Abstract
Numerous gatherings of a new species of the genus Gibellula, closely resembling the monotypic, neotropical G. mirabilis were encountered in Thailand. The taxon was cultured successfully although no in vitro sporulation was observed. The new species, Gibellula gamsii, could be distinguished from closely related other Gibellula species on the basis of morphological features and phylogenetic inferences recruiting concatenated sequences of five DNA loci including ITS, LSU, RPB1, RPB2, and EF1-α. The secondary metabolites of G. gamsii, strain BCC47868, were studied concurrently after preparative separation of the crude extract by preparative high-performance liquid chromatography (HPLC). Two new 1,3-disubstituted β-carboline alkaloids, for which we propose the trivial names, gibellamines A (1) and B (2), were isolated. The chemical structures of these compounds were elucidated by interpretation of spectral data, generated by nuclear magnetic resonance spectroscopy (NMR) and mass spectrometry (MS). The alkaloid 1 also exhibited moderate anti-biofilm activity against Staphylococcus aureus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Gibellula Cavara 1894 (Cordycipitaceae) comprises parasites of spiders that produce cylindrical synnemata arising from host cadavers, exclusively attached to the underside of living leaves, conidiophores with slender apices, obovoid vesicles bearing a number of broadly obovoid metulae, and narrowly clavate phialides bearing conidia found (Mains 1950; Samson and Evans 1992). The genus was established to accommodate G. pulchra and revised by Petch (1932) and Mains (1950). Since then, numerous further species have been proposed (Humber and Rombach 1987; Kobayasi and Shimizu 1982; Samson and Evans 1992; Tzean et al. 1998), and the genus now comprises ca. 30 taxa.
The generic concept of Gibellula mainly relies on its morphological features, while the phylogenetic relationships among species remain poorly understood due to lack of molecular data. Attempts to sequence the ITS rDNA barcode of Gibellula species collected in Thailand were made (BIOTEC fungal barcoding project, unpublished), while Johnson et al. (2009) constructed a multi-gene phylogeny of Torrubiella by using SSU, LSU, EF1-α, RPB1, and RPB2 sequences with only few links to Gibellula. Aside from the fact that access to holotype specimens has been limited, no ex-type strains are available for most Gibellula species. In general, only a few attempts to generate DNA sequence data of Gibellula have so far been made, mostly of the ITS region, and especially poorly represented are type materials as only the ex-type culture of G. clavulifera var. alba was sequenced previously (Kepler et al. 2012; Spatafora et al. 2007), which is known to provide insufficient resolution for species discrimination of Sordariomycetes and many other Ascomycota (Helaly et al. 2018; Hongsanan et al. 2018; Wendt et al. 2018).
In the course of surveying invertebrate-pathogenic fungi and searching for biologically active fungal metabolites, a fungus resembling the Latin American Gibellula mirabilis was collected from various localities throughout Thailand. Its salient morphological traits are its clavate brush-like synnemata with golden brown sterile tips. Samson and Evans (1992) failed to culture the holotype specimen of this fungus as no conidia germinated, and the species has not been reported ever since.
Herein, we report the morphology of a new Gibellula species and describe its phylogenetic placement based on concatenated sequences of five loci (ITS, LSU, RPB1, RPB2, and EF1-α). We also isolate and characterize two secondary metabolites from cultures of G. gamsii, which proved to constitute unprecedented natural products.
Materials and methods
Collection of material and isolation of strains
Specimens were collected from rain forests in various localities throughout Thailand. Spider cadavers colonized by fungi attached on the underside of living leaves were carefully removed from leaves and transported to the laboratory for isolation performed immediately on the same day as described by Kuephadungphan et al. (2014). For microscopic examination, lactophenol cotton blue was used for staining. The specimens were deposited in the BIOTEC Bangkok Herbarium (BBH), while the living cultures were deposited in the BIOTEC Culture Collection (BCC) and the Thailand Biological Resource Center (TBRC).
Molecular phylogenetic analyses
Fungal isolates were grown on potato dextrose agar (PDA; Laboratorios CONDA, Madrid, Spain) at 24 °C for 1–4 weeks, depending on the growth rate of individual strains. After incubation, about 60 mg of fungal mycelium was harvested and subsequently transferred to a 1.5mL screw cap reaction tube with 6–10 sterile 1.4mm ceramic beads (Precellys®). The DNA extraction was performed using the EZ-10 Spin Column Genomic DNA Minipreps Kit (BIO BASIC INC., Canada), following the manufacturer’s instructions. The PCR amplification of five nuclear DNA regions including internal transcribed spacer (ITS), the nuclear ribosomal large subunit DNA (LSU), translation elongation factor 1-alpha (EF1-α), and the largest and second largest subunits of RNA polymerase II (RPB1 and RPB2) were done in 25 μL volume consisting of 12.5 μL of 2× JumpStart Taq ReadyMix (20 mM Tris-HCl, pH 8.3, 100 mM KCl, 3 mM MgCl2, 0.002% gelatin, 0.4 mM of each dNTP (dATP, dCTP, dGTP, TTP), stabilizers, 0.1 unit/μL Taq DNA Polymerase, JumpStart Taq antibody, Sigma-Aldrich, Deisenhofen, Germany), 0.5 μL of each primer, 9.5 μL of sterile milli-Q water, and 2 μL of DNA template. Primers ITS1F (Gardes and Bruns 1993) and ITS4 (White et al. 1990) were used for ITS, 983F and 2218R (Rehner and Buckley 2005) for EF1-α, LR5 or LR7 (Vilgalys and Hester 1990) and LROR (Bunyard et al. 1994) for LSU, RPB1-Ac and RPB1-Cr (Murata et al. 2014) for RPB1, and fRPB2-5F and fRPB2-7cR (Liu et al. 1999) for RPB2. The amplification was performed with the programs as described by Luangsa-ard et al. (2005), Sung et al. (2001), Chaverri and Samuels (2003), Luangsuphabool et al. (2016), and Sung et al. (2007b) for ITS, LSU, EF1-α, RPB1, and RPB2, respectively. The PCR products were purified using an EZ-10 Spin Column PCR purification Kit (BIO BASIC INC., Canada) according to the manufacturer’s instructions. The sequencing was conducted in both forward and reverse directions by the in-house sequencing service of the Helmholtz-Centre of Infection Research, Germany, and Macrogen (South Korea).
