At time of dissection, we placed each whole M. beccarii ant-plant into an 8.5-L sealed plastic container with five cotton balls soaked in approximately 10 mL total ethyl acetate (Sigma Aldrich, St. Louis, MO, USA) to euthanise the ant colony. The stems, leaves, and roots of the ant-plants were removed from the domatium and discarded. Each domatium was sliced vertically into approximately 10 mm cross-sections with a knife (flame-sterilised using 99.5% ethanol between slices). Three slices were selected for sample collection including one slice from each side/end of the domatium and one slice from the middle of the domatium. For each slice, 4–5 chambers of each chamber type (waste, nursery, and ventilation) were collected using a scalpel to lift the chambers away from the domatium and placed into individual 1.5-mL tubes (total sample weight of 25 mg ± 5 mg for each chamber type for each slice). We identified the different chambers based on their characteristic features: waste chambers being dark brown/black with wart-like surface structures; nursery chambers being yellow coloured with smooth (wart-free) surfaces; and ventilation chambers being brown-coloured with smooth surfaces and a honeycomb type structure near the outer edge of the domatium (Fig. 2). We flame-sterilised the scalpel and forceps during dissections using 99.5% ethanol between each sample collected. A 1.5-mL control tube was left open during each ant-plant dissection to account for potential contaminants during sample processing for DNA extraction. Nine chamber samples per ant-plant (three of each chamber type) were collected during each ant-plant dissection except for Cowley Beach which had only three chamber samples (one of each type of chamber) per ant-plant (being a combination of the chambers collected from three slices as above). The Cowley Beach M. beccarii ant-plants were the first set of ant-plants to be dissected (pilot study).
a Myrmecodia beccarii ant-plant on Melaleuca tree at Cardwell, Queensland. b M. beccarii (i) waste chamber with wart-like structures that absorb nutrients from waste deposited by ant workers; (ii) nursery chamber with smooth surface where the brood of the colony is kept (eggs, larvae, and pupae visible in this photo); and (iii) ventilation chambers, which allow air to flow into the domatium chambers through pores on the surface of the ant-plant. Photos: M. Greenfield
DNA extraction and sequencing
We extracted DNA from the samples to determine the identity of fungi in the different chamber samples, using the Qiagen DNeasy Plant Mini Kit (Qiagen Pty Ltd, Victoria, Australia) following manufacturer’s instructions, except at the final step where we eluted 50 μL of purified DNA instead of 100 μL. We performed polymerase chain reaction (PCR) using the forward primer ITS1Fngs (GGTCATTTAGAGGAAGTAA) (Tedersoo et al. 2015) and reverse primer ITS4ngs (TTCCTSCGCTTATTGATATGC) (Tedersoo et al. 2014) to target the full internal transcribed spacer (ITS) region (ITS1-5.8S-ITS2), the formal barcode for identification of fungi in molecular studies (Schoch et al. 2012). The primers were tagged with 10-11 base pair unique identifiers for multiplexing (Online Resource Table S1). The PCR cocktail consisted of 2 μL DNA extract, 0.5 μL each of the primers (20 μM), 5 μL of 5 × HOT FIREPol® Blend Master Mix (Solis Biodyne, Tartu, Estonia) and 17 μL of double-distilled water. The HOT FIREPol Blend® Master Mix contains HOT FIREPol DNA polymerase (modified Taq polymerase, 99.5% units, error rate 0.011% per base) and a modified proofreading polymerase (0.5% units, estimated 5 × error rate reduction). This enzyme mixture has both 5′-3′ exonuclease activity and 3′-5′ proofreading activity. All samples were amplified in duplicate, and PCRs were carried out in the following thermo-cycling conditions: an initial 15 min at 95 °C, followed by 25 cycles of 95 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min, and a final cycle of 10 min at 72 °C. PCR products for each of the duplicate samples were combined, and the relative quantity of the product was estimated by running gel electrophoresis of 5 μL of DNA sample on 1% agarose gel for approximately 20 min. DNA samples yielding no visible band were reamplified by using 30 or 35 cycles to obtain enough PCR product. Based on visual estimates, the product quantity was categorised into three groups and, accordingly, 5 to 10 μL of PCR products were subjected to library preparation. Negative controls and positive controls (Cantharellus sp. from Africa that does not occur naturally in Australia) were used throughout all procedures (for DNA extraction, PCRs, and sequencing). Pooled amplicons were purified using a FavorPrep™ PCR Purification Kit (FavorGen® Biotech Corp., Vienna, Austria). The amount of DNA in each tube was quantified using Qubit®. Purified PCR products were arranged in four libraries and subjected to SMRTbell library preparation following Pacific Biosciences Amplicon library preparation protocol. The libraries were loaded to 8 SMRT cells using the diffusion method and sequenced on a PacBio Sequel instrument using Sequel Polymerase v2.1, Sequencing chemistry v2.1, and movie time of 600 min following the manufacturer’s recommendations. Circular consensus sequences (CCS) pipeline on SMRT Link (v5.1.0.26412, SMRT Link Analysis Services and GUI v5.1.0.26411) with default settings (minPasses = 3, minAccuracy = 0.9) was used for generating CCS reads.
Bioinformatics
The 8 SMRT cells yielded CCS reads totalling 50461 (library 1), 49839 (library 2), 44336 (library 3), and 52476 (library 4). Bioinformatics analyses of the PacBio sequencing data (for full ITS region) were performed using PipeCraft (v1.0) (Anslan et al. 2017). This analysis platform incorporates required tools for quality filtering, demultiplexing, chimera filtering, clustering, and taxonomy annotation. Quality filtering of the CCS reads were conducted using vsearch (v1.11.1) (Rognes et al. 2016) (fastq_maxee 1, fastq_minlen = 50, fastq_maxns = 0). The filtered data was demultiplexed based on unique identifiers using mothur (v1.36.1) (Schloss et al. 2009) (allowed barcode differences = 1, primer differences = 2). Potential chimeric sequences were detected and removed with vsearch (v1.11.1) using de novo and reference database-based filtering (against UNITE UCHIME release v7.2) (Abarenkov et al. 2010). Multiprimer artefacts (chimeric reads where full primer sequences were found in the middle of the read) were also removed using PipeCraft built-in module. Full ITS region sequences (without flanking genes of 18S and 28S; primer binding sites) were extracted with ITSx (v1.0.9) (Bengtsson-Palme et al. 2013). The full ITS reads were clustered to operational taxonomic units (OTUs) with CD-HIT (v4.6) (Li and Godzik 2006) at a threshold of 97% similarity, as commonly set in fungal molecular ecology (Kõljalg et al. 2013; Taylor et al. 2014). We used BLASTn search for the most abundant sequence of each non-singleton cluster (i.e. OTU) against GenBank and UNITE (v7.2) reference databases for taxonomic assignment of OTUs (e-value = 0.001, word size = 7, reward = 1, penalty = − 1, gap opening cost = 1, gap extension cost = 2).
