Abstract
Background
DNA methyltransferase inhibitors (DNMTis) improve survival for patients with myelodysplastic syndromes (MDS) and those with acute myeloid leukemia (AML) unable to receive standard cytotoxic chemotherapy and are, accordingly, the backbone of standard-of-care treatment for these conditions. Standard regimens with DNMTIs, decitabine (DEC) or azacitidine (AZA) include daily subcutaneous (s.c.) or intravenous (i.v.) administration for 5–7 consecutive days. Attempts to provide the therapy orally have been limited given rapid clearance of the agents by the enzyme cytidine deaminase (CDA), which is ubiquitous in the gut and liver as part of first-pass metabolism. Recently, cedazuridine (CDZ), an oral inhibitor of CDA, was successfully combined with DEC to approximate the pharmacokinetics of i.v. DEC in patients.
Objective
To determine if an oral dosing strategy might be feasible in the clinic with AZA, we attempted to increase the bioavailability of oral AZA through the use of CDZ, in a murine model.
Methods
Following pharmacokinetic and pharmacodynamic assessment of oral AZA dosed with CDZ in murine and monkey models, we tested this regimen in vivo with a human cell line-derived xenograft transplantation experiment (CDX). Following this we combined the regimen with venetoclax (VEN) to test the efficacy of an all-oral regimen in a patient-derived xenograft (PDX) model.
Results
Parenteral AZA and oral AZA + CDZ exhibited similar pharmacokinetic profiles, and efficacy against human AML cells. Tumor regression was seen with AZA + CDZ in MOLM-13 CDX and PDX models.
Conclusions
We conclude that oral AZA when combined with CDZ achieves successful tumor regression in both CDX and PDX models. Furthermore, the combination of AZA + CDZ with VEN in a PDX model emulated responses seen with VEN + AZA in the clinic, implying a potential all-oral VEN-based therapy opportunity in myeloid diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Oral azacitidine + cedazuridine achieved similar dose-dependent responses to those achieved with intraperitoneal azacitidine. |
The addition of venetoclax to oral azacitidine + cedazuridine resulted in significant decreases in tumor expansion in an acute myeloid leukemia patient-derived xenograft model. |
1 Introduction
Acute myeloid leukemia (AML) and myelodysplastic syndromes (MDS) are related myeloid neoplasms hallmarked by ineffective hematopoiesis and proliferation of malignant clones. Specifically targeting malignant clones while allowing for the preservation of normal hematopoiesis is critical in the design of new therapies. In recent years, next-generation sequencing (NGS) has considerably improved our understanding of the molecular epidemiology and biologic roles of somatic alterations associated with AML and MDS; however, only a few targeted therapies have been successfully developed to address these liabilities.
Thus, the DNA methyltransferase inhibitors (DNMTis) azacitidine (AZA) and decitabine (DEC) have remained the standard first-line disease-modifying therapies across mutational subtypes in MDS, dysplastic chronic myelomonocytic leukemia, and AML for which induction chemotherapy is found to be inappropriate. At low doses, these drugs alter methylation within the transcriptionally active regions of the genome, reduce transcription of pro-growth genetic programs, and ultimately force cells out of the cell cycle. Standard regimens with the DNMTis AZA or DEC include daily subcutaneous (s.c.) or intravenous (i.v.) administration for 5–7 consecutive days [1,2,3,4]. The demands on patients for 5 consecutive days of a parenteral dosing strategy are not trivial. These patients experience pain associated with multiple i.v. punctures, associated risks of infection, bleeding, and thrombosis, and inconveniences associated with central catheters or s.c. infusion ports [5,6,7]. Moreover, when oral versus i.v. treatment strategies have been assessed in other malignancies, patients strongly preferred oral agents as long as efficacy was not significantly compromised [8, 9]. To this end, there has been significant interest in the development of an oral formulation of DNMTi, but attempts to provide the therapy orally have been limited given rapid clearance of the agents by the enzyme cytidine deaminase (CDA), which is ubiquitous in the gut and liver and critical in the function of first-pass metabolism [10].
CC-486 is an oral analog of AZA, and has been in development for over a decade. In phase I, dose-finding studies, the maximum tolerated dosage was 480 mg a day, but patients experienced grade 3–4 diarrhea, despite poor mean relative oral bioavailability [11]. Longer schedules of lower doses of CC-486 led to improved tolerability and area under the curve (AUC), but vastly different pharmacokinetics and methylation changes than seen with standard AZA, and disappointing efficacy as primary treatment [12, 13]. While the use of CC-486 as maintenance therapy post-induction for untreated AML has been encouraging and likely will lead to approval of CC-486 for patients who achieve a complete response (CR) after induction and have less than standard cytarabine consolidation cycles, oral CC-486 is not bioequivalent to i.v./s.c. AZA [14].