Each sequence read was checked for ambiguous bases and assembled using BioEdit v.7.2.5 (Hall 1999; Hall et al. 2011). The alignment was conducted using MAFFT 7.017 with G-INS as algorithm and default settings for gap open and gap extension penalties (Katoh and Toh 2008), and manual adjustments were made in BioEdit. The multigene alignment was subsequently created by concatenation of all regions in Geneious® 7.1.9 (http://www.geneious.com, Kearse et al. 2012). The best fitting substitution model for the multigene alignment was determined by using jModeltest 2.1.7 (Darriba et al. 2012; Guindon and Gascuel 2003; Posada 2008). The phylogenetic relationships were inferred using maximum likelihood (ML) with GTR+GAMMA+I in RAxML 7.2.8 (Stamatakis 2006) and the rapid bootstrap analysis algorithm. Relative support for the branches was obtained from bootstrap analysis with 1000 replicates.
Instrumentation for spectral measurements
High-resolution electrospray ionization mass spectrometry (HRESIMS) was conducted, using an Agilent 1200 series HPLC system in combination with an electrospray ionization time-of-flight mass spectrometric (ESI-TOF-MS; Maxis, Bruker, Bremen, Germany) module [column 2.1 × 50 mm, 1.7 μm, C18 Acquity UPLC BEH (Waters, Eschborn, Germany), solvent A: 95% 5 mM ammonium acetate buffer (pH 5.5, adjusted with 1 M acetic acid) with 5% acetonitrile; solvent B: 95% acetonitrile with 5% 5 mM ammonium acetate buffer]. Nuclear magnetic resonance (NMR) spectra were recorded on Bruker Avance III 500 MHz spectrometer with a BBFO (plus) SmartProbe (1H 500 MHz, 13C 125 MHz) and a BrukerAvance III 700 MHz spectrometer with a 5-mm TCI cryoprobe (1H 700 MHz, 13C 175 MHz). Chemical shifts δ were referenced to the solvents: chloroform-d (1H, δ = 7.27 ppm;13C, δ = 77.0 ppm) and MeOH-d4 (1H, δ = 3.31 ppm; 13C, δ = 49.15 ppm). UV spectra were recorded using a Shimadzu (Duisburg, Germany) UV-vis spectrophotometer UV-2450.
Isolation and structure elucidation of the secondary metabolites
Gibellula gamsii strain BCC47868 was grown on yeast-malt-glucose (YMG) medium (Rupcic et al. 2018). Seven mycelial plugs (1 × 1 cm2) from actively growing colonies were transferred into 20 × 500 mL Erlenmeyer flasks containing 200 mL YMG medium and incubated on shaker operating at 120 rpm, 24 °C for 5 weeks. The culture filtrate (broth) obtained after separation from the fungal mycelia was extracted with ethyl acetate (4 L) to give a pale yellowish powder (170 mg). A portion of the crude broth extract was dissolved in methanol and chemically profiled on Agilent 1260 UHPLC Infinity Systems (Santa Clara, CA, USA) with diode array detector and Waters C18 Acquity UPLC BEH column (2.1 × 50 mm, 1.7 μm) using a previously described gradient elution condition (Noumeur et al. 2017) combined with ion trap MS (amaZon speed, Bruker, Bremen, Germany), and HR-ESIMS spectra on a time-of-flight (TOF) MS (Maxis, Bruker, Germany) to optimize chromatographic purification conditions. Isolation of compounds was achieved with Agilent 1100 series HPLC system (Agilent Technologies, Wilmington, DE, USA) with diode array detection. A reverse-phase C18 column (Kromasil 250 × 20 mm, 7 μm; MZ Analysentechnik, Mainz, Germany) was used as stationary phase. The mobile phase was composed of acid-free deionized water (Milli-Q, Millipore, Schwalbach, Germany: solvent A) and acid-free acetonitrile (ACN: solvent B). Separation of compounds from the crude extract (150 mg) was carried out in 2× injection in accord with the following gradients: linear gradient from 30 to 45% solvent B in 20 min, followed by linear gradient to 70% solvent B in 20 min and linear gradient to 100% solvent B in 10 min, and finally, isocratic conditions at 100% for 10 min, with a flow rate of 20 mL/min. UV detection was carried out at 210, 280, and 354 nm. Fractions were collected and pooled according to the observed peaks. Compounds 1 (18.2 mg) and 2 (1.0 mg) were afforded at retention times, (Rt) = 14–15 and 32–33 min, respectively.