We further manually filtered the remaining OTUs based on BLASTn values where e-values of <e−50 were used to assign sequences as reliable to the fungal kingdom and e-values >e−20 were considered unknown and removed from the dataset. E-values between e−20 and e−50 were manually checked against the ten best matches for assignment to kingdom Fungi or removal (resulting in another 33 OTUs being detected as chimeric sequences and removed). A further 16 OTUs were detected as chimeric sequences (artefacts of PCR amplification). Two non-fungal OTUs were removed (one insect and one plant). Global singletons (394 OTUs in total, each with only one occurrence in the dataset) were removed to avoid potentially erroneous sequences. The single positive PCR control OTU was removed along with five OTUs (all singletons) that were found only in the positive controls. We also removed two OTU doubletons (only 2 occurrences across the dataset) with low sequence coverage and low sequence similarity. This left a dataset containing 374 OTUs with minimum read abundances of 2 (Greenfield 2020) which we further filtered to remove any OTUs with total read abundances < 10, leaving a dataset with a total of 164 fungal OTUs (Online Resource Table S2). This final step was performed because the focus of this study is on the dominant fungal taxa inside the domatium chambers of M. beccarii and, according to previous studies, excluding rare species makes the community matrix more coherent and less noisy and hence strengthens the statistical power (Tedersoo et al. 2015). We used sequence similarity thresholds of > 97%, > 90%, > 85%, > 80%, > 75%, and > 70% to match OTUs roughly to species, genus, family, order, class, and phylum levels, respectively (Nilsson et al. 2019). Of the 164 OTUs, 70 OTUs (42.7%) matched the taxonomic identity of > 97% to pre-existing fungal ITS sequences in existing databases (GenBank and UNITE). A further 54 OTUs (32.9%) matched at 90–97%, and the remaining 40 OTUs (24.4%) matched to closest taxa at < 90% sequence similarity.
We sequenced 371 samples including 335 fungal DNA amplicon samples collected from the domatium chambers of M. beccarii and 36 laboratory controls (4 positive controls and 4 negative PCR controls, and 28 dissection/extraction controls). The 335 chamber samples were comprised of 116 nursery chambers (27 from Annan River, 26 from Cardwell, 13 from Cowley Beach, 20 from Kutini-Payamu National Park, and 30 from Port Douglas), 97 ventilation chambers (26 from Annan River, 15 from Cardwell, 9 from Cowley Beach, 17 from Kutini-Payamu, and 30 from Port Douglas), and 119 waste chambers (27 from Annan River, 11 from Cowley Beach, 30 from Cardwell, 21 from Kutini-Payamu, and 30 from Port Douglas). Three chamber samples failed to amplify fungi, so they were excluded from the dataset. We pooled the remaining 332 chamber samples to reduce the multiple number of chambers per plant to 3 samples per plant—resulting in there being one nursery chamber sample, one ventilation chamber sample, and one waste chamber sample for each plant collected. This was necessary because the interconnectedness of the chambers in this ant-plant mean multiple samples (for each chamber type) per plant are not independent. This left a total of 135 samples comprising 46 nursery chambers, 44 ventilation chambers, and 45 waste chambers from the 46 ant-plants (Greenfield 2020). Twenty-seven of the 28 dissection/extraction controls contained no contamination and were removed. One dissection control tube (open in the lab during the dissection of the plant AN03 from Annan River National Park) contained a single occurrence of OTU1029 (Tremellomycetes sp.) so this OTU was removed from the AN03 chamber samples. OTU1029 was not removed from any other plant chamber samples because it had not contaminated any other dissection or extraction controls. The positive and negative controls for the PCRs (8 in total) were also removed.
Statistical analyses
Statistical analyses were conducted in R version 3.6.1 (R Core Team. 2019) using the dataset containing sequence abundance data for the 164 fungal OTUs from 135 samples (Online Resource Table S2). Unique and shared fungal OTUs were investigated by creating a Venn diagram using the R package ‘VennDiagram’ (v1.6.20) (Chen 2018). We used the multivariate abundance analysis package ‘mvabund’ (v4.0.1) (Wang et al. 2012; Wang et al. 2020) to test for significant differences among the fungal OTU communities in the three chambers across the five locations. The manyglm function in mvabund was used to fit a model which included chamber type, location, and an interaction term for chamber type and location, with default arguments including family = “negative binomial”, test = “LR” (likelihood-ratio-test), and resamp = ”pit.trap”. We used the anova function in mvabund to compute an analysis of deviance table for the model with pairwise comparisons among the three chamber types (all locations combined). To identify fungal OTUs that were significantly abundant in the chambers and across locations, we used the “p.uni” argument to calculate univariate test statistics and their p values (adjusted for multiple testing using a step-down resampling procedure). We also ran separate models for each of the three chamber types to test for differences in fungal communities across the five locations for each chamber type using pairwise comparisons.
We used the packages ‘phyloseq’ (v1.28.0) (McMurdie and Holmes 2013), ‘vegan’ (v.2.5.6) (Oksanen et al. 2019) and ‘ggplot2’ (v.3.3.0) (Wickham 2016) to create ordination plots in order to visualise differences in the fungal communities in the three different chambers across the five locations. First, we standardised our OTU matrix with a Hellinger-transformation (to account for varying sampling and sequencing depth) and then performed non-metric multidimensional scaling (NMDS) with Bray-Curtis distance measure on the whole dataset (all chambers), and then each chamber type separately to examine differences across locations.
The packages ‘DESeq2’ (v1.24.0) (Love et al. 2014) and ‘phyloseq’ were used to further investigate differentially abundant fungal OTUs in the different chambers. Abundance OTU data was first loaded into Phyloseq and imported into DESEq2 using the phyloseq_to_deseq2 function. The DESeq2 model included both chamber type and location with significance test set to ”Wald”, fitType set to ”local” and multiple inference correction set to “Benjamini-Hochberg”. Pairwise contrasts on chamber type were then carried out with DESeq2 to identify differentially abundant OTUs. The bar plots were created with the package ‘phyloseq’ and ‘ggplot2’. We used FUNGuild (v1.0) (Nguyen et al. 2016) to assign trophic modes to the significantly abundant fungal OTUs.
We used the dataset containing 374 OTUs (minimum read abundances of 2) (Greenfield 2020) to compare fungal OTU richness among chambers across geographic locations. First, we tested if OTU richness is dependent on sample sequencing depth using the packages ‘car’ (v.3.0.10) (Fox and Weisberg 2019) and ‘stats’ (v3.6.3) (R Core Team. 2020). We found OTU richness was significantly positively correlated to sequencing depth (Online Resource Table S3, Fig. S1), so we calculated the residuals for richness for each sample from the regression to account for the relationship in our subsequent analysis. We used the packages ‘lme4’ (v.1.1.26) (Bates et al. 2015) and ‘lmerTest’ (v.3.1.3) (Kuznetsova et al. 2017) to determine whether fungal OTU richness (residuals) varies with chamber type and geographic location. Fixed effects included location and chamber type and an interaction term for location and chamber type, and we included plant ID as a random effect (i.e. the individual plant from which the chamber samples were collected). We used the emmeans package (v1.5.3) (Lenth 2020) for pairwise comparisons between locations for each of the chamber types.
Results
Dominant fungal orders in the domatium chambers
We detected a total of 42,747 sequences from 164 distinct fungal operational taxonomic units (OTUs) across the 135 samples collected from the domatium chambers of M. beccarii (Online Resource Table S2). Seventy percent of the 164 OTUs were classified into 25 fungal taxonomic orders. Below the order level, 33.5% of the 164 OTUs were classified to a family, 32.3% to genus, and 15.9% were classified to species level using available public databases GenBank and UNITE. Sixteen of the 25 orders were from phylum Ascomycota, eight from phylum Basidiomycota, and one from phylum Mortierellomycota. The orders with the highest number of OTUs were Chaetothyriales, Capnodiales, and Eurotiales, which collectively accounted for 57 of the total 164 OTUs (Fig. 3a). Of the 164 OTUs, 46 were unidentifiable at the order level and 15 of these could not be assigned beyond kingdom Fungi. Chaetothyriales was the dominant order in terms of sequence abundance making up 28% of the total (Fig. 3b).
