Abstract
The taxonomic and genetic diversity of Cortinarius section Calochroi in one of the most biodiversity-rich regions in Europe, the Iberian Peninsula, was investigated through morphological and phylogenetic methods. This combined methodological approach allowed the identification of 15 known species and one new species, Cortinarius ortegae, which is described here. A dichotomous key is provided for field recognition of Calochroi species inhabiting different Iberian and Balearic Island woodlands. Polymorphism analyses within some studied species showed that the distribution of intraspecific lineages is dependent on geography and, in some cases, tree host. Furthermore, the dating analysis suggested that diversification within Calochroi started in the Pliocene and most of the current genetic diversity originated in the Pleistocene. This temporal scenario supports hypotheses in which climatic oscillations in the Quaternary may have been driving the evolution of this group of ectomycorrhizal fungi.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Ballarà J, Cadiñanos Aguirre JA, Campos JC, Escánez LL, Fernández Sasia R, Gutiérrez C, Macau N, Mahiques R, Mateos A, Meléndez A, Pérez A, Pérez-de-Gregorio MÀ, Reyes JD, Salom JC, Santamaría N, Serrano S, Suárez E (2014) Cortinarius ibero-insulares 4. Fungi non Delineati, Pars LXXI–LXXII. Edizioni Candusso, Alassio
Ballarà J, Mahiques R (2010) Phlegmacium raros o nuevos asociados a Dryas octopetala L. J JEC 12:57–68
Ballarà J, Mahiques R, Garrido-Benavent I (2016) Estudi de Cortinariaceae del Parc natural del Cadí-Moixeró (III). Moixeró 8:20–50
Ballarà J, Mahiques R, Garrido-Benavent I (2017) Estudi de Cortinariaceae del Parc natural del Cadí-Moixeró (IV). Moixeró 9:20–49
Bellanger JM (2015) Les cortinaires calochroïdes: une mise au point taxinomique. Documents Mycologiques 36:3–34
Bidaud A, Moënne-Loccoz P, Reumaux P (2001) Atlas des Cortinaires. Pars XI (2), sous-genre Phlegmacium (Fr.) Trog, section Calochroi Moser & Horak. Éditions Fédération Mycologique Dauphiné-Savoie, Annecy, France
Bjorbækmo M, Carlsen T, Brysting A et al (2010) High diversity of root associated fungi in both alpine and arctic Dryas octopetala. BMC Plant Biol 10:244. https://doi.org/10.1186/1471-2229-10-244
Bödeker ITM, Clemmensen KE, de Boer W, Martin F, Olson Å, Lindahl BD (2014) Ectomycorrhizal Cortinarius species participate in enzymatic oxidation of humus in northern forest ecosystems. New Phytol 203:245–256. https://doi.org/10.1111/nph.12791
Cadiñanos Aguirre JA (2004) Cortinarius subgen. Phlegmacium raros o interesantes. Fungi non delineati, Pars XXIX. Edizioni Candusso. Alassio, Italy
Cailleux A (1981) Code des couleurs des sols. Boubée, France
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552. https://doi.org/10.1093/oxfordjournals.molbev.a026334
Clericuzio M, Dovana F, Bellanger J-M, Brandrud TE, Dima B, Frøslev TG, Boccardo F, Jeppesen TS, Vizzini A (2017) Cortinarius parasuaveolens (= C. pseudogracilior): new data and a synonymy of a very poorly known species of section Calochroi. Sydowia 69. https://doi.org/10.12905/0380.sydowia69-2017-0213
Delaporte A, Eyssartier G, Moenne-Loccoz P (2002) Cortinarius rapaceotomentosus sp. nov., un nouveau cortinaire proche de Cortinarius europaeus. Bulletin de la Société Mycologique de France 118:11–18
Doyle JJ, Doyle JL (1987) A rapid procedure for DNA purification from small quantities of fresh leaf tissue. Phytochemical Bulletin 19:11–15