Another approach in the development of oral DNMTis is the co-administration of an oral DNMTi with a CDA inhibitor (CDAi) to limit degradation of AZA during first-pass metabolism after oral dosing. Inhibiting CDA with tetrahydrouridine (THU) has been attempted in sickle cell disease, preceding low-dose oral DEC; however, THU must be given 1 h prior to DNMTi, and is unstable in acidic environments [15]. In a recent dose-escalation study (NCT02103478), the simultaneous oral administration of the novel CDAi cedazuridine (CDZ) together with DEC approximated the pharmacokinetics of i.v. DEC in patients [16]. Global levels of demethylation as measured by bisulfate sequencing in long interspersed nuclear element-1 (LINE-1) demethylation assays revealed similar reduction [16]. In a phase III, randomized study, an oral fixed-dose combination (FDC) of DEC + CDZ 35 mg/100 mg (ASTX727) confirmed DEC systemic exposure equivalence with body-surface-area-adjusted i.v. DEC, with the intrapatient oral/i.v. AUC ratio being 98.9% (90% confidence interval 92.7–105.6) and similar efficacy between ASTX727 and i.v. DEC [17].
Similarly, here we show that the combination of oral AZA and CDZ is feasible in the preclinical setting and can be safely pursued in a clinical setting. The AUC achieved with AZA + CDZ was dose dependent, and pharmacokinetics in mice were similar between oral AZA + CDZ and intraperitoneal (i.p.) AZA. Likewise, decreases in LINE-1 methylation with oral AZA + CDZ were comparable to those seen with i.p. AZA. Subsequently, human cell line-derived and primary patient-derived xenograft (PDX) models reveal that AZA + CDZ led to decreases in leukemic burden and improved survival of mice. Furthermore, the addition of venetoclax (VEN) to AZA + CDZ resulted in significant decreases in tumor expansion in an AML PDX, implying a potential all-oral administration of this emerging standard of care.
2 Methods
2.1 Patient Samples
Experiments were conducted on primary patient samples, which were provided by the Vanderbilt-Ingram Cancer Center Hematopoietic Malignancies Repository in accordance with the tenets of the Declaration of Helsinki, and were approved by the Vanderbilt University Medical Center Institutional Review Board (approval number 151710).
2.2 Cell Lines
AML cell lines THP-1 and MV-4-11 were purchased from the American Type Culture Collection (ATCC) (Manassas, VA). The MOLM-13 cell line was purchased from Deutsche Sammelung von Mikroorganismen und Zellkulturen (DSMV) (Braunschweig, Germany).
ATCC and DSMV cell lines are authenticated by short tandem repeat profiling and cytochrome C oxidase gene analysis. Cultured cells were split every 3 days and maintained in exponential growth phase. Cell lines were tested for mycoplasma as per lab standard of practice using the Universal Mycoplasma Detection Kit (ATCC). Cells were used for the experiments presented here within 10–30 passages from thawing. Cell lines were cultured in RPMI (Gibco, Carlsbad, CA) supplemented with 10–20% fetal bovine serum (Gibco) and 100 U/mL penicillin (Gibco) and 100 µg/mL streptomycin (Gibco). Cells were kept at 37 °C in a 5% CO2 incubator.
2.3 Cell Viability Assay
Compounds were diluted in dimethyl sulfoxide (DMSO) (< 0.2%) and dispensed into a 384-well plate using the Echo 555 liquid handler (Labcyte, San Jose, CA). Following the addition of compounds, cells were pipetted into the 384-well plates at a concentration of 3000 cells per well in RPMI media, as noted above, supplemented with 10% FBS and incubated at 37 °C, 5% CO2 in a tissue culture incubator. Plates were incubated for 72 h, and cell viability was measured using the Cell TiterGlo reagent (Promega, Madison, WI). Percentage viability was defined as the relative luminescence units (RLU) of each well divided by the RLU of cells in DMSO control. Dose–response curves and the 50% growth inhibition concentration (GI50) values were determined using non-linear regression of double-log transformed data using GraphPad Prism version 6.0 h.