Gibellamine A, 1-(3-(hydroxymethyl)-9H-pyrido[3,4-b]indol-1-yl)ethan-1-one (1): yellow powder; UV (MeOH) (log ε) 217 nm (6.14, λmax), 257 (5.67), 284 (5.79), 306 (5.41), 381 (5.36) nm; 1H NMR (500 MHz, MeOH-d4) see Table 2, 13C NMR (125 MHz, CDCl3) see Table 2; HRESIMS m/z 241.0970 [M + H]+ (calcd for C14H13N2O2, 241.0972)
Gibellamine B, (1-acetyl-9H-pyrido[3,4-b]indol-3-yl)methyl acetate (2): yellow syrup; UV (MeOH) (log ε) 217 nm (6.05, λmax), 253 (5.62), 283 (5.71), 307 (5.31), 381 (5.26) nm; 1H NMR (700 MHz, CDCl3) see Table 2, 13C NMR (175 MHz, CDCl3) see Table 2; HRESIMS m/z 283.1115 [M + H]+ (calcd for C16H15N2O3, 283.1109)
Biological activity screening
The minimum inhibitory concentrations (MICs) of alkaloids 1 and 2 against Bacillus subtilis DSM10, Escherichia coli DSM498, Candida tenuis MUCL29892, and Mucor plumbeus MUCL49355 were determined using a broth microdilution method previously described by Kuephadungphan et al. (2017). Their nematicidal activity against Caenorhabditis elegans and anti-biofilm activity against Staphylococcus aureus DSM1104 (ATCC25923) as well as Pseudomonas aeruginosa PA14 were also investigated using the protocols according to Helaly et al. (2017) and Chepkirui et al. (2018). The cytotoxicity assay was performed on mouse fibroblast (L929), HeLa (KB3.1), human breast adenocarcinoma (MCF-7), human prostate cancer (PC-3), squamous carcinoma (A431), human lung carcinoma(A549), and ovarian carcinoma (SKOV-3) cell lines using a 5-day MTT assay and IC50 values were expressed in micrograms per milliliter (Chepkirui et al. 2017).
Results and discussion
Taxonomy
Gibellula gamsii Kuephadungphan, Tasanathai & Luangsa-ard, sp. nov. Fig. 1.
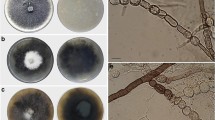
Gibellula gamsii: a, b synnema arising from the host, c part of synnema with golden brown sterile tip, d conidiophores showing conidial head, e part of conidiophore showing rough walls, f conidiophore head bearing conidia, g conidia, and h colonies obverse and reverse on PDA at 25 °C after 40 days. Scale bars: d 50 μm, e, f 20 μm, g 10 μm
Mycobank MB 825141
Etymology: in honor of the late Prof. Walter Gams, for his outstanding contribution to our knowledge of anamorphic hypocrealean fungi
Diagnosis: differs from Gibellula mirabilis, to which it is morphologically similar, in lacking a granulomanus-like conidial state, in mostly forming single synnemata and in having longer, more variable conidiophores with larger conidial heads and smaller conidia. Differs from G. alata and G. unica in synnemata that terminate in a non-sporulating, wing-like structure and by forming numerous synnemata.
Holotype: Thailand, Trang, Khao Banthat Wildlife Sanctuary, on Araneida on unidentified dicot leaf, 18 Sep. 2007, K. Tasanathai; B. Thongnuch (BBH31372, ex-type culture BCC27968)
Habitat: occurring on unidentified, small spiders found on the underside of living leaves
Description: mycelium white to yellowish-white, completely covering spider body. Synnemata yellowish-white to pale yellow, solitary or in groups of three, 5–15 mm long, with clavate, brush-like fertile area, 3–4 × 0.8–1.2 mm; narrowing to a slender, short stipe (ca. 1–2 mm) and terminating in a wing-like, rounded, yellow to golden brown sterile tip that consists of a delicate network of loose, hyaline, anastomosing hyphae (0.5–1 mm long). Conidiophores numerous arising from surface hyphae, scattered, verrucose, hyaline, (10–) 24–57 (− 91) × (2–) 3.5–5.5 (− 6) μm, narrowing to a slender apex and terminating in a swollen vesicle. Conidial head spherical, (31–) 37–42.5 (−48) diam. Vesicles ellipsoidal to globose, (6–) 7–9 (−10) × (5–) 6.5–8.5 (−10), bearing a number of metulae on the upper portion. Metulae ellipsoidal to obovoidal, (3.5–) 6–8.5 (−10) × (3.5–) 4–6 (−8) μm. Phialides mostly oblong-elliptical, (5.5–) 6.5–8.5 (−11) × (2–) 1.5–3 μm. Conidia fusiform to fusiform-elliptical, smooth-walled, hyaline, (3–) 3.5–5 (−5.5) × (1–) 1.5–2.5 (−3) μm. Granulomanus-like synanamorph not observed
Culture characteristics: isolates on PDA slow growing, attaining a diam of 1 cm in 3–4 weeks at 25 °C. Colonies white to yellowish-white; reverse pale yellow to brown at the center, or entirely brown with age. No sporulation observed
Additional specimens examined: Thailand, Trang, Khao Banthat Wildlife Sanctuary, on Araneida on unidentified leaves, 18 Sep. 2007, K. Tasanathai; B. Thongnuch (BBH27524, living culture BCC27970). Kamphaeng Phet, Khlong Lan National Park, on Araneida on unidentified leaves, 6 Nov. 2007, K. Tasanathai; S. Mongkolsamrit; P. Srikitikulchai; B. Thongnuch; R. Ridkaew; A. Khonsanit; W. Chaygate (BBH22756, living culture BCC28798; BBH23098, living culture BCC29228; BBH27573, living culture BCC28797; BBH28376, living culture BCC30449; BBH28378, living culture BCC30396; BBH28379, living culture BCC30397; BBH28372, living culture BCC28801). Chiang Mai, Huai Nam Dang National Park, on Araneida, underside of unidentified living leaf, 5 Jul. 2008, K. Tasanathai; S. Mongkolsamrit; B. Thongnuch; P. Srikitikulchai; A. Khonsanit (BBH27583). Nakhon Ratchasima, Khao Yai National Park, on Araneida, underside of unidentified living leaf, 5 Feb. 2009, K. Tasanathai; S. Mongkolsamrit; J. J. Luangsa-ard; W. Chaygate, P. Puyngain; R. Ridkaew; T. Chohmee (BBH27242); 16 Aug. 2009, K. Tasanathai; S. Mongkolsamrit; T. Chohmee; R. Ridkaew (BBH27250, living culture BCC39922; BBH27251; BBH27252); 12 Sep. 2009, K. Tasanathai; P. Srikitikulchai; S. Mongkolsamrit; T. Chohmee; R. Ridkaew (BBH27258, living culture BCC39024); 13 Sep. 2009, K. Tasanathai; P. Srikitikulchai; S. Mongkolsamrit; T. Chohmee; R. Ridkaew (BBH27259, living culture BCC39038); 6 Oct. 2009, K. Tasanathai; P. Srikitikulchai; S. Mongkolsamrit; T. Chohmee, R. Ridkaew, P. Puyngain; M. Sudhadham (BBH27274, living culure BCC39953); 7 Oct. 2009, K. Tasanathai; P. Srikitikulchai; S. Mongkolsamrit; T. Chohmee; R. Ridkaew; P. Puyngain; M. Sudhadham (BBH27275, living culture BCC39966); 10 Nov. 2009, K. Tasanathai; S. Mongkolsamrit; T. Chohmee; R. Ridkaew; M. Sudhadham; A. Khonsanit (BBH27281, living culture BCC40856); 12 Nov. 2009, K. Tasanathai; S. Mongkolsamrit; T. Chohmee; R. Ridkaew; M. Sudhadham; A. Khonsanit (BBH27286); 7 Jan. 2010, K. Tasanathai; P. Srikitikulchai; S. Mongkolsamrit; T. Chohmee; R. Ridkaew; M. Sudhadham; A. Khonsanit (BBH27351, living culture BCC40909); 3 Mar. 2010, K. Tasanathai; P. Srikitikulchai; S. Mongkolsamrit; T. Chohmee A. Khonsanit (BBH28510, living culture BCC42026); 20 May 2010, A. Khonsanit; R. Somnuk (BBH28560); 31 May 2010, K. Tasanathai, P. Srikitikulchai; S. Mongkolsamrit; T. Chohmee; A. Khonsanit; R. Somnuk; K. Sansatchnon (BBH29333). Songkhla, Ton-Nga-Chang Wildlife Sanctuary, on Araneida, underside of unidentified living leaf, 20 Jun. 2009, W. Kuephadungphan; W. Phetmanee (EPF034, living culture BCC47868)
Notes: Gibellula gamsii shows morphological similarities with G. mirabilis (Samson and Evans 1992). Both produce clavate, brush-like synnemata that terminate into a sterile tip. Most specimens of G. gamsii produced a single synnema per specimen; only in some specimens were the synnemata found in pairs or in groups of three. Samson and Evans (1992) reported synnemata in groups of two for G. mirabilis. Moreover, in G. gamsii, the length of conidiophores varied from (10–) 24–57 (− 91) μm, while for G. mirabilis, they are up to 80 μm long. Conidial heads were larger while conidia were smaller in G. gamsii [(31–) 37–42.5 (−48) μm diam, (3–) 3.5–5 (−5.5) × (1–) 1.5–2.5(−3) μm] compared to those of G. mirabilis [25–40 μm diam, 5–7 × 2–3.5 μm]. Furthermore, the granulomanus-like synanamorph reported for G. mirabilis could not be seen in any of the Thai specimens at any developmental stage. The significance of a granulomanus-like conidial state for species discrimination of Gibellula is unclear and requires study on larger numbers of specimens and strains. It is known, however, that Gibellula and Granulomanus naturally appear either concurrently or independently on the same spider hosts (Samson et al. 1989). Several species of Gibellula has been reported from Asia, such as G. alata from Sri Lanka (Petch 1932) and G. unica from Taiwan (Tzean et al. 1997). Although the conidial sizes of G. gamsii fall into the range of those reported for G. alata (4–9 × 2–4 μm) and G. unica (4–6.8 × 1.6–2.2 μm), G. gamsii can be easily distinguished from both species. G. alata typically produces wing-like structures at the swollen tip of synnema while G. unica forms numerous cylindric synnemata as well as granulomanus-like synanamorph. Some Asian Gibellula species such as G. clavulifera var. major produce long conidia (7.1–12 × 2.4–4 μm) and penicillium-like conidiophores (Tzean et al. 1997).
Gibellula was previously distinguished from Pseudogibellula by its specialized growth requirements as it could not be established in culture (Samson and Evans 1973). Samson and Evans (1992) pointed out that the majority of Gibellula species are very hard to isolate although they previously succeeded in growing G. clavulifera on mealworm agar. Tzean et al. (1997) failed to culture G. clavulifera, G. leiopus, G. pulchra, and G. unica collected in Taiwan from single or mass conidia or mycelia on diverse artificial media. In our study, cultures of G. gamsii were successfully isolated from conidia even on the standard medium PDA. This property may also turn out to be characteristic for the new species.
Molecular phylogeny
Thirty-one fungal taxa were included in the analyses, 10 of which resulted from our own work and 21 datasets were taken from GenBank (Table 1). The calculated MAFFT alignments consisted of 4108 characters, 664, 860, 718, 937, and 869 of which were derived from the ITS, LSU, RPB1, RPB2, and EF1-α datasets, respectively.
The attempt to include the β-tubulin coding gene (TUB) which was generated using standard primers introduced by Liu et al. (1999) (RPB2-5F and RPB2-7cR) in the analyses was made. The concatenated multiple gene dataset (ITS, LSU, RPB1, RPB2, EF1-α, and TUB) was created and the phylogenetic tree was then generated according to the protocol described above. Interestingly, the multigene tree which is provided in the Supplementary Material showed very long branches at some certain nodes as well as uncorrelated topologies compared with those previously described by Chiriví-Salomón et al. (2015), D’Alessandro et al. (2013), and Kepler et al. (2017). Therefore, TUB was excluded from the dataset but it is included in the Supplementary Material.
