Abstract
Ochroconis globalis, a novel species of the melanized genus Ochroconis (Sympoventuriaceae, Venturiales), is described and illustrated and distinguished phenotypically and molecularly from existing ochroconis-like species. Phylogenetic analyses using nuclear ribosomal DNA genes (nuSSU, ITS, nuLSU) and coding gene fragments (ACT1, BT2, TEF1) revealed clustering of all strains as a monophyletic clade which was well separated from known Ochroconis species. Most strains of the new species were obtained from human-made environments, while the natural habitat of the species remains enigmatic. The new combination Ochroconis musae is introduced for one of the most commonly encountered Ochroconis species, and a phenotypic key to all species is provided.
Similar content being viewed by others
Introduction
The genus Ochroconis, typified by O. constricta, was described by de Hoog and von Arx (1973) for melanized fungi with sympodial conidiogenesis and septate, ellipsoidal conidia. The genus was differentiated from the sister genus Scolecobasidium Abbott (1927) which was originally described with T- or Y-shaped conidia. Samerpitak et al. (2014) showed that the status of Scolecobasidium was doubtful and combined species with forked conidia in Ochroconis on phylogenetic grounds as members of Sympoventuriaceae, although some of the species with elaborate morphology still remain outside the family and are in need of correct classification. Colonies of members of the expanded genus Ochroconis are characteristically rust- to olivaceous-brown and produce small, brownish conidiophores bearing small numbers of mostly rough-walled conidia. All members of the genus share rhexolytic conidial liberation (Ellis 1971). Several species with more elaborate, branched conidia had been added (Matsushima 1971; 1975), and during the following decades more species, such as O. tshawytschae (Kirilenko and All-Achmed 1977) and O. gamsii (de Hoog 1985), were recognized in addition to the classical species O. anellii, O. constricta and O. humicola. Samerpitak et al. (2014) introduced a new genus, Verruconis, for a group of thermophilic species around Ochroconis gallopava. Almost synchronously with the paper of Samerpitak et al. (2014), Hao et al. (2013) proposed a new species of Scolecobasidium, and being unaware of the doubtful status of the generic type species S. terreum opted for maintenance of Scolecobasidium at the expense of Ochroconis for the entire species group. Given the ambiguity of Scolecobasidium this choice is less appropriate and is corrected in the present paper; some necessary name changes are proposed.
Machouart et al. (2014) investigated conserved genes (nuSSU, nuLSU, mtSSU, and RPB2) of the species at hand and found that both Ochroconis and Verruconis belonged to the order Venturiales, family Sympoventuriaceae. Detailed taxonomy was elaborated by Samerpitak et al. (2014) using nuclear ribosomal (nuSSU, nuLSU, ITS) and partial coding genes (actin: ACT1, β-tubulin: BT2, translation elongation factor 1-α: TEF1). In this revision, 13 species were recognized in Ochroconis and three in Verruconis. The latter genus comprised the human opportunistic neurotroph, Verruconis gallopava and two related species. Remarkably large phylogenetic distances were noted among and within the species of Ochroconis and Verruconis, which indicated the possible existence of additional, presently unrecognized taxa.
Some basic ecological trends can be observed in Ochroconis and Verruconis. Ochroconis species are mesophilic and often oligotrophic. Ochroconis anellii was isolated from a stalactite (Graniti 1962), O. lascauxensis from ancient drawings on a cave wall, and O. anomala from sediment in the same cave, i.e. the Lascaux Cave in France (Martin-Sanchez et al. 2012). Ochroconis species morphologically similar to O. humicola were reported from wet areas in the domestic environment, such as bathrooms. Several ochroconis-like strains were isolated from soil or water (Lian and de Hoog 2010). Verruconis species differ by being thermophilic and have repeatedly been recovered from hot water and from brains of warm-blooded animals including humans.
The present paper presents a taxonomic study of eleven ochroconis-like strains from various sources. Phenotypic and genotypic characters of the strains were evaluated in view of refined species delimitations and a novel Ochroconis species is introduced.