Dominant fungal taxonomic orders in the chambers of the ant-plant Myrmecodia beccarii. a Relative proportion of fungal operational taxonomic units (OTUs) found in the chambers of M. beccarii and assigned to fungal orders, showing Chaetothyriales, Capnodiales, and Eurotiales are dominant. The numbers in brackets indicate the number of fungal OTUs assigned to each order (total 164 OTUs). b Relative abundances of the 164 fungal OTUs found in the chambers of M. beccarii and assigned to fungal orders, showing Chaetothyriales is the most abundant order
Unique and shared fungal OTUs in the domatium chambers
Of the total 164 OTUs detected, there were 125 OTUs in the waste chambers, 142 OTUs in the nursery chambers, and 138 OTUs in the ventilation chambers. Ninety-four OTUs (57.3% of the 164 OTUs) were shared among the three chamber types (Fig. 4). The read abundances of these 94 shared OTUs made up 88.8% of the total 42747 sequences. The nursery and ventilation chambers shared 28 OTUs which made up 3.9% of the sequence abundances. The nursery and waste chambers shared 16 OTUs, comprising 5.6% of the sequences, and the ventilation and waste chambers shared the fewest OTUs (9 in total) making up only 0.7% of the abundances. The 17 OTUs unique to one of the chamber types collectively accounted for 1.1% of sequences (Fig.4).
Venn diagram of unique and shared fungal operational taxonomic units (OTUs) among the three different chambers of the ant-plant Myrmecodia beccarii. The numbers displayed in plain font represent the number of fungal OTUs shared between and among chambers (overlapping shaded areas) and unique (not overlapping) to each chamber type (total 164 fungal OTUs). The numbers in italics in brackets are the corresponding OTU sequence abundances (totalling 42747). The total number of OTUs associated with each of the chambers is: waste chambers 125 OTUs; nursery chambers 142 OTUs; and ventilation chambers 138 OTUs
Fungal communities among domatium chambers
Our analysis indicated significant differences in the fungal communities among the chamber types (LRT = 2546, p < 0.001). Pairwise comparisons of the chamber types showed that the fungal community in the waste chambers was different from the fungal communities in both the nursery (LRT = 1300, p < 0.001) and ventilation chambers (LRT = 1872, p < 0.001). Fungal communities in the waste chambers formed a cluster which was distinct from the other two chambers (Fig. 5a). Significantly different fungal communities were also found between the nursery and ventilation chambers (LRT = 609, p < 0.001) (Fig. 5a).
Non-metric multidimensional scaling (NMDS) ordinations displaying fungal OTU community compositions in the waste, nursery, and ventilation chambers of Myrmecodia beccarii across five locations. Plot a is for the three chamber types for all locations combined, showing the fungal communities in the waste chambers (grey) are distinct from the fungal communities in the nursery (orange) and ventilation (blue) chambers. Plots b, c, and d are separate NMDS plots for the nursery, ventilation, and waste chambers respectively, showing differences across the five locations surveyed: Annan River National Park (purple), Cardwell (yellow), Cowley Beach (light blue), Kutini-Payamu National Park (red), and Port Douglas (dark grey). Each point on a plot is a sample of a fungal community collected from one of the three chambers from one of the five locations. This ordination plot includes 164 fungal operational taxonomic units (OTUs). A Hellinger transformation was used to account for varying sampling and sequencing depth. Bray Curtis distance was used with k = 3 dimensions
Fungal community differences across the geographic distribution of Myrmecodia beccarii
We analysed each of the three chambers of M. beccarii separately to determine if the fungal OTU communities differed among locations for each of the chambers. The fungal OTU community composition varied with location for the waste chambers (LRT = 939.9, p < 0.001), nursery chambers (LRT = 990.5, p < 0.002), and ventilation chambers (LRT = 1165, p < 0.001) (Fig. 5b–d). Our analysis also indicated an interaction between chamber type and location (LRT = 887, p < 0.001). Pairwise comparisons for the waste chambers differed across locations for all but one of the pairwise comparisons, and for the nursery chambers, the fungal OTU communities differed for three of the pairwise comparisons of locations (Table 1). The fungal OTU community composition in the ventilation chambers differed across all locations (Table 1). Fungal OTU richness was significantly higher in the ventilation chambers at Port Douglas (mean 40.0 ± 3.27 SE) compared to the ventilation chambers at Cardwell (mean 22.3 ± 2.13 SE, p = 0.002) (Online Resource Tables S4, S5 and S6, Fig. S2). All other pairwise comparisons of fungal OTU richness between the five locations for each of the three chamber types were not significant (Online Resource Table S6).
To better understand why there were differences in the fungal community compositions across locations, we identified fungal OTUs individually that were significantly abundant in one or more of the three different chamber types. Our DESeq2 analysis found 41 OTUs to be significantly abundant in one or more chambers and of these, the mvabund analysis identified 22 OTUs significant for chamber type only, 10 OTUs significant for chamber type and location, 6 OTUs significant for location only, and 2 OTUs significant for chamber type, location, and an interaction effect (Table 2).
Dominant fungal taxa
We identified 27 common OTUs (from the dataset of 164 OTUs) by selecting those OTUs that occurred in at least 50% of the samples for a chamber type (Fig. 6, Table 3- values in italic). These 27 OTUs were a subset of the 41 fungal OTUs previously identified as significantly differentially abundant (Table 2) and 10 of these 27 fungal OTUs recorded the highest sequence abundances (Online Resource Fig. S3). Each of the 27 OTUs occurred in more than one chamber type but with significantly different abundances (Fig. 6, Tables 2 and 3). For example, OTU0283 (Eurotiomycetes sp) was found in all waste chamber samples, 34.8% of nursery chambers, and 9.1% of ventilation chambers; but in terms of abundance, the waste chambers contained most (96.4%) of the total abundance for this OTU (Table 3).
a Raw (sequence) abundances and b Relative (sequence) abundances of the most common fungal operational taxonomic units (OTUs) in the three different chambers of Myrmecodia beccarii (27 fungal OTUs in total) across the five locations surveyed. Fungal Taxa are the best match found in available databases (GenBank and UNITE). To be selected as one of the most common 27 OTUs, the OTU had to occur in at least 50% of at least one of the chamber types. Under this definition of ‘most common’, there were 6 OTUs in the nursery chambers, 12 OTUs in the ventilation chambers, and 13 OTUs in the waste chambers. Note that each of the most common OTUs for a chamber type may also occur in one, or both, of the other chamber types.