The multigene tree (Fig. 2) comprised seven different genera belonging to the family Cordycipitaceae including Akanthomyces, Beauveria, Blackwellomyces, Cordyceps, Gibellula, Hevansia, and outgroup Engyodontium. The analyses showed the genera segregated corresponding to the most recent phylogenetic classification of the Cordycipitaceae (Mongkolsamrit et al. 2018). Gibellula has been inferred as a monophyletic group in previous phylogenetic studies (Johnson et al. 2009; Kepler et al. 2017). A Gibellula clade composed of G. clavulifera cf. alba, G. cf. alba, G. leiopus, G. longispora, G. pulchra, and G. gamsii was strongly supported (99%). Sequences of all isolates of the new species also formed a strongly supported (100% BS value) species clade and were inferred as the phylogenetic sister of G. leiopus.
RAxML tree based on the concatenated five gene datasets (ITS, LSU, RPB1, RPB2, and EF1-α) showing the relationship among Gibellula and related genera. Bootstrap proportions ≥ 50% are provided above corresponding nodes; nodes with a BS of 100% are shown as thick lines. The ex-type strains are marked with a superscript T (T) and the isolates encountered in this study are highlighted
Isolation and structure elucidation of alkaloids 1 and 2
G. gamsii was fermented using YMG medium and the fermentation broth was extracted with ethyl acetate and analyzed by HPLC-(DAD)-UV-TOFMS. Two sets of metabolites with characteristic UV absorption bands were observed upon analysis of chromatograms (Fig. 3) and spectroscopic data (UV and MS). Subsequent HPLC-(DAD)-UV-guided fractionation led to the isolation of two new compounds, 1 and 2, belonging to β-carboline class of alkaloids.
Gibellamine A (1) was isolated as pale yellow solid. Its molecular formula was determined as C14H12N2O2 by HRESIMS, requiring 10 degrees of unsaturation. The UV spectrum of 1 exhibited characteristic absorption bands at 217, 284, and 381 nm, diagnostic of a β-carboline chromophore (Wang et al. 2008). The 1H and 13C spectroscopic data of 1 (Table 1) displayed signals assignable to one methyl (C-11), one oxygenated methylene (C-12), five aromatic methines, six unsaturated quaternary carbons, and one carbonyl functionality (C-10). The 1H–1H COSY and HSQC NMR experiments suggested the presence of 1,2-disubstituted benzenoid substructure (C-5–C-8) (Fig. 4). The methyl singlet at δH 2.82 (H3–11) showed HMBC correlation to the carbonyl at δC 203.7 (C-10), suggesting the presence of an acetyl group. Additional HMBC correlation of H3–11/C-1 located the acetyl group at C-1 to form the 1-acetyl-β-carboline moiety in 1. The attachment of the acetyl carbonyl group (C-10) at C-1 with a beta-nitrogen atom in the pyridine sub-structure positioned the chemical shift of H3–11 to a lower field. In the 1H NMR spectrum of 1, a downfield singlet was readily discernible at δH 8.38, indicating the proton was localized at H-4, corresponding to the peri-position of the β-carboline skeleton. The HMBC correlations from H-4 to C-9a and C-4a supported this assumption (Fig. 4). The singlet proton chemical shifts at H-4 and H2–12 also showed HMBC correlations to an oxygenated methylene moiety at δC 66.6 (C-12) and δC 117.5, respectively, suggesting C-12 was connected to C-3. Therefore, a 1,3-disubsituted β-carboline core structure was established. Hence, the gross structure of 1 was constructed and determined as 1-(3-(hydroxymethyl)-9H-pyrido[3,4-b]indol-1-yl)ethan-1-one with the assigned trivial name, gibellamine A.
Gibellamine B (2) was obtained as a yellow syrup. It exhibited a prominent pseudomolecular ion peak at m/z 283.1115 [M + H]+ in HRESIMS, indicating the molecular formula C16H14N2O3, which is 42 mass units greater than that of 1 and corresponds to the addition of an acetyl group. The UV, 1H, and 13C NMR spectroscopic data of 2 strongly resembled those of 1 (Table 1), except additional signals corresponding to the acetyl methyl at δH 2.21 (H3-2′)/δC 21.1 (C-2′) and carboxyl group at δC 170.9 (C-1′) were observed in 2. These changes are supported by HMBC correlations noted between H3-2′ with C-1′ and C-12 (δC 67.6) (Fig. 4). Therefore, compound 2 was identified as (1-acetyl-9H-pyrido[3,4-b]indol-3-yl)methyl acetate and named gibellamine B.
The COSY, HSQC-DEPT, HMBC, 1H, and 13C NMR spectra of compounds 1–2 are available as Supplementary Material.
While a number of β-carboline alkaloid derivatives have been reported from different natural sources, most 1,3-disubstituted carbazole alkaloids bearing an acetyl group at C-1 and a carboxylic acid derivative functionality at C-3 such as 3 (Fig. 5) have been exclusively reported in many alkaloid-containing plant genera (Cao et al. 2012; Wen et al. 2014; Zhang et al. 2015) and bacteria (Huang et al. 2011; Kornsakulkarn et al. 2013). In the fungal kingdom, isolates of marine origin produce harman-type β-carboline alkaloids such as derivatives observed in a Capnodium sp. (He et al. 2016) and Neosartorya pseudofischeri (currently valid name: Aspergillus fischeri; Lan et al. 2016). Interestingly, harman alkaloids (4) (Fig. 5) have also been identified from the insect-associated fungus Ophiocordyceps sinensis (Yang et al. 2011) and from the hypocrealean species, Trichoderma virens (Liu et al. 2012). The new compounds are the first examples of C-16-reduced derivatives compared to other naturally occurring alkaloids of similar type which features a carboxyl group instead of an oxymethylene moiety.