Materials and methods
Phenotypic studies
Eleven strains of an unknown Ochroconis species (Table 1) were cultured on oatmeal (OA) and malt extract agars (MEA), respectively, and incubated at 24 °C for 14 days. Morphological observations were carried out as described by Samerpitak et al. (2014). To investigate the optimal temperature for growth, all strains were cultured on MEA and incubated for three weeks at temperatures varying from 4 to 40 °C with 3 °C intervals. Colony diameters were measured after 3, 7, 11, 14, 18, and 21 days.
Phylogeny
Eleven unknown strains including 18 type and reference strains of Ochroconis and Verruconis species (Table 2, Samerpitak et al. 2014) were included in phylogenetic analyses. DNA extraction was performed according to Feng et al. (2014). Six markers, viz. nuSSU, nuLSU, ITS, ACT1, BT2, and TEF1 were amplified and sequenced by PCR using primers and conditions as reported earlier (Badali et al. 2008; Najafzadeh et al. 2009; Feng et al. 2014). BioNumerics v. 4.61 (Applied Maths, Sint-Martens-Latem, Belgium) was employed for first iterative alignments. Sequences of nuLSU, nuSSU and ITS were aligned with the web-based program Muscle (www.ebi.ac.uk/Tools/msa/muscle), and ACT1, BT2, and TEF1 with the program Mafft (http://www.ebi.ac.uk/Tools/msa/mafft). Sequence alignments were adjusted using BioEdit v. 7.0.5.2. Mole% G + C of ITS was calculated using BioEdit v. 7.0.5.2, and distances between species by Mega5 (Tamura et al. 2011). Sequences were concatenated following Samerpitak et al. (2014). Multi-locus analysis was performed using the Bayesian approach with MrBayes v. 3.1.2 from the CIPRES Science Gateway (Miller et al. 2010). Two parallel runs of 10,000,000 generations were done with a sampling frequency of 1,000 trees. A burnin tree sample of 10 % was discarded. Maximum likelihood (ML) using Tamura-Nei and GTR+I as the best model with 1,000 bootstrap replicates, and maximum parsimony (MP) with 1,000 bootstrap replicates were also carried out in Mega5 (Tamura et al. 2011). Presented tree was obtained with Bayesian approach. Tree reconstruction, visualization, and editing were done with TreeView v. 1.6.6, FigTree v. 1.1.2 and Mega5.
Results
The overall mean distance of ITS sequences among 15 Ochroconis species and the unidentified Ochroconis was 13.4 %. ITS sequences of the unidentified Ochroconis were 687 bp in length including 317 bp for ITS1, 157 bp for 5.8S, and 213 bp for ITS2. All 11 strains of the unknown species were 99.8 − 100 % identical in rDNA genes (data not shown). Judging from ITS phylogeny of all Ochroconis and Verruconis species sequenced to date (Samerpitak et al. 2014), the strains were at considerable distance to the described species, O. tshawytschae, O. anellii, O. anomala, O. verrucosa, and O. lascauxensis. A minimum distance of 6.5 % was found with O. tshawytschae. ITS length and mole% G + C seem to be specific to each Ochroconis-Verruconis species including the unidentified one, ranging from 566 bp (O. cordanae) to 754 bp (O. anomala), and from 48.82 % (O. sexualis) to 60.96 % (O. tshawytschae) (Table 3).
The dataset of six concatenated sequences contained 4,913 characters of which 1,271 were parsimony-informative. A multi-locus analysis, applying all algorithms mentioned above confirmed that the investigated strains formed a separate, strongly supported cluster at 1/100 %/100 % (BI/ML/MP) and that the 1 strains represented a hitherto undescribed member of the O. tshawytschae species group with smallest distances to O. anellii and O. lascauxensis (Fig. 1).
Taxonomy
Ochroconis globalis Samerpitak, Duarte, Attili-Angelis & de Hoog, sp. nov. – Figs. 2 and 3. Mycobank MB807506
Etymology: the name refers to the fungus’ wide geographical distribution.
Holotype: CBS 119644 (living) = CBS H-21940 (dried), from indoor sample, dwelling house, Germany, Düsseldorf, 2002.