2.4 Pharmacokinetic Studies
All animal experiments were conducted in accordance with guidelines approved by the Institutional Animal Care and Use Committee (IACUC) at Vanderbilt University Medical Center under protocol M1500038-01. The pharmacokinetics of AZA in NSGS (NOD-scid IL2Rgnull3Tg [hSCF/hGM-CSF/hIL3]) male mice (The Jackson Laboratory, Bar Harbor, ME) (n = 4/group) were assessed in whole blood after dosing with AZA alone via oral (10 mg/kg) or i.p. (2.5 mg/kg) administration, and via oral administration (2.5 mg/kg) in combination with CDZ (1–30 mg/kg). Whole blood samples (10 µL) were collected over up to 24 h in tubes containing 10 µL of 19 mg/mL aqueous ethylenediaminetetraacetic acid (EDTA) and 10 µL of 30 µg/mL aqueous THU for stabilization. The pharmacokinetics of AZA in cynomolgus monkeys (n = 3, male, aged 3–4 years, 3.9–4.9 kg) were assessed in plasma using a crossover design after dosing on each occasion with AZA alone, via s.c. or oral (6.25 mg/kg) administration, and oral administration (6.25 mg/kg) in combination with CDZ (10 mg/kg), with serial blood samples for plasma collected over 24 h into THU-spiked EDTA tubes and centrifuged within 30 min of collection (2000g for 10 min at 5 °C) for plasma. The plasma samples from monkeys and diluted whole blood samples from mice were stored at − 70 °C until bioanalysis with a qualified liquid chromatography tandem mass spectrometry (LC–MS/MS) method.
2.5 DNA Isolation and Pyrosequencing Method
Global DNA methylation was assessed with LINE-1 methylation bisulfite sequencing analysis. For this, DNA was harvested from blood collected in Paxgene tubes. Cells were resuspended in 1X proteinase K digestion buffer with proteinase K and incubated at 50 °C for 30 min. Cell debris was pelleted by centrifugation at 14,000×g for 10 min, and 20 μL was used in the bisulfite conversion reaction using the EZ DNA Methylation-Direct kit (Zymo Research, Irvine, CA). For DNA methylation analysis, 500 ng of extracted genomic DNA was bisulfite treated using the EZ DNA Methylation kit (Zymo Research). Bisulfite-treated DNA was purified according to the manufacturer’s protocol. Polymerase chain reaction (PCR) was performed using 1 μL of bisulfite-treated DNA and 0.2 μM of each primer. One primer was biotin labeled and high performance liquid chromatography (HPLC) purified in order to purify the final PCR product using Sepharose beads. PCR product was bound to Streptavidin Sepharose HP (GE Healthcare Life Sciences, Chicago, IL). The immobilized PCR products were purified, washed, denatured with a 0.2 μM NaOH solution, and rewashed using the Pyrosequencing Vacuum Prep Tool (Qiagen, Hilden, Germany), as per the manufacturer’s protocol. Next, 0.5 μM of sequencing primer was annealed to the purified single-stranded PCR products. 10 μL of the PCR products was sequenced by pyrosequencing on the PSQ96 HS System (Qiagen) following the manufacturer’s instructions. The methylation status of each CpG site was determined individually as an artificial C/T single nucleotide polymorphism (SNP) using QCpG software (Qiagen). The methylation level at each CpG site was calculated as the percentage of the methylated alleles divided by the sum of all methylated and unmethylated alleles. The mean methylation level was calculated using methylation levels of all measured CpG sites within the targeted region of each gene. Each experiment included non-CpG cytosines as internal controls to detect incomplete bisulfite conversion of the input DNA. In addition, a series of unmethylated and methylated DNA were included as controls in each PCR.
2.6 In Vivo Murine Modeling of AML
Female NSGS mice (The Jackson Laboratory), 6–8 weeks old were irradiated with 100 cGy radiation. Twenty-four hours later, mice were transplanted intravenously with cells of interest. In the cell line model experiments, 3 × 103 MOLM-13 cells were transplanted in each irradiated mouse. At day 7 post-transplant, engrafted mice were randomized to receive 2.5 mg/kg i.p. AZA, 2.5 mg/kg oral AZA, 2.5 mg/kg oral AZA + 3 mg/kg CDZ, or CDZ alone at 30 mg/kg (to test for off-target toxicity of CDZ). Oral AZA and CDZ were administered via oral gavage daily for 7 consecutive days; i.p. therapy was similarly given for 7 consecutive days. For the PDX model, 2 × 106 mononucleated cells from primary patient samples were transplanted via tail vein injections.
