Abstract
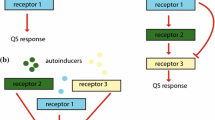
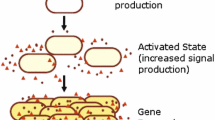
Bacterial quorum sensing (QS) is a form of intercellular communication that relies on the production and detection of diffusive signaling molecules called autoinducers. Such a mechanism allows the bacteria to track their cell density in order to regulate group behavior, such as biofilm formation and bioluminescence. In a number of bacterial QS systems, including V. harveyi, multiple signaling pathways are integrated into a single phosphorylation–dephosphorylation cycle. In this paper, we propose a weight control mechanism, in which QS uses feedback loops to ‘decode’ the integrated signals by actively changing the sensitivity in different pathways. We first use a slow/fast analysis to reduce a single-cell model to a planar dynamical system involving the concentrations of phosphorylated signaling protein LuxU and a small non-coding RNA. In addition to identifying the weight control mechanism, we show that adding a feedback loop can lead to a bistable QS response in certain parameter regimes. We then combine the slow/fast analysis with a contraction mapping theorem in order to reduce a population model to an effective single-cell model, and show how the weight control mechanism allows bacteria to have a finer discrimination of their social and physical environment.











Similar content being viewed by others
References
Bressloff PC (2016) Ultrasensitivity and noise amplification in a model of V. harveyi quorum sensing. Phys Rev E 93:062418
Cornforth DM, Popat R, McNally L, Gurney J, Scott-Phillips DM, Ivens A, Diggle SP, Brown SP (2014) Combinatorial quorum sensing allows bacteria to resolve their social and physical environment. Proc Natl Acad Sci USA 111:4280–4284
Davies DG, Parsek MR, Pearson JP, Iglewski BH, Costerton JW, Greenberg EP (1998) The involvement of cell-to-cell signals in the development of bacterial biofilm. Science 280:295–298
Dunlap PV (1999) Quorum regulation of luminescence in Vibrio fischeri. J Mol Microbiol Biotechnol 1:5–12
Fan G, Bressloff PC (2017) Population model of quorum sensing with multiple parallel pathways. Bull Math Biol 79:2599–2626
Fuqua C, Winans SC, Greenberg EP (1996) Census and consensus in bacterial ecosystems: the LuxR-LuxI family of quorum-sensing transcriptional regulators. Annu Rev Microbiol 50:727–751
Ge H, Qian M (2008) Sensitivity amplification in the phosphorylation-dephosphorylation cycle: nonequilibrium steady states, chemical master equation and temporal cooperativity. J Chem Phys 129:015104
Goldbeter A, Koshland DE (1981) An amplified sensitivity arising from covalent modification in biological systems. Proc Natl Acad Sci USA 78:684–6844
Goryachev AB (2009) Design principles of the bacterial quorum sensing gene networks. WIRE Syst Biol Med 1:45–60
Haseltine EL, Arnold FH (2008) Implications of rewiring bacterial quorum sensing. Appl Environ Microbiol 74(2):437–445
Hense BA, Kuttler C, Muller J, Rothballer M, Hartmann A, Kreft JU (2007) Does efficiency sensing unify diffusion and quorum sensing? Nat Rev Microbiol 5:230–239
Hunter GAM, Guevara Vasquez F, Keener JP (2013) A mathematical model and quantitative comparison of the small RNA circuit in the Vibrio harveyi and Vibrio cholerae quorum sensing systems. Phys Biol 10:046007
Jung SA, Hawver LA, Ng WL (2016) Parallel quorum sensing signaling pathways in Vibrio cholerae. Curr Genet 62(2):255–260
Lenz DDH, Mok KC, Lilley BN, Kulkarni RV, Wingreen NS, Bassler BL (2004) The small RNA chaperone Hfq and multiple small RNAs control quorum sensing in Vibrio harveyi and Vibrio cholerae. Cell 118:69–82
Lohmiller W, Slotine JJE (1998) On contraction analysis for non-linear systems. Automatica 34:683–696
Long T, Tu KC, Wang Y, Mehta P, Ong NP, Bassler BL, Wingreen NS (2009) Quantifying the integration of quorum-sensing signals with single-cell resolution. PLoS Biol 7(3):e1000068