Description based on CBS 119644 at 24 °C after 2 weeks in darkness.
On OA, colonies 25–30 mm in diameter after 2 weeks, moderately expanding, smooth, dry, flat, greyish brown to dark brown. On MEA, colonies attaining 18–20 mm in diameter after 2 weeks, flat, velvety to floccose with some shallow radial fissures, brownish olive green to dark olive green with a 0.5 mm submerged colony margin, reverse as dark olive, green and brown in the central portion, on MEA, hyphae subhyaline to pale brown, smooth- and thick-walled; 1.4–3.0 μm wide, coiled hyphae usually present. Conidiophores mostly arising laterally from vegetative hyphae, erect or flexuous, cylindrical with 1–2 septa, 15–60 × 2–4.7 μm, brown, smooth-walled, with sympodially proliferating conidiogenous cells bearing one or more denticles in the apical region, denticles cylindrical, subhyaline, up to 2 μm long. Conidia ellipsoidal to cylindrical sometimes slightly apiculate at the base, 5.5–10.5 × 3.0–5.5 μm, smooth-walled, pale brown, mostly 2-celled, becoming verrucose and constricted at the median septum when old. Conidial secession rhexolytic, frills remaining visible on denticle and on conidial base. Cardinal temperatures on MEA: minimum at 4 °C, optimum at 24–27 °C, maximum at 33 °C. No growth at 37 °C, leading to death after 4 weeks of incubation.
Note: To clarify the new taxonomic position of the new taxon O. globalis in the Ochroconis lineage, the available type and representative strains of species in the genera Ochroconis and Verruconis (Table 2) were included in a phylogenetic comparison. Ochroconis atlantica ATCC 32684 (Wellman 1975) was not available for study, but with its muriform conidia and its different habitat, this species was judged to be significantly different from O. globalis. Morphologically, O. globalis resembles O. constricta, but conidia of O. globalis are larger and more variable in shape, i.e. ellipsoidal to cylindrical, 2- to 4-celled, with less rough walls at young age, and with longer, cylindrical conidiophores. Conidia of O. globalis are also similar to those of O. musae and O. cordanae, but have a constriction at the median septum and have more prominently verrucose walls at later stages of development. Conidiophore lengths varied from long cylindrical – similar to those of O. gamsii and O. humicola – to shorter, like those of O. cordanae and O. musae. Therefore, given the high degree of variation (Fig. 3), morphological characters are insufficient to unambiguously differentiate O. globalis. However, when compared within members of the O. tshawytschae species group (O. anellii, O. lascauxensis, O. anomala, O. verrucosa, and O. tshawytschae), O. globalis can be differentiated morphologically from other species by features listed in the key and summarized in Fig. 4.
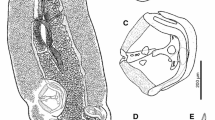
Overview of Ochroconis species with septate conidia. a. O. constricta CBS 211.53. b. O. globalis CBS 119644. c. O. anellii CBS 284.64. d. O. lascauxensis CBS 131815. e. O. anomala CBS 131816. f. O. verrucosa CBS 383.81. g. O. tshawytschae CBS 100438. h. O. gamsii CBS 239.78. i. O. humicola CBS 116655. j. O. cordanae CBS 475.80. k. O. musae CBS 729.95. l. O. minima CBS 510.71. Scale bar = 10 μm
Ochroconis musae (G.Y. Sun & Lu Hao) Samerpitak & de Hoog, comb. nov. Mycobank MB808843
≡ Scolecobasidium musae G.Y. Sun & Lu Hao, Mycol. Prog. 12: 492, 2013 (basionym)
= Ochroconis mirabilis Samerpitak & de Hoog, Fungal Divers. 65: 110, 2014.
Holotype (dried culture): HMAS 243664 from fruit surface of Japanese fiber banana, Musa basjoo, China, Hainan Province, Haikou City, Ledong county, L. Hao; culture ex-type CGMCC 3.14990 = 0HLHKBJ-22.