Prior to treatment, peripheral micro-chimerism was documented at week 1 in the cell line model. For the AML PDX model, peripheral chimerism was established by 2 weeks. Mice showing no peripheral micro-chimerism by 1 week in the cell line model or 4 weeks in the AML PDX model were removed from the study. Chimeric mice were randomized pre-treatment. Upon establishing micro-chimerism, and after 35 days, mice were treated with either AZA (Selleckchem, Houston, TX) by daily gavage or i.p. injection, CDZ by gavage, or VEN (Chemietek, Indianapolis, IN) by gavage. VEN was dissolved in polyethylene glycol (PEG) and ethanol and diluted with Phosal 50 PG for gavage dosing. CDZ was dissolved in 5% methylcellulose. Engrafted mice received 2.5 mg/kg i.p. AZA, 2.5 mg/kg oral AZA, 2.5 mg/kg oral AZA + 3 mg/kg CDZ, or CDZ alone as vehicle at 30 mg/kg. Oral drugs were administered via oral gavage daily for 7 consecutive days; i.p. therapy was similarly given for 7 consecutive days. Peripheral blood was assessed weekly for human chimerism. Spleen/body ratio was calculated as organ weight (gram) per gram of body weight.
2.7 Flow Cytometry
For flow cytometry, red blood cells were lysed with EL Buffer on ice (Qiagen), and the remaining cells were washed and resuspended in 1 × phospahte-buffered saline (PBS) with 1% bovine serum albumin (BSA) and stained for 15 min with the following antibodies: human CD45-APC (clone 2D1), human CD33-PE-Cy7 (clone P67.6), murine CD45-PE (clone 30-F11), and DAPI (all from Biolegend, San Diego, CA).
2.8 Immunohistochemistry
Tissues were fixed in 4% paraformaldehyde for 48 h and stored in 70% ethanol before being embedded in paraffin and sectioned at 5 µm. The bone tissue was decalcified prior to being embedded in paraffin. Sections were de-waxed in xylene and rehydrated in successive ethanol baths. Standard Mayer’s hematoxylin and eosin (H&E) staining was performed. Antigen retrieval using a standard pH 6 sodium citrate buffer (BioGenex, Fremont, CA) was performed, and sections were stained with Monoclonal Mouse Anti-Human CD45 (Dako, Santa Clara, CA, M0701, dilution 1:200) using M.O.M. Kit (Vector, Burlingame, CA).
2.9 Measurement of In Vivo Hypomethylation
Global DNA methylation was assessed in murine blood at baseline and 8, 19, and 29 days after treatment using the LINE-1 methylation bisulfite sequencing assay as previously described [18].
2.10 Statistics
Unless otherwise noted, data were summarized using the mean (± standard deviation [SD]). Per group sample sizes are presented in figures and results reported from two separate experiments, unless stated otherwise. To avoid normality assumptions, pairwise group comparisons were made using the non-parametric Mann–Whitney U test. The non-parametric Spearman correlation was used to assess pairwise variable associations. Data were analyzed using GraphPad Prism 6.0 for Windows (GraphPad Software, www.graphpad.com) and R (R Core Team 2017; R: a language and environment for statistical computing; R Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org/). Significance was listed as *p < 0.05, **p < 0.01, and ***p < 0.001.
3 Results
3.1 CDZ Does Not Enhance Effects of AZA In Vitro
We measured GI50 in AML cell lines treated with vehicle (DMSO), AZA, and AZA/CDZ combinations. After 72 h of direct treatment of cells, no differences were noted in viability of cell lines between AZA and AZA + CDZ in vitro, as expected. The GI50 in MV-4-11 (7.247 µM), MOLM-13 (0.815 µM), and THP-1 (7.217 µM) cells remained unchanged with the addition of CDZ (Fig. 1). Although cancer cell lines produce CDA, total levels are negligible in comparison to that of the gut and liver. This result led us to more directly pursue an in vivo approach to testing the combination of oral AZA and CDZ.
3.2 Pharmacokinetics of Oral AZA + CDZ Suggests On-Target AUC for AZA
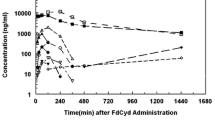
To compare the bioavailability of AZA when dosed orally with CDZ or with traditional parenteral use, we conducted pharmacokinetic studies in both primate and murine models. In male cynomolgus monkeys, dosing with oral AZA alone at 6.25 mg/kg (monkey equivalent dose converted from clinical AZA dose of 75 mg/m2) resulted in mean (SD) systemic AUC last measurable concentration (AUClast) of 18.9 ± 10.1 ng∙h/mL, compared to 369 ± 42 after the same dose was administered s.c. (Fig. 2a). However, when the same oral dose (AZA 6.25 mg/kg) was co-administered with 10 mg/kg CDZ, AZA exposures increased 1634%, with AUClast of 309 ± 268 ng∙h/mL, a level comparable to that seen with s.c.