Miyashiro T, Ruby EG (2012) Shedding light on bioluminescence regulation in Vibrio fischeri. Mol Microbiol 84:795–806
Ng WL, Bassler BL (2009) Bacterial quorum-sensing network architectures. Annu Rev Genet 43:197–222
Papenfort K, Bassler BL (2016) Quorum sensing signal-response systems in Gram-negative bacteria. Nat Rev Microbiol 1:577–588
Popat R, Cornforth DM, McNally L, Brown SP (2015) Collective sensing and collective responses in quorum-sensing bacteria. J R Soc Interface 12:20140882
Qian H (2012) Cooperativity in cellular biochemical processes. Annu Rev Biophys 41:179–204
Redfield RJ (2002) Is quorum sensing a side effect of diffusion sensing? Trends Microbiol 10:365–370
Russo G, Slotine JJE (2010) Global convergence of quorum sensing network. Phys Rev E 82:041919
Seger J, Brockman H (1987) What is bet-hedging? In: Harvey PH, Partridge L (eds) Oxford surveys in evolutionary biology, vol 4. Oxford University Press, Cambridge, pp 182–211
Swem LR, Swem DL, Wingreen NS, Bassler BL (2008) Deducing receptor signaling parameters from in vivo analysis: LuxN/AI-1 quorum sensing in Vibrio harveyi. Cell 134:461–473
Teng SW, Schaffer JN, Tu KC, Mehta P, Lu W, Ong NP, Bassler BL, Wingreen NS (2011) Active regulation of receptor ratios controls integration of quorum-sensing signals in Vibrio harveyi. Mol Syst Biol 7:491
Tu KC, Long T, Svenningsen SL, Wingreen NS, Bassler BL (2010) Negative feedback loops involving small regulatory RNAs precisely control the Vibrio harveyi quorum-sensing response. Mol Cell 37:567–579
Wei Y, Ng WL, Cong J, Bassler BL (2012) Ligand and antagonist driven regulation of the Vibrio cholerae quorum-sensing receptor CqsS. Mol Microbiol 83:1095–1108
West SA, Winzer K, Gardner A, Diggle SP (2012) Quorum sensing and the confusion about diffusion. Trends Microbiol 20:586–594
Williams JW, Cui X, Levchenko A, Stevens AM (2008) Robust and sensitive control of a quorum sensing circuit by two interlocked feedback loops. Mol Syst Biol 4:234
Zhu J, Chai Y, Zhong Z, Li S, Winans SC (2003) Agrobacterium bioassay strain for ultrasensitive detection of N-acylhomoserine lactone-type quorum-sensing molecules: detection of autoinducers in Mesorhizobium huakuii. Appl Environ Microbiol 69(11):6949–6953
Acknowledgements
PCB was supported by the National Science Foundation (DMS-1613048). GF was supported by the National Science Foundation (DMS-RTG 1148230).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix A
Appendix A
In this appendix, we show that a sufficient condition for the equilibrium of the single-cell model to be unique is given by Eq. (40). We already showed the uniqueness for the cases of \(u_1>u_2\) and \(u_1=u_2\), in Sect. 2.3. Next, we will work on the case of \(u_1<u_2\). Since the equilibrium satisfies \(f_1(\phi )=f_2(\phi )\), it follows after some algebra that
Note that
Here, because the two sides of (A.1) have to be the same sign, we have the following cases,
Since we are studying the case \(u_1<u_2\), it follows that \(u_1<h(\phi )<u_2\), which implies \(f_3(\phi )>0\).
By some algebra on Eq. (A.1), we get
Recall from Eq. (33) that \(h(\phi )\ge 0\). Since the AI concentration \(u_2\ge 0\), we have \(1-f_3(\phi )>0\). Uniqueness is guaranteed by the monotonicity of the function \(f_4(\phi )\). By taking the derivative with respect to \(\phi \) of \(u_2=f_4(\phi )\), we find
where,
Since \(0<f_3<1\) and \(1+h>0\), one sufficient condition for \(f_4^\prime <0\) to be true is having \(f_3^\prime <0\). Also,
Since \(0\le \phi \le 1\), we require \(h(\phi )-h^\prime (\phi )\phi ^2+\phi h^\prime (\phi )<0\). By direct substitution, we have
Therefore, Eq. (40) is a sufficient condition for the response to be unique.
Rights and permissions
About this article
Cite this article
Fan, G., Bressloff, P.C. Modeling the Role of Feedback in the Adaptive Response of Bacterial Quorum Sensing. Bull Math Biol 81, 1479–1505 (2019). https://doi.org/10.1007/s11538-019-00570-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-019-00570-8