Abstract
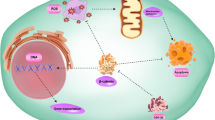
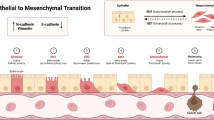
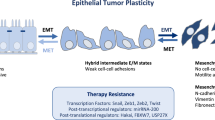
Carcinomas often utilize epithelial-mesenchymal transition (EMT) programs for cancer progression and metastasis. Numerous studies report SNAIL-induced miR200/Zeb feedback circuit as crucial in regulating EMT by placing cancer cells in at least three phenotypic states, viz. epithelial (E), hybrid (h-E/M), mesenchymal (M), along the E-M phenotypic spectrum. However, a coherent molecular-level understanding of how such a tiny circuit controls carcinoma cell entrance into and residence in various states is lacking. Here, we use molecular binding data and mathematical modeling to report that the miR200/Zeb circuit can essentially utilize combinatorial cooperativity to control E-M phenotypic plasticity. We identify minimal combinatorial cooperativities that give rise to E, h-E/M, and M phenotypes. We show that disrupting a specific number of miR200 binding sites on Zeb as well as Zeb binding sites on miR200 can have phenotypic consequences—the circuit can dynamically switch between two (E, M) and three (E, h-E/M, M) phenotypes. Further, we report that in both SNAIL-induced and SNAIL knock-out miR200/Zeb circuits, cooperative transcriptional feedback on Zeb as well as Zeb translation inhibition due to miR200 are essential for the occurrence of intermediate h-E/M phenotype. Finally, we demonstrate that SNAIL can be dispensable for EMT, and in the absence of SNAIL, the transcriptional feedback can control cell state transition from E to h-E/M, to M state. Our results thus highlight molecular-level regulation of EMT in miR200/Zeb circuit and we expect these findings to be crucial to future efforts aiming to prevent EMT-facilitated dissemination of carcinomas.








Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article [and its supplementary information files].
References
Aiello NM, Kang Y (2019) Context-dependent EMT programs in cancer metastasis. J Exp Med 216 (5):1016–1026. https://doi.org/10.1084/JEM.20181827
Bastid J (2012) EMT in carcinoma progression and dissemination: facts, unanswered questions, and clinical considerations. Cancer Metastasis Rev 31 (1–2):277–283. https://doi.org/10.1007/S10555-011-9344-6
Brabletz S, Brabletz T (2010) The ZEB/miR-200 feedback loop-a motor of cellular plasticity in development and cancer? EMBO Rep 11 (9):670–677. https://doi.org/10.1038/embor.2010.117
Brabletz S, Bajdak K, Meidhof S et al (2011) The ZEB1/miR-200 feedback loop controls Notch signalling in cancer cells. EMBO J. https://doi.org/10.1038/emboj.2010.349
Brooks MD, Burness ML, Wicha MS (2015) Therapeutic implications of cellular heterogeneity and plasticity in breast cancer. Cell Stem Cell 17 (3):260–271. https://doi.org/10.1016/J.STEM.2015.08.014
Bu P, Chen KY, Chen JH et al (2013) A microRNA miR-34a-regulated bimodal switch targets notch in colon cancer stem cells. Cell Stem Cell. https://doi.org/10.1016/j.stem.2013.03.002
Burk U, Schubert J, Wellner U et al (2008) A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. https://doi.org/10.1038/embor.2008.74
Chaffer CL, Marjanovic ND, Lee T et al (2013) Poised chromatin at the ZEB1 promoter enables breast cancer cell plasticity and enhances tumorigenicity. Cell 154 (1):61–74. https://doi.org/10.1016/J.CELL.2013.06.005