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29:1969–1973. https://doi.org/10.1093/molbev/mss075
Fernández CW, Nguyen NH, Stefanski A, Han Y, Hobbie SE, Montgomery RA, Reich PB, Kennedy PG (2017) Ectomycorrhizal fungal response to warming is linked to poor host performance at the boreal-temperate ecotone. Glob Chang Biol 23:1598–1609. https://doi.org/10.1111/gcb.13510
Flores-Rentería D, Curiel Yuste J, Rincón A, Brearley FQ, García-Gil JC, Valladares F (2015) Habitat fragmentation can modulate drought effects on the plant–soil–microbial system in Mediterranean Holm oak (Quercus ilex) forests. Microb Ecol 69:798–812. https://doi.org/10.1007/s00248-015-0584-9
Frøslev TG, Brandrud TE, Dima B (2017) Cortinarius stjernegaardii and C. kristinae (Basidiomycota, Agaricales), two new European species with a mainly northern distribution. Mycol Prog 16:145–153. https://doi.org/10.1007/s11557-016-1264-y
Frøslev TG, Jeppesen TS, Læssøe T (2006) Seven new calochroid and fulvoid species of Cortinarius. Mycol Res 110:1046–1058. https://doi.org/10.1016/j.mycres.2006.05.012
Frøslev TG, Jeppesen TS, Læssøe T, Kjøller R (2007) Molecular phylogenetics and delimitation of species in Cortinarius section Calochroi (Basidiomycota, Agaricales) in Europe. Mol Phylogenet Evol 44:217–227. https://doi.org/10.1016/j.ympev.2006.11.013
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes—application for the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294x.1993.tb00005.x
Garnica S, Schön ME, Abarenkov K et al (2016) Determining threshold values for barcoding fungi: lessons from Cortinarius (Basidiomycota), a highly diverse and widespread ectomycorrhizal genus. FEMS Microbiol Ecol 92:fiw045. https://doi.org/10.1093/femsec/fiw045
Garnica S, Spahn P, Oertel B, Ammirati J, Oberwinkler F (2011) Tracking the evolutionary history of Cortinarius species in section Calochroi, with transoceanic disjunct distributions. BMC Evol Biol 11:213. https://doi.org/10.1186/1471-2148-11-213
Garnica S, Weiß M, Oertel B, Ammirati J, Oberwinkler F (2009) Phylogenetic relationships in Cortinarius, section Calochroi, inferred from nuclear DNA sequences. BMC Evol Biol 9(1). https://doi.org/10.1186/1471-2148-9-1
Garrido-Benavent I, Ballarà J, Mahiques R (2015) New insights into subg. Phlegmacium sect. Calochroi: adding morphological and molecular data from Mediterranean representatives, with special regard to Cortinarius prasinus, C. natalis and C. murellensis species complexes. Journal des J.E.C. 17:8–78
Garrido-Benavent I, Ballarà J, Mahiques R (2016) Dos nuevas especies de Cortinarius, subgénero Telamonia, del Parque Natural del Cadí-Moixeró (noreste de la Península Ibérica), basadas en caracteres morfológicos y moleculares. Journal des J.E.C. 18:66–85
Geml J, Timling I, Robinson CH, Lennon N, Nusbaum HC, Brochmann C, Noordeloos ME, Taylor DL (2011) An arctic community of symbiotic fungi assembled by long-distance dispersers: phylogenetic diversity of ectomycorrhizal basidiomycetes in Svalbard based on soil and sporocarp DNA. J Biogeogr 39:74–88. https://doi.org/10.1111/j.1365-2699.2011.02588.x
Han LH, Feng B, Wu G et al (2017) African origin and global distribution patterns: evidence inferred from phylogenetic and biogeographical analyses of ectomycorrhizal fungal genus Strobilomyces. J Biogeogr. https://doi.org/10.1111/jbi.13094
Harrower E, Ammirati JF, Cappuccino AA et al (2011) Cortinarius species diversity in British Columbia and molecular phylogenetic comparison with European specimen sequences. Botany 89:799–810. https://doi.org/10.1139/b11-065