In the waste chambers, we found 13 common fungal OTUs from the orders Chaetothyriales (4), Eurotiales (1), Saccharomycetales (2), Mycosphaerellales (2), and from the class Eurotiomycetes (1) and Tremellomycetes (3). The percentage abundances of each of these 13 common waste chamber OTUs were more than 90% of the total abundances for each of these OTUs across all chambers (Table 3). The 13 common OTUs were either very low in abundance or absent in the nursery and ventilation chambers (Table 3, Fig. 6). Five of the 13 common fungal OTUs in the waste chambers were assigned to trophic modes by FUNGuild with three assigned as saprotrophs and two as symbiotrophs (Table 3). The nursery and ventilation chambers had four fungal OTUs that were common to both these chambers. These four OTUs were from the orders Chaetothyriales (1), Hysteriales (1), Hypocreales (1), and one OTU classified to class Eurotiomycetes (Table 3). Chaetothyreales OTU0347 was found in 71.7% of nursery chambers and 56.8% of ventilation chambers but the abundance of this OTU in the nursery chambers was 83.3% of the total abundance. Likewise, the Hysteriales OTU0563 was found in just over half of both the nursery and ventilation chambers but with highest abundance in the nursery chambers (72.8%). Despite occurring in at least 50% of both nursery and ventilation chambers, the Eurotiomycetes OTU0438 and the Hypocrealeas OTU0544 (Fusarium sp.) occurred in low abundances in the nursery chambers (< 27%) compared to the ventilation chambers (> 70% of the total abundances) (Table 3). The nursery chambers also contained two common OTUs, both from the order Eurotiales, that were found in > 50% of the nursery chambers with relatively high abundances of 84.5% (OTU0281) and 77.2% (OTU0300). The ventilation chambers contained eight other common fungal OTUs belonging to the orders Exobasidiales (3), Capnodiales (3), and Tremellales (2). These eight fungal OTUs had the highest percentage abundances in the ventilation chambers (Table 3).
The 13 common OTUs in the waste chambers were significantly abundant (for chamber type), and five (OTU0202, OTU0214, OTU0263, OTU0302, and OTU0469) were also significantly abundant for location, (Table 2). Also, two OTUs were significant for location and had an interaction effect with chamber type including OTU0283 and OTU1029. Of the six common OTUs in the nursery chambers and 12 OTUs common to ventilation chambers, two were also significant for location including OTU0347 and OTU0898 (Table 2, Fig. 6).
Discussion
This study is among the first to identify and compare the long overlooked fungal communities in the domatium chambers of an epiphytic ant-plant. We found strong associations between chamber type and fungal community composition across our five surveyed geographic locations. The fungal community in the waste chambers of M. beccarii was consistently distinct from the nursery and ventilation chambers across all five locations surveyed which spanned 675 km. The fungal communities within the nursery and ventilation chambers overlapped more than either did with the waste chambers but were also distinct from each other. The fungal OTUs found in each of the chamber types were generally not unique to each chamber type; however, differences in OTU abundances drove the patterns we found in the fungal communities for each of the chamber types. As with other ant-plant systems studied to-date, Chaetothyrialeas fungi dominated in terms of the numbers and abundances of fungal OTUs in the chambers of M. beccarii.
Dominant fungal orders in the domatium chambers
Fungi from the order Chaetothyriales were dominant in the domatium chambers of M. beccarii in terms of the number of fungal OTUs and the abundances of OTUs. The dominance of Chaetothyriales fungi in M. beccarii is consistent with other studies of non-epiphytic ant-plant domatia in Cameroon, Malaysia and French Guiana (Blatrix et al. 2012; Blatrix et al. 2013; Defossez et al. 2009; Moreno et al. 2019; Nepel et al. 2016; Voglmayr et al. 2011). There were also high numbers of fungal OTUs from the order Capnodiales in the chambers of M. beccarii which, together with Chaetothyriales fungi, have also been found associated with ant-carton (Voglmayr et al. 2011) and ant nests (Schlick-Steiner et al. 2008). The presence of Eurotiales fungi in M. beccarii is not surprising given this order of fungi are ubiquitous in nature and include saprotrophic species as well as animal-associated genera (Chen et al. 2015). Forty-six OTUs were unclassifiable at the order level suggesting there are species of fungi in this ant-plant that have never been sequenced before according to the online databases available for comparison (GenBank and UNITE). Our results suggest that epiphytic ant-plants may provide habitats for multiple novel fungal families and potentially order-level taxa; however, it may be at least partly related to our poor understanding about the fungi inhabiting tree canopies in Australia.
The fungal communities in the domatium chambers
The waste, nursery, and ventilation chambers harboured different fungal communities that varied somewhat across locations. Between chambers, differences in fungal communities were driven primarily by variation in the relative sequence abundances of specific OTUs, rather than by unique differences in the identity or number of fungal OTUs in the different chambers. This is clear from the high number of fungal OTUs (94 of the 164 OTUs) that were shared among the three chambers that also collectively made up most (88.8%) of the total abundances. The high number of shared OTUs is not unexpected given the interconnectedness of the domatium chambers and the movement of ant workers among chambers potentially spreads fungal particles across other chamber types. However, despite many OTUs being found across chambers, some fungal OTUs occurred significantly more often in one chamber or another.
The waste chambers of M. beccarii contained 13 fungal OTUs that were significantly abundant and common across the five locations surveyed. The high abundances of these 13 common waste chamber OTUs (and low abundances in, or absence from, the other chambers) suggest that ant workers are maintaining and/or transporting fungi to the waste chambers (e.g. in faeces or other waste), and/or creating an environment suitable for specific fungi. Maintenance of fungi by ant workers in an ant-plant was first noticed by Miehe (1911) who observed fungal mats in the waste chambers of the epiphytic ant-plant Myrmecodia tuberosa (in Java) that had been cut neatly, and the only possible explanation was that the ant workers were trimming fungal hyphae. We also observed dense brown to black thick mats on most of the waste chamber surfaces of all dissected ant-plants. The waste chambers contain the colony’s waste deposits and represent sources for plant nutrient acquisition (Huxley 1978, 1982). Therefore, it is reasonable to expect that at least some of these fungi are involved in the breakdown and releasing of nutrients from waste. Alternatively, fungi in the waste chambers may be cultivated as food or used for their secondary metabolites such as antimicrobial compounds that could be used by the ant colony as defence compounds against pathogens.
Common fungal OTUs in the waste chambers included four fungi from the order Chaetothyriales. One of these (OTU0202) matched at 99% similarity to a sequence found in the domatium of the ant-plant Keetia hispida (Rubiaceae) in Cameroon (accession number HQ634649) (Voglmayr et al. 2011). Although these two ant-plant species are from the family Rubiaceae, the ant species are from different sub-families (Philidris cordata (Dolichoderinae) in M. beccarii and Crematogaster sp. (Myrmicinae) in K. hispida) and these ant-plant systems have evolved separately on different continents. A Chaetothyriales fungal OTU found in Azteca sp. nests on Cecropia trees in Costa Rica was also isolated from domatia of K. hispida occupied by Crematogaster margaritae in Cameroon (Vasse et al. 2017; Nepel et al. 2016). Our research supports the recent phylogenetic study reporting that some ant-associated Chaetothyriales fungi do not cluster according to the ant species, host ant-plant, or geographic origin (Vasse et al. 2017). The other three Chaetothyriales fungal OTUs found in the waste chambers across the five locations had sequence similarities that allowed identification of two of these OTUs to the family Trichomeriaceae (both as putative symbiotrophs according to FUNGuild) and the other to the order Chaetothyriales. Future research could investigate the chambers of other epiphytic ant-plants in the Australasian region to determine if these Chaetothyriales fungi are widespread in other epiphytic ant-plant systems, or whether they show any host plant and/or ant specificity.