Charitou T, Bryan K, Lynn DJ (2016) Using biological networks to integrate, visualize and analyze genomics data. Genet Select Evol. https://doi.org/10.1186/s12711-016-0205-1
Fares J, Fares MY, Khachfe HH, Salhab HA, Fares Y (2020) Molecular principles of metastasis: a hallmark of cancer revisited. Signal Transduct Target Ther. https://doi.org/10.1038/s41392-020-0134-x
Fuqua SAW (2021) Metastasis: complexity thwarts precision targeting. Br J Cancer 125 (8):1033–1035. https://doi.org/10.1038/s41416-021-01401-1
Gras B, Jacqueroud L, Wierinckx A et al (2014) Snail family members unequally trigger EMT and thereby differ in their ability to promote the neoplastic transformation of mammary epithelial cells. PLoS ONE. https://doi.org/10.1371/journal.pone.0092254
Guaita S, Puig I, Francí C et al (2002) Snail induction of epithelial to mesenchymal transition in tumor cells is accompanied by MUC1 repression and ZEB1 expression. J Biol Chem 277 (42):39209–39216. https://doi.org/10.1074/jbc.M206400200
Gupta PB, Pastushenko I, Skibinski A, Blanpain C, Kuperwasser C (2019) Phenotypic plasticity: driver of cancer initiation, progression, and therapy resistance. Cell Stem Cell. https://doi.org/10.1016/j.stem.2018.11.011
Hari K, Sabuwala B, Subramani BV et al (2020) Identifying inhibitors of epithelial–mesenchymal plasticity using a network topology-based approach. NPJ Syst Biol Appl. https://doi.org/10.1038/s41540-020-0132-1
Hill L, Browne G, Tulchinsky E (2012) ZEB/miR-200 feedback loop: at the crossroads of signal transduction in cancer. Int J Cancer 132 (4):745–754. https://doi.org/10.1002/ijc.27708
Jolly MK, Boareto M, Huang B et al (2015) Implications of the hybrid epithelial/mesenchymal phenotype in metastasis. Front Oncol. https://doi.org/10.3389/fonc.2015.00155
Jolly MK, Tripathi SC, Jia D et al (2016) Stability of the hybrid epithelial/mesenchymal phenotype. Oncotarget 7 (19):27067–27084
Jolly MK, Preca BT, Tripathi SC et al (2018) Interconnected feedback loops among ESRP1, HAS2, and CD44 regulate epithelial-mesenchymal plasticity in cancer. APL Bioeng. https://doi.org/10.1063/1.5024874
Jolly MK, Mani SA, Levine H (2018) Hybrid epithelial/mesenchymal phenotype (s): the ‘fittest’ for metastasis? Biochim Biophys Acta Rev Cancer 1870 (2):151–157. https://doi.org/10.1016/j.bbcan.2018.07.001
Jolly MK, Somarelli JA, Sheth M et al (2019) Hybrid epithelial/mesenchymal phenotypes promote metastasis and therapy resistance across carcinomas. Pharmacol Ther. https://doi.org/10.1016/j.pharmthera.2018.09.007
Kim NH, Kim HS, Li XY et al (2011) A p53/miRNA-34 axis regulates Snail1-dependent cancer cell epithelial-mesenchymal transition. J Cell Biol 195 (3):417–433. https://doi.org/10.1083/jcb.201103097
Korpal M, Kang Y (2008) The emerging role of miR-200 family of microRNAs in epithelial-mesenchymal transition and cancer metastasis. RNA Biol. https://doi.org/10.4161/rna.5.3.6558
Kröger C, Afeyan A, Mraz J et al (2019) Acquisition of a hybrid E/M state is essential for tumorigenicity of basal breast cancer cells. Proc Natl Acad Sci U S A 116 (15):7353–7362. https://doi.org/10.1073/pnas.1812876116
Kurahara H, Takao S, Maemura K et al (2012) Epithelial-mesenchymal transition and mesenchymal-epithelial transition via regulation of ZEB-1 and ZEB-2 expression in pancreatic cancer. J Surg Oncol. https://doi.org/10.1002/jso.23020