Harrower E, Bougher NL, Henkel TW, Horak E, Matheny PB (2015) Long-distance dispersal and speciation of Australasian and American species of Cortinarius sect. Cortinarius. Mycologia 107:697–709. https://doi.org/10.3852/14-182
Hebert PD, Cywinska A, Ball SL (2003) Biological identifications through DNA barcodes. Proc R Soc Lond B Biol Sci 270:313–321. https://doi.org/10.1098/rspb.2002.2218
Hewitt GM (2011) Mediterranean peninsulas: the evolution of hotspots. In: Zachos F, Habel J (eds) Biodiversity hotspots. Springer, Berlin, Heidelberg, pp 123–147. https://doi.org/10.1007/978-3-642-20992-5_7
Horton BM, Morag G, Davidson NJ, Ratkowsky DA, Close DC, Wardlaw TJ, Mohammed C (2017) An assessment of ectomycorrhizal fungal communities in Tasmanian temperate high-altitude Eucalyptus delegatensis forest reveals a dominance of the Cortinariaceae. Mycorrhiza 27:67–74. https://doi.org/10.1007/s00572-016-0725-0
Katoh K, Standley DM (2013) MAFFT: iterative refinement and additional methods. Methods Mol Biol 1079:131–146. https://doi.org/10.1007/978-1-62703-646-7_8
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf43
Kyaschenko J, Clemmensen KE, Hagenbo A, Karltun E, Lindahl BD (2017) Shift in fungal communities and associated enzyme activities along an age gradient of managed Pinus sylvestris stands. ISME J 11:863–874. https://doi.org/10.1038/ismej.2016.184
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701. https://doi.org/10.1093/molbev/mss020
Leavitt SD, Divakar PK, Ohmura Y, Wang L, Esslinger TL, Lumbsch HT (2015) Who’s getting around? Assessing species diversity and phylogeography in the widely distributed lichen-forming fungal genus Montanelia (Parmeliaceae, Ascomycota). Mol Phylogenet Evol 90:85–96. https://doi.org/10.1016/j.ympev.2015.04.029
Leavitt SD, Esslinger TL, Lumbsch HT (2012a) Neogene-dominated diversification in neotropical montane lichens: dating divergence events in the lichen-forming fungal genus Oropogon (Parmeliaceae). Am J Bot 99:1764–1777. https://doi.org/10.3732/ajb.1200146
Leavitt SD, Esslinger TL, Divakar PK, Lumbsch H (2012b) Miocene and Pliocene dominated diversification of the lichen-forming fungal genus Melanohalea (Parmeliaceae, Ascomycota) and Pleistocene population expansions. BMC Evol Biol 12:176. https://doi.org/10.1186/1471-2148-12-176
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Liimatainen K, Carteret X, Dima B, Kytövuori I, Bidaud A, Reumaux P, Niskanen T, Ammirati JF, Bellanger J-M (2017) Cortinarius section Bicolores and section Saturnini (Basidiomycota, Agaricales), a morphogenetic overview of European and North American species. Persoonia 39:175–200. https://doi.org/10.3767/persoonia.2017.39.08
Liimatainen K, Niskanen T, Dima B, Kytövuori I, Ammirati JF, Frøslev TG (2014) The largest type study of Agaricales species to date: bringing identification and nomenclature of Phlegmacium (Cortinarius) into the DNA era. Persoonia 33:98–140. https://doi.org/10.3767/003158514x684681
Liu K, Raghavan S, Nelesen S, Linder CR, Warnow T (2009) Rapid and accurate large-scale coestimation of sequence alignments and phylogenetic trees. Science 324:1561–1564. https://doi.org/10.1126/science.1171243
Liu K, Warnow TJ, Holder MT, Nelesen SM, Yu J, Stamatakis AP, Linder CR (2011) SATe-II: very fast and accurate simultaneous estimation of multiple sequence alignments and phylogenetic trees. Syst Biol 61:90–106. https://doi.org/10.1093/sysbio/syr095