Other (non-Chaetothyriales) fungi were also common and abundant in the waste chambers of M. beccarii and consistently found across the five locations surveyed. Three common OTUs were identified to the class Tremellomycetes which contains mostly yeasts that are mycoparasites or animal pathogens (Weiss et al. 2014). Their role (if any) in the waste chambers is yet to be determined, but it is possible that these yeasts act as mycoparasites on the mycelium of Chaetothyriales species. It has been suggested that the occurrence of fungi from orders such as Eurotiales, Hypocreales, Pleosporales, and Saccharomycetales are most likely contaminants (Vasse et al. 2017), opportunistic, or non-symbiotic competitors in ant-plant domatia (Blatrix et al. 2013). However, we found these non-Chaetothyriales fungal OTUs in more than 50% of the waste chambers (some with high abundance) but with very little occurrence in the other chamber types, suggesting a yet-to-be-established functional role.
The differences in the fungal communities between nursery and ventilation chambers were not so pronounced compared with the waste chambers; however, the abundances of most of the common fungal OTUs differed between the former two chamber types. The ventilation chambers lead into the nursery chambers in the lower/middle part of the domatium of M. beccarii, whereas the waste chambers tend to be concentrated more towards the upper/middle portion of the domatium beneath the stem. This might explain why the nursery and ventilation chambers shared some fungal taxa that are relatively uncommon (or absent) in the waste chambers. Also, we often found brood in both the nursery and ventilation chambers, but rarely in the waste chambers (and then only pupae) and we observed ant workers moving brood between the nursery and ventilation chambers. This may be in response to temperature/humidity changes in this ant-plant, and the movement of brood may further explain why there was overlap between these two chamber types as some fungi may be associated with the brood.
Three fungi from the order Chaetothyriales were found in high abundances in the nursery chambers. Chaetothyriales OTU0347 was common at four of our locations and had a 97% match to a sequence isolated from domatia of the ant-plant Leonardoxa africana letouzeyi (ant species: Aphomomyrmex after) in Cameroon (accession number KC951221) (Blatrix et al. 2013). The nursery chambers also contained two other Chaetothyriales fungi (OTUs 372 and 373) with high abundances but low frequency. These OTUs were matched with > 98% identity similarity to sequences isolated from the ant-plant Keetia hispida (Rubiaceae) in Cameroon and Saraca thaipingensis (Fabaceae) in Malaysia respectively (Voglmayr et al. 2011). All domatium symbiont fungi isolated and sequenced previously are closely related to each other (Nepel et al. 2014), and the four Chaetothyriales fungal OTUs from this study support these findings. However, we also found other Chaetothyriales fungi that have not been recorded in other ant-plants.
The ventilation chambers were dominated by OTUs from the order Exobasidiales and Capnodiales. Fungi from Exobasidiales are known to be plant pathogens and are divided into four groups based on their morphology and the plant host range they parasitize, including plants from Ericanae, Lauraceae, monocots, and palms (Begerow 2002). The Exobasidiales sequences found in this study could only be identified to the order level and have never been recorded before. Capnodiales fungi have been found in ant-carton in Cameroon and Malaysia (Voglmayr et al. 2011), and we often observed ant-carton in the ventilation chambers during this study, which may explain the occurrence of Capnodiales fungi. The greater exposure of ventilation chambers to the outside environment increases the likelihood of harbouring opportunistic fungi such as Fusarium OTU0544 which was abundant in all ventilation chambers.
Consistency in fungal OTU communities across the five locations
The significant geographic variation in abundances of 18 of the fungal OTUs and interaction between chamber and location for two of these OTUs indicate large variation in abundances across the five locations surveyed. The abundances and occurrences of any fungus in the domatium chambers of M. beccarri are likely to be influenced by interactions with other fungi and possibly other microorganisms such as bacteria, and this could vary across locations due to, for example, different micro-climates outside domatia. Different numbers of ant workers in different ant-plants are also likely to alter the abundances of fungi in the domatium chambers they occupy. Seven of the 13 common waste chamber fungal OTUs were significantly more abundant at some locations and this may be due to the ant workers transporting/depositing different types (and amounts) of waste into the waste chambers at different locations. The fungal communities in the ventilation chambers were different across all locations and were the only chamber type to have a significant difference in fungal OTU richness at a location, being higher at Port Douglas compared to Cardwell. It is not surprising that the ventilation chambers had the most variation, given they are the most exposed of all chambers to the outside environment and therefore to a range of different fungi. We also observed ant workers entering domatia via large pores that are sometimes present on domatium, and it is possible ant workers transport fungi from the outside environment into the ventilation chambers that could differ across locations. Only three of the pairwise comparisons of geographic locations for the nursery chambers were significant, suggesting the fungal communities in the nursery chambers are the most stable of the three chamber types. This could be because the brood of the colony are tended by ant workers in these chambers, and it is likely the workers keep these chambers free of unwanted fungi. Despite these differences across locations, patterns in the occurrences and abundances of the common fungal OTUs discussed here were found in the domatium chambers of M. beccarii.
Conclusions
The consistent patterns in fungal communities among ant-plant chambers are extraordinary given their fragmented distribution across a broad range and the inclusion of specimens of M. beccarii from both national parks and suburban populations. The different chambers of this epiphytic ant-plant serve different purposes for the ant colony and the plant. It is in the waste chambers where the three potential players in this mutualism intersect: the ants deposit waste in the waste chambers, the fungal community is distinct in the waste chambers, and the plant absorbs nutrients from the waste chambers. While we have not yet unequivocally determined what role/s fungi play in this ant-plant, we have achieved the first step in determining whether a multipartite mutualism exists by showing that the waste chambers contain a specific fungal community that is constant over a large portion of the distribution of this ant-plant. The role of fungi in this mutualism is likely to include the breakdown of organic waste in the waste chambers. However, fungi are involved in so many different interactions with other organisms, and their role in this ant-plant may include various other functions such as the production of antibiotic compounds that keep the brood in the nursery chambers safe from bacterial or fungal pathogens, or fungi may be used as a source of food for the ant colony. It is also probable that some fungi are parasitic or opportunistic. Whether fungi perform any, or all, of these functions in ant-plants should be the focus of future research. Sampling of fungi in the chambers of other epiphytic ant-plants, as well as their resident ant workers, and the host trees and habitat in which epiphytic ant-plants live, could help explain how widespread and common (or not) fungi are in these ant-plants and in the environment generally. Answering these questions could ultimately unravel whether fungi are important in the evolution, maintenance, and stability of epiphytic ant-plant mutualisms.
Data availability
The sequencing data has been deposited in Sequence Read Archive (SRA) under BioProject PRJNA661078. Online Resource Table S2 contains a list of the 164 fungal operational taxonomic units (OTUs) with minimum read abundance of 10, including taxonomic assignment and (closest match in GenBank and UNITE databases), Accession number (GenBank), Species Hypothesis number (UNITE), e-value, % sequence coverage, % sequence similarity. The full dataset of 374 fungal OTUs (minimum read abundance of 2) has been deposited in the James Cook University Tropical Data Hub and will be made publicly available at time of publication (https://doi.org/10.25903/5f713cd732757) (Greenfield 2020). This 374 OTU dataset includes the representative sequence and taxonomic information for each of the fungal OTUs, and additional data including OTU sequence abundances across each of the samples, and sample data including metadata relating to each sample.