Lambert AW, Weinberg RA (2021) Linking EMT programmes to normal and neoplastic epithelial stem cells. Nat Rev Cancer 21 (5):325–338. https://doi.org/10.1038/s41568-021-00332-6
Lambert AW, Pattabiraman DR, Weinberg RA (2017) Emerging biological principles of metastasis. Cell 168 (4):670–691. https://doi.org/10.1016/J.CELL.2016.11.037
Larsen JE, Nathan V, Osborne JK et al (2016) ZEB1 drives epithelial-to-mesenchymal transition in lung cancer. J Clin Invest. https://doi.org/10.1172/JCI76725
Levine E, Ben JE, Levine H (2007) Target-specific and global effectors in gene regulation by microRNA. Biophys J 93 (11):L52–L54
Li R, Liang J, Ni S, Zhou T, Qing X, Li H, Pei D (2010) A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 7 (1):51–63. https://doi.org/10.1016/j.stem.2010.04.014
Liao TT, Yang MH (2020) Hybrid epithelial/mesenchymal state in cancer metastasis: clinical significance and regulatory mechanisms. Cells. https://doi.org/10.3390/cells9030623
Lu W, Kang Y (2019) Epithelial-mesenchymal plasticity in cancer progression and metastasis. Dev Cell 49 (3):361–374. https://doi.org/10.1016/j.devcel.2019.04.010
Lu M, Jolly MK, Gomoto R, Huang B, Onuchic J, Ben-Jacob E (2013a) Tristability in cancer-associated microRNA-TF chimera toggle switch. J Phys Chem B 117 (42):13164–13174. https://doi.org/10.1021/jp403156m
Lu M, Jolly MK, Levine H, Onuchic JN, Ben-Jacob E (2013b) MicroRNA-based regulation of epithelial-hybrid-mesenchymal fate determination. Proc Natl Acad Sci U S A 110 (45):18144–18149. https://doi.org/10.1073/pnas.1318192110
Mani SA, Guo W, Liao MJ et al (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133 (4):704–715. https://doi.org/10.1016/J.Cell.2008.03.027
Nordick B, Chae-Yeon Park M, Quaranta V, Hong T (2022) Cooperative RNA degradation stabilizes intermediate epithelial-mesenchymal states and supports a phenotypic continuum. iScience. 25 (10):105224. https://doi.org/10.1016/j.isci.2022.105224
Park SM, Gaur AB, Lengyel E, Peter ME (2008) The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. https://doi.org/10.1101/gad.1640608
Pastushenko I, Blanpain C (2019) EMT transition states during tumor progression and metastasis. Trends Cell Biol. https://doi.org/10.1016/j.tcb.2018.12.001
Paul MC, Schneeweis C, Falcomatà C et al (2023) Non-canonical functions of SNAIL drive context-specific cancer progression. Nat Commun 14 (1):1–21. https://doi.org/10.1038/s41467-023-36505-0
Peinado H, Olmeda D, Cano A (2007) Snail, Zeb and bHLH factors in tumour progression: an alliance against the epithelial phenotype? Nat Rev Cancer 7 (6):415–428. https://doi.org/10.1038/nrc2131
Rashid M, Hari K, Thampi J, Santhosh NK, Jolly MK. Network topology metrics explaining enrichment of hybrid epithelial/mesenchymal phenotypes in metastasis. Faeder JR, ed. PLoS Comput Biol. 2022;18 (11):1010687. https://doi.org/10.1371/journal.pcbi.1010687
Rozum J, Albert R (2022) Leveraging network structure in nonlinear control. NPJ Syst Biol Appl. https://doi.org/10.1038/s41540-022-00249-2
Samavarchi-Tehrani P, Golipour A, David L, Sung HK, Beyer TA, Datti A, Wrana JL (2010) Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 7 (1):64–77. https://doi.org/10.1016/j.stem.2010.04.015
Sampson VB, David JM, Puig I et al (2014) Wilms’ tumor protein induces an epithelial-mesenchymal hybrid differentiation state in clear cell renal cell carcinoma. PLoS ONE. https://doi.org/10.1371/journal.pone.0102041