Determination of biological activities
A number of 1,3-disubstituted β-carboline alkaloids have been reported to exhibit a wide range of biological activities including anti-acetylcholinesterase (Zhou et al. 2015), anti-allergenic (Sun et al. 2004), anti-bacterial (Wang et al. 2012), anti-cancer (Tian et al. 2012), anti-inflammatory (Chen et al. 2010), anti-malaria (Huang et al. 2011), molluscicidal (Adewunmi et al. 1989), and neurochemical-modulating (Cain et al. 1982) activities. Thus, 1 and 2 were evaluated for their anti-microbial, anti-biofilm, cytotoxic, and nematicidal activities. Weak to insignificant antimicrobial activity was exhibited by both compounds against yeasts, filamentous fungi, and Gram-positive and Gram-negative bacteria. The alkaloid 1 significantly inhibited biofilm formation in Staphylococcus aureus (ATCC 25923) with an MIC value of 62.5 μg/mL.
The cytotoxic activities (Table 2) were rather weak, but a broader range of cytotoxic effects in comparison to compound 1 was observed for gibellamine B (2) against human lung carcinoma A549, mouse fibroblast L929, and squamous carcinoma A431 cell-lines, with IC50 values of 13, 16, and 24 μg/mL, respectively. Weak cytotoxicity was observed for the major alkaloid gibellamine A (1) against L929 with IC50 value of 28 μg/mL. In addition, both alkaloids did not demonstrate nematicidal activity against C. elegans and antimicrobial activity at any concentration of 10–100 and 2.34–300 μg/mL, respectively.
References
Adewunmi CO, Monache FD, Marquis BB (1989) Molluscicidal activities of some alkaloids. Bull Chem Soc Ethiop 3:103–106
Bunyard BA, Nicholson MS, Royse DJ (1994) A systematic assessment of Morchella using RFLP analysis of the 28S ribosomal RNA gene. Mycologia 86:762–772
Cain M, Weber RW, Guzman F, Cook JM, Barker SA, Rice KC, Crawley JN, Paul SM, Skolnick P (1982) β-Carbolines: synthesis and neurochemical and pharmacological actions on brain benzodiazepine receptors. J Med Chem 25:1081–1091
Cao L-H, Zhang W, Luo J-G, Kong L-Y (2012) Five new β-carboline-type alkaloids from Stellaria dichotoma var. lanceolata. Helv Chim Acta 95:1018–1025
Chaverri P, Samuels GJ (2003) Hypocrea/Trichoderma (Ascomycota, Hypocreales, Hypocreaceae): species with green ascospores. Stud Mycol 48:1–116
Chen Y-F, Kuo P-C, Chan H-H, Kuo I-J, Lin F-W, Su C-R, Yang M-L, Li D-T, Wu T-S (2010) β-Carboline alkaloids from Stellaria dichotoma var. lanceolata and their anti-inflammatory activity. J Nat Prod 73:1993–1998
Chepkirui C, Matasyoh JC, Decock C, Stadler M (2017) Two cytotoxic triterpenes from cultures of a Kenyan Laetiporus sp. (Basidiomycota). Phytochem Lett 20:106–110
Chepkirui C, Yuyama K, Wanga L, Decock C, Matasyoh J, Abraham WR, Stadler M (2018) Microporenic acids A-G, biofilm inhibitors and antimicrobial agents from the basidiomycete Microporus sp. J Nat Prod 81:778–784
Chiriví-Salomón J, Danies G, Restrepo S, Sanjuan T (2015) Lecanicillium sabanense sp. nov. (Cordycipitaceae) a new fungal entomopathogen of coccids. Phytotaxa 234:63–74
D’Alessandro CP, Jones LR, Humber RA, Lastra CCL, Sosa-Gomez DR (2013) Characterization and phylogeny of Isaria spp. Strains (Ascomycota: Hypocreales) using ITS1-5.8S-ITS2 and elongation factor 1-alpha sequences. J Basic Microbiol 53:1–1
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Guindon S, Gascuel O (2003) A simple, fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hall T (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hall T, Biosciences I, Carlsbad C (2011) BioEdit: an important software for molecular biology. GERF Bull Biosci 2:60–61
He H, Ma Z, Wang Q, Liu Y, Xu H (2016) Chemical constituents of the mangrove-associated fungus Capnodium sp. SZ-F22. A new eremophilane sesquiterpene. Nat Prod Res 30:1526–1531
Helaly SE, Kuephadungphan W, Phongpaichit S, Luangsa-ard JJ, Rukachaisirikul V, Stadler M (2017) Five unprecedented secondary metabolites from the spider parasitic fungus Akanthomyces novoguineensis. Molecules 22:991–1001
Helaly SE, Thongbai B, Stadler M (2018) Diversity of biologically active secondary metabolites from endophytic and saprotrophic fungi of the ascomycete order Xylariales. Nat Prod Rep, in press (https://doi.org/10.1039/c8np00010g)
Hongsanan S, Xie N, Liu JK, Dissanayake A, Ekanayaka AH, Raspé O Jayawardena RS, Hyde KD, Jeewon R, Purahong W, Stadler M, Peršoh D (2018) Can we use environmental DNA as holotypes? Fungal Divers, in press :https://doi.org/10.1007/s13225-018-0404-x
Huang H, Yao Y, He Z, Yang T, Ma J, Tian X, Li Y, Huang C, Chen X, Li W, Zhang S, Zhang C, Ju J (2011) Antimalarial β-carboline and indolactam alkaloids from Marinactinospora thermotolerans, a deep sea isolate. J Nat Prod 74:2122–2127