Note: Full descriptions of this species were given by Hao et al. (2013) and by Samerpitak et al. (2014) under its synonymous name O. mirabilis. The species was classified in Scolecobasidium (Hao et al. 2013) without consideration of the doubtful identity of the genus Scolecobasidium. The large phylogenetic distances among Ochroconis and Verruconis species were neglected. Samerpitak et al. (2014) described O. mirabilis for the most common Ochroconis species, which mostly had been reported under the name of the phenotypically similar species O. constricta; the authors were unaware of the almost synchronous description of the same species as S. musae by Hao et al. (2013). The ITS sequence of strain GS-2012 (=0HLHKBJ-22) had been deposited at GenBank as Scolecobasidum sp. (JQ364738) and was included by Samerpitak et al. (2014) under O. mirabilis. LSU sequence (JQ364739) of O. musae CGMCC 3.14990 (=HLHKBJ-22) is almost identical to that of O. mirabilis CBS 729.95 (KF156144). Consequently, to solve this taxonomic dilemma with two names having been introduced for the same fungus, the new combination Ochroconis musae is proposed in the present study.
Key to species of Ochroconis
1a. Conidia ovoid to cylindrical……………………………2
1b. Conidia muriform………………………...O. atlantica
2a. Conidia mostly 2-celled…….…………………..……3
2b. Conidia mostly 4-celled………………….…..……12
3a. Conidia smooth-walled, clavate to T- or Y-shaped…................……………………….O. minima
3b. Conidia smooth-walled to verrucose, ovoid to cylindrical……..............…………………......…..…4
4a. Conidiophores flask-shaped to cylindrical………..…5
4b. Conidiophores long cylindrical mostly unbranched…....................................................….…..10
5a. Conidia constricted at the septum…...................….…6
5b. Conidia rarely constricted at the septum….............…7
6a. Conidia verrucose, cylindrical, mostly 2-celled………….........................................O. constricta
6b. Conidia smooth-walled to verrucose, cylindrical, sometimes 3- or 4-celled…………......…………O. globalis
7a. Conidia ovoid to cylindrical, 2- to 3-celled................................................……O. anomala
7b. Conidia cylindrical, mostly 2-celled…....…...........…8
8a. Conidiophores subhyaline to pale brown, flask-shaped to cylindrical, branched………............….…O. musae
8b. Conidiophores brown, cylindrical, erect, mostly unbranched………………….............………….…..9
9a. Conidiophores 10–50 × 2.5–3.5 μm, mostly arising laterally; conidia cylindrical..................…O. cordanae
9b. Conidiophores 10–100 × 2.5–3.5 μm, arising both terminally and laterally; conidia ovoid to cylindrical …......................………......….....…O. crassihumicola
10a. Conidiophores brown, cylindrical, 20–100 × 2–3 μm, erect, sometimes short rhizoid present; conidia long cylindrical, 8–20 × 3–5 μm ……...……..O. humicola
10b. Conidiophores dark- to red brown, cylindrical, 20–80 × 2–3 μm, erect to flexuous; conidia mostly one septum, straight to curved and unilaterally flattened…………..........................................…….11
11a. Conidia 6–9 × 2–3 μm, straight to curved……........................................…..O. gamsii
11b. Conidia larger, 8–12 × 3–4 μm, mostly straight………..................……….O. macrozamiae
12a. Conidia cylindrical, 2- to 4-celled.…….………….13
12b. Conidia cylindrical to slightly fusiform, mostly 4-celled.....................……………………………….15
13a. Conidiophores cylindrical; conidia long cylindrical, 20–50 × 10–20 μm, 2- to 4-celled, round at both ends, verruculose….….………….................O. longiphora
13b. Conidiophores cylindrical to acicular; conidia smaller than 20–50 × 10–20 μm, smooth-walled, verrucose or tuberculat..............................................................…14