Saxena M, Stephens MA, Pathak H, Rangarajan A (2011) Transcription factors that mediate epithelial–mesenchymal transition lead to multidrug resistance by upregulating ABC transporters. Cell Death Disease. 2 (7):e179–e179. https://doi.org/10.1038/cddis.2011.61
Singh A, Settleman J (2010) EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene 29 (34):4741–4751. https://doi.org/10.1038/onc.2010.215
Subbalakshmi AR, Kundnani D, Biswas K et al (2020) NFATc acts as a non-canonical phenotypic stability factor for a hybrid epithelial/mesenchymal phenotype. Front Oncol. https://doi.org/10.3389/fonc.2020.553342
Thiery JP, Acloque H, Huang RYJ, Nieto MA (2009) Epithelial-mesenchymal transitions in development and disease. Cell 139 (5):871–890. https://doi.org/10.1016/J.CELL.2009.11.007
Tian XJ, Zhang H, Xing J (2013) Coupled reversible and irreversible bistable switches underlying TGFβ-induced epithelial to mesenchymal transition. Biophys J 105 (4):1079–1089. https://doi.org/10.1016/j.bpj.2013.07.011
Title AC, Hong SJ, Pires ND et al (2018) Genetic dissection of the miR-200–Zeb1 axis reveals its importance in tumor differentiation and invasion. Nat Commun. https://doi.org/10.1038/s41467-018-07130-z
Tripathi S, Kessler DA, Levine H, Balázsi G, Albert R (2022) Minimal frustration underlies the usefulness of incomplete regulatory network models in biology. Published Online. https://doi.org/10.1073/pnas
Vega S, Morales AV, Ocaña OH, Valdés F, Fabregat I, Nieto MA (2004) Snail blocks the cell cycle and confers resistance to cell death. Genes Dev 18 (10):1131–1143. https://doi.org/10.1101/GAD.294104
Ye X, Weinberg RA (2015) Epithelial-mesenchymal plasticity: a central regulator of cancer progression. Trends Cell Biol 25 (11):675–686. https://doi.org/10.1016/J.TCB.2015.07.012
Zhang J, Tian XJ, Zhang H et al (2014) TGF-β-induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops. Sci Signal. 7 (345):ra91. https://doi.org/10.1126/SCISIGNAL.2005304/SUPPL_FILE/7_RA91_SM.PDF
Zhang Y, Donaher JL, Das S et al (2022) Genome-wide CRISPR screen identifies PRC2 and KMT2D-COMPASS as regulators of distinct EMT trajectories that contribute differentially to metastasis. Nat Cell Biol 24 (4):554–564. https://doi.org/10.1038/s41556-022-00877-0
Zheng X, Carstens JL, Kim J et al (2015) Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature 527 (7579):525–530. https://doi.org/10.1038/nature16064
Acknowledgements
We benefited from useful discussions with Prof. Mohit Kumar Jolly, Department of Bioengineering, Indian Institute of Science, Bangalore, India. We gratefully acknowledge funding by the Department of Science and Technology (DST), Govt. of India, to MR (grant number: DST/INSPIRE/04/2020/001492).
Funding
The Funding was provided by DST, Govt of India, DST/INSPIRE/04/2020/001492, Mubasher Rashid
Author information
Authors and Affiliations
Contributions
MR: Conceptualization, methodology, simulations, data analysis, supervision, funding acquisition, writing—original draft as well as revision. BMD: Data analysis and simulations. MB: discussions and simulations.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rashid, M., Devi, B.M. & Banerjee, M. Combinatorial Cooperativity in miR200-Zeb Feedback Network can Control Epithelial–Mesenchymal Transition. Bull Math Biol 86, 48 (2024). https://doi.org/10.1007/s11538-024-01277-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11538-024-01277-1