Humber RA, Rombach MC (1987) Torrubiella ratticaudata sp. nov. (Pyrenomycetes: Clavicipitales) and other fungi from spiders on the Solomon Islands. Mycologia 79:375–382
Johnson D, Sung GH, Hywel-Jones NL, Luangsa-ard J, Bischoff JF, Kepler RM, Spatafora JW (2009) Systematics and evolution of the genus Torrubiella (Hypocreales, Ascomycota). Mycol Res 113:279–289
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9:286–298
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Kepler RM, Luangsa-ard JJ, Hywel-Jones NL, Quandt CA, Sung GH, Rehner SA, Aime MC, Henkel TW, Sanjuan T, Zare R, Chen M, Li Z, Rossman AY, Spatafora JW, Shrestha B (2017) A phylogenetically-based nomenclature for Cordycipitaceae (Hypocreales). IMA Fungus 8:335–353
Kepler RM, Sung GH, Ban S, Nakagiri A, Chen MJ, Huang B, Li Z, Spatafora JW (2012) New teleomorph combinations in the entomopathogenic genus Metacordyceps. Mycologia 104:182–197
Kobayasi Y, Shimizu D (1982) Monograph of the genus Torrubiella. Bull Natn Sci Mus Tokyo Ser B 8:43–78
Kornsakulkarn J, Saepua S, Boonruangprapa T, Suphothina S, Thongpanchang C (2013) New β-carboline and indole alkaloids from actinomycete Actinomadura sp. BCC 24717. Phytochem Lett 6:491–494
Kuephadungphan W, Helaly SE, Daengrot C, Phongpaichit S, Luangsa-ard JJ, Rukachaisirikul V, Stadler M (2017) Akanthopyrones A-D, α-pyrones bearing a 4-O-methyl-β-D-glucopyranose moiety from the spider-associated ascomycete Akanthomyces novoguineensis. Molecules 22:1202–1211
Kuephadungphan W, Phongpaichit S, Luangsa-ard JJ, Rukachaisirikul V (2014) Antimicrobial activity of invertebrate-pathogenic fungi in the genera Akanthomyces and Gibellula. Mycoscience 55:127–133
Lan W-J, Fu S-J, Xu M-Y, Liang W-L, Lam C-K, Zhong G-H, Xu J, Yang D-P, Li H-J (2016) Five new cytotoxic metabolites from the marine fungus Neosartorya pseudofischeri. Mar Drugs 14:18
Liu T, Li Z, Wang Y, Wang A, Tian L, Pei Y, Hua H (2012) Secondary metabolites from marine-derived fungus Hypocrea virens (II). Zhongguo Yaowu Huaxue Zazhi 22:124–126, 130
Liu YL, Whelen S, Hall BD (1999) Phylogenetic relationship among ascomycetes: evidence from an RNA polymerase II subunit. Mol Biol Evol 16:1799–1808
Luangsa-ard JJ, Hywel-Jones NL, Manoch L, Samson RA (2005) On the relationships of Paecilomyces sect. Isarioidea species. Mycol Res 109:581–589
Luangsuphabool T, Lumbsch HT, Aptroot A, Piapukiew J, Sangvichien E (2016) Five new species and one new record of Astrothelium (Trypetheliaceae, Ascomycota) from Thailand. Lichenol 48:727–737
Mains EB (1950) The genus Gibellula on spiders in North America. Mycologia 42:306–321
Mongkolsamrit S, Noisripoom W, Thanakitpipattana D, Wutikhun T, Spatafora JW, Luangsa-ard JJ (2018) Disentangling cryptic species with isaria-like morphs in Cordycipitaceae. Mycologia 110:230–257
Murata N, Aoki T, Kusaba M, Tosa Y, Chuma I (2014) Various species of Pyricularia constitute a robust clade distinct from Magnaporthe salvinii and its relatives in Magnaporthaceae. J Gen Plant Pathol 80:66–72
Noumeur SR, Helaly SE, Jansen R, Gereke M, Stradal TEB, Harzallah D, Stadler M (2017) Preussilides A-F, bicyclic polyketides from the endophytic fungus Preussia similis with antiproliferative activity. J Nat Prod 80:1531–1540
Petch T (1932) Gibellula. Annls Mycol 30:386–393
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Rehner SA, Buckley E (2005) A Beauveria phylogeny inferred from nuclear ITS and EF1-α sequences: evidence for cryptic diversification and links to Cordyceps teleomorphs. Mycologia 97:84–98
Rehner SA, Minnis AM, Sung GH, Luangsa-ard JJ, Devotto L, Humber RA (2011) Phylogeny and systematic of the anamorphic, entomopathogenic genus Beauveria. Mycologia 103:1055–1073
Rupcic Z, Rascher M, Kanaki S, Köster RW, Stadler M, Wittstein K (2018) Two new cyathane diterpenoids from mycelial cultures of the medicinal mushroom Hericium erinaceus and the rare species, Hericium flagellum. Int J Mol Sci 19:740
Samson RA, Evans HC (1973) Notes on entomogenous fungi from Ghana I. The genera Gibellula and Pseudogibellula. Acta Bot Neerl 22:522–528
Samson RA, Evans HC (1992) New species of Gibellula on spiders (Araneida) from South America. Mycologia 84:300–314
Samson RA, van Reenen-Hoekstra ES, Evans HC (1989) New species of Torrubiella (Ascomycotina: Clavicipitales) on insects from Ghana. Stud Mycol 31:123–132
Spatafora JW, Sung GH, Sung JM, Hywel-Jones NL, White F (2007) Phylogenetic evidence for an animal pathogen origin of ergot and the grass endophytes. Mol Ecol 16:1701–1711
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Sun B, Morikawa T, Matsuda H, Tewtrakul S, Wu L-J, Harima S, Yoshikawa M (2004) Structures of new β-carboline-type alkaloids with antiallergic effects from Stellaria dichotoma. J Nat Prod 67:1464–1469