a Pharmacokinetic (PK) profiling reveals increased bioavailability and similar AUC between parenteral AZA and oral AZA + CDZ, but decreased availability with oral AZA alone in cynomolgus monkeys. b Murine PK studies reveal CDZ dose-dependent increases in AUC of oral AZA and in comparison to dosing of standard i.p. AZA. c LINE-1 demethylation induced by oral AZA only approaches methylation reduction induced by parenteral AZA when combined with CDZ. AZA azacitidine, AUC area under the curve, CDZ cedazuridine, i.p. intraperitoneal, LINE-1 long interspersed nuclear element-1, s.c. subcutaneous
In male NSGS mice, dose-normalized systemic exposures (AUC curve to infinite time (AUC0–inf) [ng∙h/mL] divided by dose [mg/kg]) were 26 and 647 after a single 10-mg/kg oral and 2.5-mg/kg i.p. AZA dose. When given orally at 2.5 mg/kg in combination with CDZ at increasing doses from 1 to 30 mg/kg, AZA dose-normalized AUC increased dose-dependently to 224–364, an 8.6- to 14-fold increase over oral AZA alone (Fig. 2b). These data show that the addition of CDZ to oral AZA leads to pharmacokinetics comparable to those of parenteral dosing, in both primate and murine models.
3.3 Pharmacodynamic Mimicry of AZA by Oral AZA + CDZ via LINE-1 Demethylation
To verify that oral distribution of AZA did not perturb its ability to affect global DNA methylation patterns, we tested the blood from mice treated with both i.p. and oral regimens of AZA and CDZ for 7 consecutive days with a LINE-1 assay measuring levels of hypomethylated residues both pre and post treatment [18]. Oral AZA + CDZ resulted in similar demethylation as i.p. AZA, whereas oral AZA alone was unable to replicate the DNA demethylation induced by i.p. AZA in the LINE-1 assay (Fig. 2c). The transient hypomethylating effect induced by oral AZA alone quickly receded.
3.4 Oral AZA + CDZ is Effective in an In Vivo Human Cell Line-Derived Xenograft Model
Following pharmacokinetic and pharmacodynamic (PK/PD) assessment of oral AZA dosed with CDZ, we used this regimen in an in vivo human cell line-derived xenograft transplantation experiment (CDX). To this end, we studied the effects of CDZ, AZA, and the AZA + CDZ combination on MOLM-13 AML cells in NSGS mice. During treatment, the kinetics of the MOLM-13 human cell line expansion were defined by detection of human CD45+ cells in the blood as detected by flow cytometry. At approximately 3 weeks after transplant, mice treated only with CDZ became moribund, and all experimental groups were sacrificed for analysis of chimerism.
In the CDX model, as expected, i.p. AZA significantly decreased leukemic expansion, exhibiting an average chimerism of 7.86% ± 9.9 and 0.43% ± 0.65 in the bone marrow and spleen, respectively, whereas CDZ alone has no effect on leukemic expansion in the CDX model, exhibiting an average chimerism of 38.32% ± 17.7 and 27.28% ± 12.8 in the marrow and spleen, respectively (Fig. 3a). Likewise, single-agent oral AZA alone failed to decrease AML expansion in both the bone marrow and spleen of treated mice, exhibiting an average chimerism of 41.02% ± 14.6 and 13.9% ± 11. 3, respectively; however, the addition of CDZ to oral AZA led to significant decreases in AML cells in both bone marrow and spleen, showing scant CD45+ cells (1.49% ± 2.66 and 0.09% ± 0.13). Differences between the oral AZA + CDZ group and traditional i.p. AZA dosing were not statistically significant in either bone marrow or splenic tissue (bone marrow: i.p. AZA 7.86 ± 9.9 vs. oral AZA + CDZ 1.49 ± 2.66, p = 0.258; spleen: 0.43 ± 0.65 vs. 0.09 ± 0.13). Immunohistochemical staining for human CD45 also revealed decreased expansion of AML in both bone marrow and spleen of i.p. AZA-treated and oral AZA + CDZ-treated mice (Fig. 3b). In a second MOLM-13 cell transplant experiment, Kaplan–Meier survival analysis revealed that both i.p. AZA and oral AZA + CDZ led to a statistically significant increase in lifespan, with no significant difference in survival between i.p. AZA-treated and oral AZA + CDZ-treated mice (Fig. 3c).