References
Abarenkov K, Nilsson RH, Larsson KH, Alexander IJ, Eberhardt U, Erland S, Hoiland K, Kjoller R, Larsson E, Pennanen T, Sen R, Taylor AFS, Tedersoo L, Ursing BM, Vralstad T, Liimatainen K, Peintner U, Koljalg U (2010) The UNITE database for molecular identification of fungi - recent updates and future perspectives. New Phytol 186(2):281–285. https://doi.org/10.1111/j.1469-8137.2009.03160.x
Anslan S, Bahram M, Hiiesalu I, Tedersoo L (2017) PipeCraft: flexible open-source toolkit for bioinformatics analysis of custom high-throughput amplicon sequencing data. Mol Ecol Resour 17(6):e234–e240. https://doi.org/10.1111/1755-0998.12692
Bailey IW (1920) Some relations between ants and fungi. Ecology 1(3):174–189. https://doi.org/10.2307/1929134
Bates D, Maechler M, Bolker BM, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67(1):1–48. https://doi.org/10.18637/jss.v067.i01
Beattie AJ (1985) The evolutionary ecology of ant-plant mutualisms. Cambridge University Press, USA
Begerow D (2002) The exobasidiales: an evolutionary hypothesis. Mycol Prog 1(2):187–199. https://doi.org/10.1007/s11557-006-0018-7
Bengtsson-Palme J, Ryberg M, Hartmann M, Branco S, Wang Z, Godhe A, De Wit P, Sánchez-García M, Ebersberger I, de Sousa F, Amend A, Jumpponen A, Unterseher M, Kristiansson E, Abarenkov K, Bertrand YJK, Sanli K, Eriksson KM, Vik U, Veldre V, Nilsson RH (2013) Improved software detection and extraction of ITS1 and ITS2 from ribosomal ITS sequences of fungi and other eukaryotes for analysis of environmental sequencing data. Methods Ecol Evol 4(10):914–919. https://doi.org/10.1111/2041-210x.12073
Blatrix R, Djiéto-Lordon C, Mondolot L, La Fisca P, Voglmayr H, McKey D (2012) Plant-ants use symbiotic fungi as a food source: new insight into the nutritional ecology of ant–plant interactions. Proc R Soc B Biol Sci 279(1744):3940–3947. https://doi.org/10.1098/rspb.2012.1403
Blatrix R, Debaud S, Salas-Lopez A, Born C, Benoit L, McKey DB, Attéké C, Djiéto-Lordon C (2013) Repeated evolution of fungal cultivar specificity in independently evolved ant-plant-fungus symbioses. PLoS One 8(7):e68101. https://doi.org/10.1371/journal.pone.0068101
Chen H (2018) VennDiagram: Generate high-resolution Venn and Euler plots. R package version 1.6.20. https://CRAN.R-project.org/package = VennDiagram
Chen K, Miadlikowska J, Katalin M, Arnold AE, U’Ren JM, Gaya E, Gueidan C, Lutzoni F (2015) Phylogenetic analyses of eurotiomycetous endophytes reveal their close affinities to Chaetothyriales, Eurotiales, and a new order – Phaeomoniellales. Mol Phylogenet Evol 85:117–130. https://doi.org/10.1016/j.ympev.2015.01.008
Chomicki G, Renner SS (2015) Phylogenetics and molecular clocks reveal the repeated evolution of ant-plants after the late Miocene in Africa and the early Miocene in Australasia and the Neotropics. New Phytol 207(2):411–424
Commonwealth of Australia. Environment Protection and Conservation Act (1999) Australian Government Department of the Environment and Water Resources. Canberra, Australia
Currie CR, Scott JA, Summerbell RC, Malloch D (1999) Fungus-growing ants use antibiotic-producing bacteria to control garden parasites. Nature (London) 398(6729):701–704. https://doi.org/10.1038/19519
Defossez E, Selosse MA, Dubois MP, Mondolot L, Faccio A, Djieto-Lordon C, McKey D, Blatrix R (2009) Ant-plants and fungi: a new threeway symbiosis. New Phytol 182(4):942–949
Defossez E, Djieto-Lordon C, McKey D, Selosse MA, Blatrix R (2011) Plant-ants feed their host plant, but above all a fungal symbiont to recycle nitrogen. Proc R Soc B-Biol Sci 278(1710):1419–1426. https://doi.org/10.1098/rspb.2010.1884
Dejean A, Solano PJ, Ayroles J, Corbara B, Orivel J (2005) Insect behaviour: arboreal ants build traps to capture prey. Nature 434(7036):973–973. https://doi.org/10.1038/434973a
Fox J, Weisberg S (2019) An R companion to applied regression, Third edn. Sage, Thousand Oaks, California, USA https://socialsciences.mcmaster.ca/jfox/Books/Companion/
Gay H (1993) Animal-fed plants: an investigation into the uptake of ant-derived nutrients by the far-eastern epiphytic fern Lecanopteris Reinw.(Polypodiaceae). Biol J Linn Soc 50(3):221–233
Greenfield M (2020) Fungal communities collected from the three distinct chambers of Myrmecodia beccarii from five locations in northern Queensland. James Cook University. (dataset). doi.org/10.25903/5f713cd732757
Hölldobler B, Wilson EO (1990) The ants. Harvard University Press, Cambridge, MA
Hussa EA, Goodrich-Blair H (2013) It takes a village: ecological and fitness impacts of multipartite mutualism. Annu Rev Microbiol 67(1):161–178. https://doi.org/10.1146/annurev-micro-092412-155723
Huxley CR (1978) The ant-plants Myrmecodia and Hydnophytum (Rubiaceae), and the relationships between their morphology, ant occupants, physiology and ecology. New Phytol 80(1):231–268. https://doi.org/10.1111/j.1469-8137.1978.tb02285.x
Huxley CR (1982) Ant-epiphytes of Australia. In:Buckley R (ed) Ant-plant interactions in Australia, vol 4. Geobotany. Springer Netherlands, pp 63-73. doi:10.1007/978-94-009-7994-9_7
Janzen DH (1972) Protection of Barteria (Passifloraceae) by Pachysima ants (Pseudomyrmecinae) in a Nigerian rain forest. Ecology 53:885–892
Janzen DH (1974) Epiphytic myrmecophytes in Sarawak: mutualism through the feeding of plants by ants. Biotropica:237–259
Jebb M (1991) Cavity structure and function in the tuberous Rubiaceae. In:Huxley CR, Cutler DF (eds) Ant-plant interactions. Oxford University Press, pp 374-389
Kemp JE, Lovatt RJ, Bahr JC, Kahler CP, Appelman CN (2007) Pre-clearing vegetation of the coastal lowlands of the Wet Tropics Bioregion, North Queensland. Cunninghamia 10(2)
Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M, Bates ST, Bruns TD, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P, Larsson E, Lindahl BD, Lücking R, Martín MP, Matheny PB, Nguyen NH, Niskanen T, Oja J, Peay KG, Peintner U, Peterson M, Põldmaa K, Saag L, Saar I, Schüßler A, Scott JA, Senés C, Smith ME, Suija A, Taylor DL, Telleria MT, Weiss M, Larsson K-H (2013) Towards a unified paradigm for sequence-based identification of fungi. Mol Ecol 22(21):5271–5277. https://doi.org/10.1111/mec.12481
Kuznetsova A, Brockhoff PB, Christensen RHB (2017) lmerTest Package: tests in linear mixed effects models. J Stat Softw 82(13):1–26. https://doi.org/10.18637/jss.v082.i13
Lenth R (2020) emmeans: estimated marginal means, aka least-squares means. R package version 1.4.5. https://CRAN.R-project.org/package = emmeans
Leroy C, Séjalon-Delmas N, Jauneau A, Ruiz-González M-X, Gryta H, Jargeat P, Corbara B, Dejean A, Orivel J (2011) Trophic mediation by a fungus in an ant–plant mutualism. J Ecol 99(2):583–590. https://doi.org/10.1111/j.1365-2745.2010.01763.x
Leroy C, Jauneau A, Martinez Y, Cabin-Flaman A, Gibouin D, Orivel J, Sejalon-Delmas N (2017) Exploring fungus-plant N transfer in a tripartite ant-plant-fungus mutualism. Ann Bot 120(3):417–426. https://doi.org/10.1093/aob/mcx064
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22(13):1658–1659. https://doi.org/10.1093/bioinformatics/btl158
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Mayer VE, Voglmayr H (2009) Mycelial carton galleries of (Formicidae) as a multi-species network. Proceedings of the Royal Society B. Biological sciences 276(1671):3265–3273. https://doi.org/10.1098/rspb.2009.0768
Mayer VE, Frederickson ME, McKey D, Blatrix R (2014) Current issues in the evolutionary ecology of ant–plant symbioses. New Phytol 202(3):749–764. https://doi.org/10.1111/nph.12690
McMurdie PJ, Holmes S (2013) phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. Plos One 8 (4). doi:https://doi.org/10.1371/journal.pone.0061217
Miehe H (1911) Über die javanische Myrmecodia und die Beziehung zu ihren Ameisen.