14a. Conidia cylindrical, mostly 4-celled, smooth-walled to verrucose…...............................…….……O. sexualis
14b. Conidia 2- to 4-celled, verrucose to tuberculated….......…......…....…...O. lascauxensis
15a. Conidia smooth-walled to verrucose, often constricted at septa………………………….............….O. anellii
15b. Conidia verrucose, rarely constricted at septa….......16
16a. Conidiophores short to long cylindrical, erect to flexuous; conidia cylindrical, verrucose, single; sometimes chlamydospores present…...…O. tshawytschae
16b. Conidiophores cylindrical; conidia cylindrical to slightly fusiform, coarsely verrucose, sometimes in branched or unbranced chains; chlamydospores absent…….....................................………O. verrucosa
Discussion
In phylogenetic analyses of both conserved and coding genes, O. globalis could easily be distinguished from described Ochroconis species. All 11 strains of the new species were 99.8 − 100 % identical in rDNA genes and three coding genes, and clustered together concordantly in all single-gene analyses (data not shown). A multi-locus analysis of 11 available strains showed strict concordance in all investigated genes. Because of the high degrees of variability found even in conserved markers, ITS and LSU were recommended as the best diagnostic and barcoding candidates for Ochroconis and Verruconis species (Samerpitak et al. 2014), which also holds true for O. globalis. Distances between species were large enough to recognize O. globolis unambiguously in all investigated genes, each having satisfactory usability as barcoding markers. The large barcoding gaps between Ochroconis and Verruconis species are reflected by differences in length and mole% G + C of ITS (Table 3). For routine diagnostics of Ochroconis and Verruconis species, ITS sequencing is the most effective tool.
The majority of investigated O. globalis strains were isolated from water or from domestic wet cells such as bathrooms. A similar habitat choice was observed in O. musae [as O. mirabilis in Samerpitak et al. (2014) or as O. humicola in Lian and de Hoog (2010) and Heinrichs et al. (2013a, b)]. These habitats suggest oligotrophism accompanied by low competitive ability, as is known for black yeasts (Sudhadham et al. 2008). Water might play an important role in their distribution (Heinrichs et al. 2013a, b). One of the investigated strains, CBS 119643, was found as a superficial opportunist on skin of a human foot, but no proven case report has as yet been published. Lian and de Hoog (2010) suggested that ability to grow at 37 °C was not necessary for cutaneous infection. Strains may have entered softened human skin during bathing, as hypothesized by Satow et al. (2008) and Lian and de Hoog (2010). CBS 135766 was isolated from a living salmon (Salmo salar) with visceral infection. The Brazilian strain of O. globalis, CBS 131956, was isolated from the cuticle of gynes of an ant species, Atta capiguara, and shared a similar habitat with some strains of O. cordanae and O. sexualis (Samerpitak et al. 2014). CBS 131956 is the only strain from a tropical climate, while remaining O. globalis strains originated from temperate zones. The tropical ant-associated strain grew optimally at 27 to 30 °C. Investigations on fungal diversity and ecology associated with social insects are currently in progress. Given the high diversity of sources of isolation of O. globalis and the scant information on their ecology other than yield in culture, the actual natural habitat of these oligotrophic fungi remains enigmatic.