Oral AZA + CDZ leads to reduction of leukemic expansion in a cell line-derived xenograft transplantation. a Spleen and bone marrow were evaluated for chimerism analysis, and a non-parametric, unpaired, two-tailed t test was used to calculate significance (mean ± SD). b Immunohistochemistry of long bones and spleen stained with monoclonal antibody for hCD45 reveal AML cells left within the bone marrow and spleen of experimental mice (× 40). c Kaplan–Meier analysis reveals the significantly extended lifespan of mice treated with either i.p. AZA or oral AZA + CDZ (vehicle vs. i.p. AZA, p < 0.0001 and vehicle vs. AZA + CDZ, p < 0.001), with no significant difference between the two treatments. Data shown from 2.5 mg/kg i.p. AZA (n = 9), 2.5 mg/kg oral AZA (n = 5), 2.5 mg/kg oral AZA + 3 mg/kg CDZ (n = 8), and 30 mg/kg of CDZ alone (n = 7) (vehicle). d Standard hematoxylin and eosin staining of both bone marrow in treated mice reveal maintenance of architecture despite therapy with AZA (× 20, × 40). AML acute myeloid leukemia, AZA azacitidine, CDZ cedazuridine, i.p. intraperitoneal, n.s. not significant, SD standard deviation. **p < 0.01, ***p < 0.001
Throughout the duration of the experiments, the mice in the treatment groups had no significant weight loss, with no group losing more than 20% in body weight (data not shown). After tissues were harvested from experimental mice, bone marrows stained with H&E were reviewed for any toxicity-induced changes in architecture. To determine if drug dosing suppressed normal mouse bone marrow, the marrow within long bones was evaluated by a veterinary pathologist, and no significant hypocellularity, dysplasia, or other signs of drug toxicity were seen in treated mice (Fig. 3d), suggesting that AZA preferentially affected transplanted MOLM-13 AML cells at the doses given. Likewise, no unforeseen effects of CDZ on normal murine bone marrow were noted.
3.5 Oral AZA + CDZ Show Efficacy in a Patient-Derived Xenograft Transplantation Model
After validating the safety and efficacy of oral AZA in a CDX model of AML, we sought to further validate this combination in a primary PDX model of AML. During treatment, the kinetics of patient AML cell expansion was defined by detection of human AML cells in the blood as detected by flow cytometry. At approximately 7 weeks after transplant, mice treated with CDZ or oral AZA alone became moribund, and all experimental groups were sacrificed for analysis of chimerism. Similar to the CDX model, in the PDX model, i.p. AZA decreased leukemic expansion relative to vehicle, having an average chimerism of 62.85% ± 14.1 in the bone marrow and 70.65% ± 2.95 in the spleen, whereas CDZ did not affect disease burden in the bone marrow or spleen, displaying an average chimerism of 90.8% ± 2.9 and 91.46% ± 4.1, respectively (Fig. 4a). More importantly, while oral AZA alone was unable to significantly decrease AML expansion, exhibiting an average chimerism of 76.88% ± 10.98 and 89.34% ± 4.4 in the bone marrow and the spleen, respectively, the addition of CDZ to oral AZA led to significant decreases in AML, displaying an average chimerism of 54.77% ± 12.7 and 58.57% ± 6.24 in bone marrow and spleen, respectively. Importantly, the addition of CDZ to oral AZA showed no significant differences in activity against AML compared to traditional AZA i.p. dosing in bone marrow, but was more effective in the leukemic proliferation in the spleen (Fig. 4a). As VEN + DNMTi therapy has emerged as a new standard of care in some cases of AML [19], we also tested a cohort of mice with VEN in addition to oral AZA + CDZ versus VEN treatment alone and saw significant decreases of tumor burden in both the bone marrow (VEN alone 83.33% ± 7.59 vs. oral AZA + CDZ + VEN 7.06% ± 3.60) and spleen (VEN alone 91.34% ± 3.88 vs. oral AZA + CDZ + VEN 32.32% ± 12.77) of treatment mice. Mice within the oral AZA + CDZ + VEN treatment groups reflected decreases in tumor burden, along with those of i.p. and oral AZA + CDZ treatments, while exhibiting minimal weight changes (Fig. 4b).