Sung GH, Hywel-Jones NL, Sung JM, Luangsa-ard JJ, Shrestha B, Spatafora JW (2007a) Phylogenetic classification of Cordyceps and the clavicipitaceous fungi. Stud Mycol 57:5–59
Sung GH, Spatafora JW (2004) Cordyceps cardinalis sp. nov., a new species of Cordyceps with an east Asian-eastern North American distribution. Mycologia 96:658–666
Sung GH, Spatafora JW, Zare R, Hodge KT, Gams W (2001) A revision of Verticillium sect. Prostrata. II. Phylogenetic analyses of SSU and LSU nuclear rDNA sequences from anamorphs and teleomorphs of the Clavicipitaceae. Nova Hedwigia 72:311–328
Sung GH, Sung JM, Hywel-Jones NL, Spatafora JW (2007b) A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): identification of localized incongruence using a combinational bootstrap approach. Mol Phylogenet Evol 44:1204–1223
Tian J, Shen Y, Li H, Liu R, Shan L, Gao J, Zhang W (2012) Carboline alkaloids from Psammosilene tunicoides and their cytotoxic activities. Planta Med 78:625–629
Tsang C-C, Chan JFW, Pong W-M, Chen JHK, Ngan AHY, Cheung M, Lai CKC, Tsang DNC, Lau SKP, Woo PCY (2016) Cutaneous hyalohyphomycosis due to Parengyodontium album gen. et comb. nov. Med Mycol 54:699–713
Tzean SS, Hsieh LS, Wu WJ (1997) The genus Gibellula on spiders from Taiwan. Mycologia 89:309–318
Tzean SS, Hsieh LS, Wu WJ (1998) Torrubiella dimorpha, a new species of spider parasite from Taiwan. Mycol Res 102:1350–1354
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246
Wang L, Gong X-J, Zhou X, Xian C, Yang S-L, Yang Z-N (2012) Chemical constituents of Psammosilene tunicoides and bacteriostatic activity. Zhongguo Zhongyao Zazhi 37:3577–3580
Wang W, Nam S-J, Lee B-C, Kang H (2008) Antimalarial β-carboline and indolactam alkaloids from Marinactinospora thermotolerans, a deep sea isolate. J Nat Prod 71:163–166
Wen B, Yuan X, Zhang W-D, Chen B-Y, Fu K-L, Li B, Ye J, Shan L, Shen Y-H (2014) Chemical constituents from the aerial parts of Psammosilene tunicoides. Phytochem Lett 9:59–66
Wendt L, Sir EB, Kuhnert E, Heitkämper S, Lambert C, Hladki AI, Romero AI, Luangsa-ard JJ, Srikitikulchai P, Peršoh D, Stadler M (2018) Resurrection and emendation of the Hypoxylaceae, recognised from a multi-gene genealogy of the Xylariales. Mycol Prog 17:115–154
White TJ, Bruns TD, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA for phylogenetics. In: Innis MA et al (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Yang M-L, Kuo P-C, Hwang T-L, Wu T-S (2011) Anti-inflammatory principles from Cordyceps sinensis. J Nat Prod 74:1996–2000
Zare R, Gams W, Culham A (2000) A revision of Verticillium sect. Prostrata I. Phylogenetic studies using ITS sequences. Nova Hedwigia 71:465–480
Zhang Y, Wang G, Lv H, Luo J, Kong L (2015) Two new β-carboline alkaloids from the roots of Gypsophila oldhamiana. Nat Prod Res 29:1207–1211
Zhou A, Hu J, Wang L, Zhong G, Pan J, Wu Z, Hui A (2015) Combined 3d-QSAR, molecular docking, and molecular dynamics study of tacrine derivatives as potential acetylcholinesterase (AChE) inhibitors of Alzheimer’s disease. J Mol Model 21:1–12
Acknowledgements
The National Park, Wildlife and Plant Conservation Department in Thailand is gratefully acknowledged for permission to conduct a study in the protected areas. We also thank Wera Collisi and Christel Kakoschke for conducting the cytotoxicity assay and NMR spectroscopic measurements, respectively.
Funding
This work was supported by the TRF through the Royal Golden Jubilee Ph.D. program (grant number PHD/0163/2556 to W.K.); the Alexander von Humboldt Foundation (AvH Georg-Forster Postdoctoral Research Fellowship to A.P.G.M.); the German Federal Ministry of Education and Research (BMBF); the National Center for Genetic Engineering and Biotechnology (BIOTEC); the National Science and Technology Development Agency (NSTDA); and the Prince of Songkla University Graduate School. This project has also received funding from the European Union’s Horizon 2020 research and innovation programme (RISE) under the Marie Skłodowska-Curie grant agreement no. 645701, project acronym “GoMyTri”; lead beneficiaries J.J.L. and M.S.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editors: Hans-Josef Schroers and Marc Stadler
This article is part of the “Special Issue on hyphomycete taxonomy and diversity in honour of Walter Gams who passed away in April 2017.”
Taxonomic novelty: Gibellula gamsii Kuephadungphan, Tasanathai & Luangsa-ard
Electronic supplementary material
ESM 1
(PDF 341 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Kuephadungphan, W., Macabeo, A.P.G., Luangsa-ard, J.J. et al. Studies on the biologically active secondary metabolites of the new spider parasitic fungus Gibellula gamsii. Mycol Progress 18, 135–146 (2019). https://doi.org/10.1007/s11557-018-1431-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-018-1431-4