Looney BP, Ryberg M, Hampe F, Sánchez-García M, Matheny PB (2016) Into and out of the tropics: global diversification patterns in a hyperdiverse clade of ectomycorrhizal fungi. Mol Ecol 25:630–647. https://doi.org/10.1111/mec.13506
Mahiques R (2011) Flora corològica i bibliogràfica dels Cortinarius Ibero-Insulars. Butlletí de la Societat Micològica Valenciana 16:121–162
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES science gateway for inference of large phylogenetic trees. In: Proceedings of the gateway computing environments workshop (GCE), pp 1–8 New Orleans
Myers N, Mittermeier RA, Mittermeier CG, Da Fonseca GA, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858. https://doi.org/10.1038/35002501
Niskanen T, Liimatainen K, Mahiques R, Ballarà J, Kytövuori I (2011) Cortinarius badiolaevis, a new conifer-associated, darkening species in the subgenus Telamonia (Basidiomycota, Agaricales). Mycol Prog 10:101–105. https://doi.org/10.1007/s11557-010-0680-7
Nouhra E, Urcelay C, Longo S, Tedersoo L (2013) Ectomycorrhizal fungal communities associated to Nothofagus species in Northern Patagonia. Mycorrhiza 23:487–496. https://doi.org/10.1007/s00572-013-0490-2
Oertel B, Schmidt-Stohn G, Saar G (2009) Die Laugenreaktion am Stielbasisfilz bei Fruchtkörpern von Cortinarius, Subgen. Phlegmacium—Eine Bestandsaufnahme 23 Jahre nach Entdeckung dieser neuartigen Reaktion. Journal des J.E.C. 11:20–31
Ortega A, Suárez-Santiago VN, Reyes JD (2008) Morphological and ITS identification of Cortinarius species (section Calochroi) collected in Mediterranean Quercus woodlands. Fungal Divers 29:73–88
Pachauri RK, Allen MR, Barros VR et al (2014) Climate change 2014: synthesis report. Contribution of working groups I, II and III to the fifth assessment report of the Intergovernmental Panel on Climate Change. IPCC
Pérez-Izquierdo L, Zabal-Aguirre M, Flores-Rentería D, González-Martínez S, Buée M, Rincón A (2017) Functional outcomes of fungal community shifts driven by tree genotype and spatial–temporal factors in Mediterranean pine forests. Environ Microbiol 19:1639–1652. https://doi.org/10.1111/1462-2920.13690
Puillandre N, Lambert A, Brouillet S, Achaz G (2012) ABGD, automatic barcode gap discovery for primary species delimitation. Mol Ecol 21:1864–1877. https://doi.org/10.1111/j.1365-294x.2011.05239.x
Rodríguez A, Curiel Yuste J, Rey A, Durán J, García-Camacho R, Gallardo A, Valladares F (2016) Holm oak decline triggers changes in plant succession and microbial communities, with implications for ecosystem C and N cycling. Plant Soil 414:247–263. https://doi.org/10.1007/s11104-016-3118-4
Rodríguez-Sánchez F, Hampe A, Jordano P, Arroyo J (2010) Past tree range dynamics in the Iberian Peninsula inferred through phylogeography and palaeodistribution modelling: a review. Rev Palaeobot Palynol 162:507–521. https://doi.org/10.1016/j.revpalbo.2010.03.008
Ronquist F, Teslenko M, van der Mark P et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Ryberg M, Matheny PB (2011) Asynchronous origins of ectomycorrhizal clades of Agaricales. P R Soc Lond B Biol 279:2003–2011. https://doi.org/10.1098/rspb.2011.2428
Ryberg M, Larsson E, Molau U (2009) Ectomycorrhizal diversity on Dryas octopetala and Salix reticulata in an alpine cliff ecosystem. Arct Antarct Alp Res 41:506–514. https://doi.org/10.1657/1938-4246-41.4.506
Salom JC, Siquier JL, Mahiques R (2015) El gènere Cortinarius a les Illes Balears (Espanya) I. Revista Catalana de Micologia 36:11–27