Oral AZA + CDZ exhibit preliminary safety and efficacy in a primary AML PDX model. a NSGS mice were engrafted and treated with 2.5 mg/kg i.p. AZA (n = 4), 2.5 mg/kg oral AZA (n = 5), 3 mg/kg CDZ + 2.5 mg/kg oral AZA (n = 5), 20 mg/kg VEN (n = 5), 3 mg/kg CDZ + 2.5 mg/kg oral AZA + VEN (n = 4), or CDZ alone at 30 mg/kg (n = 5) (vehicle), and experimental groups were sacrificed for analysis of chimerism. Data shown as mean ± SEM. b Mouse body weights of experimental mice per group per time point shown as mean ± SEM. AML acute myeloid leukemia, AZA azacitidine, CDZ cedazuridine, i.p. intraperitoneal, n.s. not significant, PDX patient-derived xenograft, SEM standard error of the mean, VEN venetoclax. *p < 0.05, **p < 0.01, ***p < 0.001
4 Discussion
Consistent with previous preclinical and clinical studies with DEC + CDZ (ASTX727) [16], oral AZA approximated bioavailability in mice and monkeys and had comparable pharmacodynamic and anti-tumor activity to i.p. AZA when given with the CDAi CDZ. Pharmacokinetic evaluation in male NSGS mice and cynomolgus monkeys demonstrated proof of concept for achieving higher AZA systemic exposures after oral co-administration with CDZ, and in monkeys, oral combination matched the exposures of AZA after the same parenteral dose. In both species, relative bioavailability of oral AZA alone is very low, but was markedly enhanced (860–1400% in mice and 1630% in monkeys) when dosed with CDZ. These data reveal higher exposures at relatively low oral AZA doses when given with CDZ, avoiding the need to overcome CDA with high doses of single-agent AZA alone—a strategy originally pursued in the development of CC-486. With CC-486, doses were escalated in attempts to match PK/PD properties of AZA i.v./s.c. at approved doses, but this goal was not met and CC-486 led to considerable toxicity at higher doses [11]. Later attempts to administer a variety of dosing regimens was successful and revealed that CC-486 is active in patients with myeloid neoplasms [11, 13, 20, 21]. However, in the alternative dosing regimens, effects on DNA methylation were very different to those with standard AZA i.v./s.c. While dosing of CC-486 to overcome CDA activity and to achieve PK/PD equivalency was not successful due to dose-dependent off-target non-hematologic toxicities [11], the use of the drug in a maintenance setting has been explored. The QUAZAR AML-001 study was a randomized controlled trial of CC-486 versus placebo for patients older than 55 years of age who reached CR after induction cytotoxic chemotherapy, but were within 90 days of achieving CR. Given this 90-day restriction (and later amended to 120 days), the average number of consolidation cycles was only 1.0. While overall survival was no different between the arms, the median overall survival (mOS) was superior for the patients treated with CC-486 (hazard ratio 0.69). Thus, QUAZAR provides the first clear maintenance use for DNMTi in AML in the post-cytotoxic therapy setting. The role of CC-486 in MDS or otherwise in AML (e.g., in de novo disease or relapsed/refractory setting) is yet to be explored.
VEN, an oral selective inhibitor of BCL2, in combination with parenteral DNMTi or low-dose cytarabine (LDAC) for older patients with AML led to striking improvements in remission rates. The approval of VEN when given together with DNMTi or LDAC in AML is based on phase Ib studies which revealed CRs/CR with incomplete count recovery (CRi) in 50–70% of patients over 65 years of age who were deemed to not be candidates for standard cytotoxic induction chemotherapy [19, 22, 23]. The historical mOS for AML in patients over 65 years was measured in months, so VEN-based therapy is quickly becoming part of a changing paradigm in the treatment of AML. To this end, as proof of principle, we tested the all-oral VEN + AZA + CDZ combination in an AML PDX model, and found it tolerable and effective in eliminating the AML. These findings are consistent with the ability to induce meaningful remissions in AML with all-oral VEN-based therapies, and as VEN + DNMTi therapy is evolving to the standard of care, preclinical signals of activity with oral analogs is necessary prior to clinical trials testing this combination.
AZA and DEC are cornerstones in the treatment of myeloid malignancies, but the daily parenteral treatment in the cancer clinic utilizes considerable resources and places potential undue burden on the patients who are committed to this therapy until toxicity or progression of disease. Oral analogs are critical advances, and learning how to best use these compounds is a priority. ASTX727 (DEC + CDZ) has been shown to be largely equivalent to DEC in randomized bioequivalence studies, and CC-486 (oral AZA) improves mOS if used for maintenance after cytotoxic chemotherapy-induced CR in older patients. These advances should be woven into standard practice, and the capacity to emulate AZA PK/PD and clinical efficacy in MDS and AML can be tested with AZA + CDZ. This is the first preclinical evaluation of oral AZA with CDZ, and these preliminary data provide rationale for the development of oral AZA + CDZ therapy as a FDC (ASTX030) for myeloid disease.
References
Lyons RM, Cosgriff TM, Modi SS, Gersh RH, Hainsworth JD, Cohn AL, et al. Hematologic response to three alternative dosing schedules of azacitidine in patients with myelodysplastic syndromes. J Clin Oncol. 2009;27(11):1850–6. https://doi.org/10.1200/JCO.2008.17.1058.