Moreno LF, Mayer V, Voglmayr H, Blatrix R, Benjamin Stielow J, Teixeira MM, Vicente VA, de Hoog S (2019) Genomic analysis of ant domatia-associated melanised fungi (Chaetothyriales, Ascomycota). Mycol Prog 18(4):541–552. https://doi.org/10.1007/s11557-018-01467-x
Nelsen MP, Lücking R, Boyce CK, Lumbsch HT, Ree RH (2020) The macroevolutionary dynamics of symbiotic and phenotypic diversification in lichens. Proc Natl Acad Sci 117(35):21495–21503. https://doi.org/10.1073/pnas.2001913117
Nepel M, Voglmayr H, Schönenberger J, Mayer VE (2014) High diversity and low specificity of Chaetothyrialean fungi in carton galleries in a neotropical ant–plant association. PLoS One 9(11):e112756. https://doi.org/10.1371/journal.pone.0112756
Nepel M, Voglmayr H, Blatrix R, Longino JT, Fiedler K, Schönenberger J, Mayer VE (2016) Ant-cultivated Chaetothyriales in hollow stems of myrmecophytic Cecropia sp. trees – diversity and patterns. Fungal Ecol 23:131–140. https://doi.org/10.1016/j.funeco.2016.07.007
Nguyen NH, Song ZW, Bates ST, Branco S, Tedersoo L, Menke J, Schilling JS, Kennedy PG (2016) FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Nilsson RH, Anslan S, Bahram M, Wurzbacher C, Baldrian P, Tedersoo L (2019) Mycobiome diversity: high-throughput sequencing and identification of fungi. Nat Rev Microbiol 17(2):95–109. https://doi.org/10.1038/s41579-018-0116-y
Oksanen J, Guillaume Blanchet F, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2019) Vegan: Community Ecology Package. R package version 2:5–6 https://CRAN.R-project.org/package = vegan
Quan Y, Muggia L, Moreno LF, Wang M, Al-Hatmi AMS, da Silva MN, Shi D, Deng S, Ahmed S, Hyde KD, Vicente VA, Kang Y, Stielow JB, de Hoog S (2020) A re-evaluation of the Chaetothyriales using criteria of comparative biology. Fungal Divers 103(1):47–85. https://doi.org/10.1007/s13225-020-00452-8
R Core Team. (2019) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/
R Core Team. (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/
Rickson FR (1979) Absorption of animal tissue breakdown products into a plant stem-the feeding of a plant by ants. Am J Bot:87–90
Rico-Gray V, Barber JT, Thien LB, Ellgaard EG, Toney JJ (1989) An unusual animal-plant interaction: feeding of Schomburgkia tibicinis (Orchidaceae) by Ants. Am J Bot 76(4):603–608
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Rosenberg E, Koren O, Reshef L, Efrony R, Zilber-Rosenberg I (2007) The role of microorganisms in coral health, disease and evolution. Nat Rev Microbiol 5(5):355–362. https://doi.org/10.1038/nrmicro1635
Rosumek FB, Silveira FA, Neves FS, NPdU B, Diniz L, Oki Y, Pezzini F, Fernandes GW, Cornelissen T (2009) Ants on plants: a meta-analysis of the role of ants as plant biotic defenses. Oecologia 160(3):537–549
Schlick-Steiner BC, Steiner FM, Konrad H, Seifert B, Christian E, Moder K, Stauffer C, Crozier RH (2008) Specificity and transmission mosaic of ant nest-wall fungi. Proc Natl Acad Sci 105(3):940–943. https://doi.org/10.1073/pnas.0708320105
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537. https://doi.org/10.1128/AEM.01541-09
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Fungal Barcoding C, Fungal Barcoding Consortium Author L, Bolchacova E, Voigt K, Crous PW, Miller AN, Wingfield MJ, Aime MC, An K-D, Bai F-Y, Barreto RW, Begerow D, Bergeron M-J, Blackwell M, Boekhout T, Bogale M, Boonyuen N, Burgaz AR, Buyck B, Cai L, Cai Q, Cardinali G, Chaverri P, Coppins BJ, Crespo A, Cubas P, Cummings C, Damm U, de Beer ZW, de Hoog GS, Del-Prado R, Dentinger B, Diéguez-Uribeondo J, Divakar PK, Douglas B, Dueñas M, Duong TA, Eberhardt U, Edwards JE, Elshahed MS, Fliegerova K, Furtado M, García MA, Ge Z-W, Griffith GW, Griffiths K, Groenewald JZ, Groenewald M, Grube M, Gryzenhout M, Guo L-D, Hagen F, Hambleton S, Hamelin RC, Hansen K, Harrold P, Heller G, Herrera C, Hirayama K, Hirooka Y, Ho H-M, Hoffmann K, Hofstetter V, Högnabba F, Hollingsworth PM, Hong S-B, Hosaka K, Houbraken J, Hughes K, Huhtinen S, Hyde KD, James T, Johnson EM, Johnson JE, Johnston PR, Jones EBG, Kelly LJ, Kirk PM, Knapp DG, Kõljalg U, Kovács GM, Kurtzman CP, Landvik S, Leavitt SD, Liggenstoffer AS, Liimatainen K, Lombard L, Luangsa-ard JJ, Lumbsch HT, Maganti H, Maharachchikumbura SSN, Martin MP, May TW, McTaggart AR, Methven AS, Meyer W, Moncalvo J-M, Mongkolsamrit S, Nagy LG, Nilsson RH, Niskanen T, Nyilasi I, Okada G, Okane I, Olariaga I, Otte J, Papp T, Park D, Petkovits T, Pino-Bodas R, Quaedvlieg W, Raja HA, Redecker D, Rintoul TL, Ruibal C, Sarmiento-Ramírez JM, Schmitt I, Schüßler A, Shearer C, Sotome K, Stefani FOP, Stenroos S, Stielow B, Stockinger H, Suetrong S, Suh S-O, Sung G-H, Suzuki M, Tanaka K, Tedersoo L, Telleria MT, Tretter E, Untereiner WA, Urbina H, Vágvölgyi C, Vialle A, Vu TD, Walther G, Wang Q-M, Wang Y, Weir BS, Weiß M, White MM, Xu J, Yahr R, Yang ZL, Yurkov A, Zamora J-C, Zhang N, Zhuang W-Y, Schindel D (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci U S A 109(16):6241–6246. https://doi.org/10.1073/pnas.1117018109
State of Queensland. Nature Conservation Act (1992) Department of Environment and Science. Brisbane, Queensland, Australia
Taylor DL, Hollingsworth TN, McFarland JW, Lennon NJ, Nusbaum C, Ruess RW (2014) A first comprehensive census of fungi in soil reveals both hyperdiversity and fine-scale niche partitioning. Ecol Monogr 84(1):3–20. https://doi.org/10.1890/12-1693.1