Sánchez-Ramírez S, Tulloss RE, Amalfi M, Moncalvo J-M (2015) Palaeotropical origins, boreotropical distribution and increased rates of diversification in a clade of edible ectomycorrhizal mushrooms (Amanita section Caesareae). J Biogeogr 42:351–363. https://doi.org/10.1111/jbi.12402
Schenk JJ (2016) Consequences of secondary calibrations on divergence time estimates. PLoS One 11:e0148228. https://doi.org/10.1371/journal.pone.0148228
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690. https://doi.org/10.1093/bioinformatics/btl446
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML Web servers. Syst Biol 57:758–771. https://doi.org/10.1080/10635150802429642
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Teasdale SE, Beulke AK, Guy PL, Orlovich DA (2013) Environmental barcoding of the ectomycorrhizal fungal genus Cortinarius. Fungal Divers 58:299–310. https://doi.org/10.1007/s13225-012-0218-1
Truong C, Mujic AB, Healy R, Kuhar F, Furci G, Torres D, Niskanen T et al (2017) How to know the fungi: combining field inventories and DNA-barcoding to document fungal diversity. New Phytol 214:913–919. https://doi.org/10.1111/nph.14509
White TJ, Bruns TD, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols. Academic Press, San Diego, pp 315–322
Zachos J, Pagani M, Sloan L, Thomas E, Billups K (2001) Trends, rhythms, and aberrations in global climate 65 Ma to present. Science 292:686–693. https://doi.org/10.1126/science.1059412
Acknowledgements
We are very grateful to J.A. Cadiñanos Aguirre, J.D. Reyes and E. Suárez for sharing some Calochroi collections. We also thank S. García and P. Alvarado for collaborating in the production of molecular data. Asunción de los Ríos (Dept. of Biogeochemistry and Microbial Ecology, National Museum of Natural Sciences) is also thanked for providing necessary resources for conducting this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Zhu-Liang Yang
Electronic supplementary material
Online Resource 1
GenBank accession numbers of nrITS sequences used in phylogenetic and population genetic analyses (PDF 294 kb)
Online Resource 2
Calochroid species for which polymorphism statistics were calculated (see the “Material and methods” section), and associated plant species and source (in square brackets). Specimens studied in the present work are highlighted in bold. The type locality is indicated when known. Codes correspond with different countries (see www.countrycode.org). “m.f.”, mixed forest (PDF 301 kb)
Online Resource 3
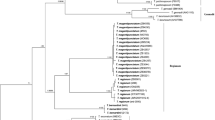
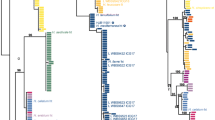
Phylogram depicting the evolutionary relationships of members in Cortinarius section Calochroi obtained with the software SATé v. 2.2.7 and based on nrITS data. Terminals belonging to one phylogenetic species and represented by two or more sequences are collapsed for better visualisation. Species found in the Iberian Peninsula are highlighted, including the new species reported in the present study. Bootstrap support values are given for each node, and branches showing strong support (BP ≥ 70%) are in bold (PDF 439 kb)
Rights and permissions
About this article
Cite this article
Mahiques, R., Ballarà, J., Salom, J.C. et al. Morphogenetic diversity of the ectomycorrhizal genus Cortinarius section Calochroi in the Iberian Peninsula. Mycol Progress 17, 815–831 (2018). https://doi.org/10.1007/s11557-018-1394-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-018-1394-5