References
Abbott EV (1927) Scolecobasidium, a new genus of soil fungi. Mycologia 19:29–31
Badali H, Gueidan C, Najafzadeh MJ, Bonifaz A, Gerrits van den Ende AHG, de Hoog GS (2008) Biodiversity of the genus Cladophialophora. Stud Mycol 61:175–191
de Hoog GS, von Arx JA (1973) Revision of Scolecobasidium and Pleurophragmium. Kavaka 1:55–60
de Hoog GS (1985) Taxonomy of Dactylaria complex IV-VI. Stud Mycol 26:1–60
Ellis MB (1971) Dematiaceous Hyphomycetes. Commonwealth Mycological Institute, Kew, pp 43–46
Feng P, Lu Q, Gerrits van den Ende AHG, Sun J, Li RY, Xi LY, Vicente VA, Lai W, Lu C, de Hoog GS (2014) Cyphellophora and its relatives in Phialophora: biodiversity and possible role in human infection. Fungal Divers 65:17–45. doi:10.1007/s13225-012-0194-5
Graniti A (1962) Scolecobasidium anellii n. sp., agenti di annerimenti superficiali di stalattiti. G Bot Ital 69:360–365
Hao L, Chen C, Zhang R, Zhu M, Sun G, Gleason ML (2013) A new species of Scolecobasidium associated with the sooty blotch and flyspeck complex on banana from China. Mycol Progress 12:489–495. doi:10.1007/s11557-012-0855-5
Heinrichs G, Hübner I, Schmidt CK, de Hoog GS, Haase G (2013a) Analysis of black fungal biofilms occurring at domestic water taps (I): Compositional analysis using tag-encoded FLX amplicon pyrosequencing. Mycopathologia 175:387–397. doi:10.1007/s11046-013-9618-3
Heinrichs G, Hübner I, Schmidt CK, de Hoog GS, Haase G (2013b) Analysis of black fungal biofilms occurring at domestic water taps (II): Potential routes of entry. Mycopathologia 175:399–412. doi:10.1007/s11046-013-9619-2
Kirilenko TC, All-Achmed MA (1977) Ochroconis tshawytschae (Doty et Slater) comb.nov. Mikrobiol Zh 39:303–306
Lian X, de Hoog GS (2010) Indoor wet cells harbour melanized agents of cutaneous infection. Med Mycol 48:622–628
Machouart M, Samerpitak K, de Hoog GS, Gueidan C (2014) A multigene phylogeny reveals that Ochroconis belongs to the family Sympoventuriaceae (Venturiales, Dothideomycetes). Fungal Divers 65:77–88. doi:10.1007/s13225-013-0252-7
Martin-Sanchez PM, Nováková A, Bastian F, Alabouvette C, Saiz-Jimenez C (2012) Two new species of the genus Ochroconis, O. lascauxensis and O. anomala isolated from black stains in Lascaux Cave, France. Fungal Biol 116:574–589
Matsushima T (1971) Microfungi of the Solomon Islands and Papua-New Guinea. Kobe, pp 50–52
Matsushima T (1975) Icones Microfungorum a Matsushima lectorum. Kobe, pp 126–131
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Proceedings of the Gateway Computing Environments Workshop (GCE), 14 Nov. 2010, New Orleans, LA 1–8
Najafzadeh MJ, Gueidan C, Badali H, Gerrits van den Ende AHG, Xi L, de Hoog GS (2009) Genetic diversity and species delimitation in the opportunistic genus Fonsecaea. Med Mycol 47:17–25
Samerpitak K, Van der Linde E, Choi H-J, Gerrits van den Ende AHG, Machouart M, Gueidan C, de Hoog GS (2014) Taxonomy of Ochroconis, genus including opportunistic pathogens on humans and animals. Fungal Divers 65:89–126. doi:10.1007/s13225-013-0253-6
Satow MM, Attili-Angelis D, de Hoog GS, Angelis DF, Vicente VA (2008) Selective factors involved in oil flotation isolation of black yeast from the environment. Stud Mycol 61:157–163
Sudhadham M, Sihanonth P, Sivichai S, Chaiyarat R, Dorrestein GM, Menken SBJ, de Hoog GS (2008) The neurotropic black yeast Exophiala dermatitidis has possible origin in the tropical rain forest. Stud Mycol 61:137–144
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Wellman AM (1975) A new species of Ochroconis isolated from pelagic tar fragments. Can J Bot 53:1630–1633
Acknowledgments
We would like to thank K. Luijsterburg for his assistance in making fungal photographs.
Author information
Authors and Affiliations
Corresponding author
Additional information
Taxonomic novelties: Ochroconis globalis Samerpitak, Duarte, Attili-Angelis & de Hoog, sp. nov., Ochroconis musae (G.Y. Sun & Lu Hao) Samerpitak & de Hoog, comb. nov.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Samerpitak, K., Duarte, A.P.M., Attili-Angelis, D. et al. A new species of the oligotrophic genus Ochroconis (Sympoventuriaceae). Mycol Progress 14, 6 (2015). https://doi.org/10.1007/s11557-015-1023-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-015-1023-5