Tedersoo L, Bahram M, Polme S, Koljalg U, Yorou NS, Wijesundera R, Ruiz LV, Vasco-Palacios AM, Thu PQ, Suija A, Smith ME, Sharp C, Saluveer E, Saitta A, Rosas M, Riit T, Ratkowsky D, Pritsch K, Poldmaa K, Piepenbring M, Phosri C, Peterson M, Parts K, Partel K, Otsing E, Nouhra E, Njouonkou AL, Nilsson RH, Morgado LN, Mayor J, May TW, Majuakim L, Lodge DJ, Lee SS, Larsson KH, Kohout P, Hosaka K, Hiiesalu I, Henkel TW, Harend H, Guo LD, Greslebin A, Grelet G, Geml J, Gates G, Dunstan W, Dunk C, Drenkhan R, Dearnaley J, De Kesel A, Dang T, Chen X, Buegger F, Brearley FQ, Bonito G, Anslan S, Abell S, Abarenkov K (2014) Global diversity and geography of soil fungi. Science 346(6213):1078. https://doi.org/10.1126/science.1256688
Tedersoo L, Anslan S, Bahram M, Polme S, Riit T, Liiv I, Koljalg U, Kisand V, Nilsson RH, Hildebrand F, Bork P, Abarenkov K (2015) Shotgun metagenomes and multiple primer pair-barcode combinations of amplicons reveal biases in metabarcoding analyses of fungi. Mycokeys 10:1–43. https://doi.org/10.3897/mycokeys.10.4852
Treseder KK, Davidson DW, Ehleringer JR (1995) Absorption of ant-provided carbon dioxide and nitrogen by a tropical epiphyte. Nature 375(6527):137
Vasse M, Voglmayr H, Mayer V, Gueidan C, Nepel M, Moreno L, de Hoog S, Selosse M-A, McKey D, Blatrix R (2017) A phylogenetic perspective on the association between ants (Hymenoptera: Formicidae) and black yeasts (Ascomycota: Chaetothyriales). Proc R Soc B Biol Sci 284(1850). https://doi.org/10.1098/rspb.2016.2519
Voglmayr H, Mayer V, Maschwitz U, Moog J, Djieto-Lordon C, Blatrix R (2011) The diversity of ant-associated black yeasts: insights into a newly discovered world of symbiotic interactions. Fungal Biology 115(10):1077–1091. https://doi.org/10.1016/j.funbio.2010.11.006
Volp TM (2016) Interactions between the epiphytic ant-plant Myrmecodia beccarii and its ant inhabitants. Honours Thesis, James Cook University, Cairns, Queensland, Australia
Volp TM, Lach L (2019) An epiphytic ant-plant mutualism structures arboreal ant communities. Environ Entomol. https://doi.org/10.1093/ee/nvz083
Wang Y, Naumann U, Wright ST, Warton DI (2012) mvabund– an R package for model-based analysis of multivariate abundance data. Methods Ecol Evol 3(3):471–474. https://doi.org/10.1111/j.2041-210X.2012.00190.x
Wang Y, Naumann U, Eddelbuettel D, Wilshire J, Warton D (2020) mvabund: statistical methods for analysing multivariate abundance data. R package version 4.1.3. https://CRAN.R-project.org/package = mvabund
Weiss M, Bauer R, Sampaio JP, Oberwinkler F (2014) 12 tremellomycetes and related groups. In:McLaughlin DJ, Spatafora JW (eds) Systematics and Evolution: Part A. Springer Berlin Heidelberg, Berlin, Heidelberg, pp 331-355. doi:https://doi.org/10.1007/978-3-642-55318-9_12
Wickham H (2016) Elegant Graphics for Data Analysis. Springer-Verlag New York, USA
Acknowledgements
The Myrmecodia beccarii ant-plants collected during this project were done so under a Scientific Purposes Permit (No. WITK16204015) granted to M. Greenfield by the Department of Environment and Heritage Protection. Douglas Shire Council granted permission to M. Greenfield to collect M. beccarii ant-plants in Port Douglas. The sequencing service was provided by the Norwegian Sequencing Centre (www.sequencing.uio.no), a national technology platform hosted by the University of Oslo and supported by the “Functional Genomics” and “Infrastructure” programs of the Research Council of Norway and the Southeastern Regional Health Authorities. We thank the traditional owners of the Northern Kuuku Ya’u Kanthanampu Aboriginal Corporation RNTBC Land Trust and the Yuku-Baja-Muliku Land and Sea Ranger Program for their support in collecting ant-plants from the Kutini-Payamu National Park and Annan River National Park, respectively. We thank Rasmus Puusepp (University of Tartu, Estonia) for technical advice and training regarding the multiplexing PCR process, and Dr. Natalie Dillon and Louise Hucks at James Cook University for their technical advice regarding molecular techniques. We also thank Professor Will Edwards for statistical advice. Volunteers who helped during this project included Ian Coldman, Pauline Lenancker, Barry and Jenn Muir, Kaylene Bransgrove, Susan Nuske, Avril Underwood, Greta Kading, and Brad Connor. We also thank the two anonymous reviewers for taking time to provide critical comments that improved the manuscript.
Funding
This study was funded by Wet Tropics Management Authority (Grant Number 913); Holsworth Wildlife Research Endowment; Linnaean Society of London Systematics Research Fund; Wildlife Preservation Society of Queensland; and James Cook University, College of Science and Engineering Higher Degree by Research Enhancement Scheme.
Author information
Authors and Affiliations
Contributions
M.J.G. designed the study, conducted the field and lab work, collected the data, analysed the data, created the figures and tables, and wrote the manuscript. S.A. performed the bioinformatics and edited the manuscript. L.T. provided laboratory space, primers, and reagents, advice on high-throughput sequencing, and edited the manuscript. M.F. provided advice on data analysis and edited the manuscript. B.C.C. provided advice on interpreting the data and edited the manuscript. S.E.A. and L.L. provided advice on the design of the study, interpretation of results, and edited the manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Section Editor: Dominik Begerow
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 201 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Greenfield, M.J., Lach, L., Congdon, B.C. et al. Consistent patterns of fungal communities within ant-plants across a large geographic range strongly suggest a multipartite mutualism. Mycol Progress 20, 681–699 (2021). https://doi.org/10.1007/s11557-021-01690-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-021-01690-z