Fenaux P, Mufti GJ, Hellstrom-Lindberg E, Santini V, Finelli C, Giagounidis A, et al. Efficacy of azacitidine compared with that of conventional care regimens in the treatment of higher-risk myelodysplastic syndromes: a randomised, open-label, phase III study. Lancet Oncol. 2009;10(3):223–32. https://doi.org/10.1016/S1470-2045(09)70003-8.
Fenaux P, Mufti GJ, Hellstrom-Lindberg E, Santini V, Gattermann N, Germing U, et al. Azacitidine prolongs overall survival compared with conventional care regimens in elderly patients with low bone marrow blast count acute myeloid leukemia. J Clin Oncol. 2010;28(4):562–9. https://doi.org/10.1200/JCO.2009.23.8329.
Silverman LR, Demakos EP, Peterson BL, Kornblith AB, Holland JC, Odchimar-Reissig R, et al. Randomized controlled trial of azacitidine in patients with the myelodysplastic syndrome: a study of the cancer and leukemia group B. J Clin Oncol. 2002;20(10):2429–40. https://doi.org/10.1200/JCO.2002.04.117.
Groeger JS, Lucas AB, Thaler HT, Friedlander-Klar H, Brown AE, Kiehn TE, et al. Infectious morbidity associated with long-term use of venous access devices in patients with cancer. Ann Intern Med. 1993;119(12):1168–74. https://doi.org/10.7326/0003-4819-119-12-199312150-00003.
Kogan AJ, Dunn JD. Myelodysplastic syndromes: health care management considerations. Manag Care. 2009;18(11 Suppl 9):25–8.
Nabhan C, Radhakrishnan A. Aplastic anemia surfacing after treatment of acute promyelocytic leukemia: the Dameshek riddle. Clin Adv Hematol Oncol. 2009;7(10):672–4.
Ishitobi M, Shibuya K, Komoike Y, Koyama H, Inaji H. Preferences for oral versus intravenous adjuvant chemotherapy among early breast cancer patients. Patient Prefer Adherence. 2013;7:1201–6. https://doi.org/10.2147/PPA.S52687.
Liu G, Franssen E, Fitch MI, Warner E. Patient preferences for oral versus intravenous palliative chemotherapy. J Clin Oncol. 1997;15(1):110–5. https://doi.org/10.1200/JCO.1997.15.1.110.
Mahfouz RZ, Jankowska A, Ebrahem Q, Gu X, Visconte V, Tabarroki A, et al. Increased CDA expression/activity in males contributes to decreased cytidine analog half-life and likely contributes to worse outcomes with 5-azacytidine or decitabine therapy. Clin Cancer Res. 2013;19(4):938–48. https://doi.org/10.1158/1078-0432.CCR-12-1722.
Garcia-Manero G, Gore SD, Cogle C, Ward R, Shi T, Macbeth KJ, et al. Phase I study of oral azacitidine in myelodysplastic syndromes, chronic myelomonocytic leukemia, and acute myeloid leukemia. J Clin Oncol. 2011;29(18):2521–7. https://doi.org/10.1200/JCO.2010.34.4226.
Savona MR, Kolibaba K, Conkling P, Kingsley EC, Becerra C, Morris JC, et al. Extended dosing with CC-486 (oral azacitidine) in patients with myeloid malignancies. Am J Hematol. 2018;93(10):1199–206. https://doi.org/10.1002/ajh.25216.
Garcia-Manero G, Scott BL, Cogle CR, Boyd TE, Kambhampati S, Hetzer J, et al. CC-486 (oral azacitidine) in patients with myelodysplastic syndromes with pretreatment thrombocytopenia. Leuk Res. 2018;72:79–85. https://doi.org/10.1016/j.leukres.2018.08.001.
de Lima M, Oran B, Champlin RE, Papadopoulos EB, Giralt SA, Scott BL, et al. CC-486 maintenance after stem cell transplantation in patients with acute myeloid leukemia or myelodysplastic syndromes. Biol Blood Marrow Transplant. 2018;24(10):2017–24. https://doi.org/10.1016/j.bbmt.2018.06.016.
Molokie R, Lavelle D, Gowhari M, Pacini M, Krauz L, Hassan J, et al. Oral tetrahydrouridine and decitabine for non-cytotoxic epigenetic gene regulation in sickle cell disease: a randomized phase 1 study. PLoS Med. 2017;14(9):e1002382. https://doi.org/10.1371/journal.pmed.1002382.
Savona MR, Odenike O, Amrein PC, Steensma DP, DeZern AE, Michaelis LC, et al. An oral fixed-dose combination of decitabine and cedazuridine in myelodysplastic syndromes: a multicentre, open-label, dose-escalation, phase 1 study. Lancet Haematol. 2019;6(4):e194–203. https://doi.org/10.1016/S2352-3026(19)30030-4.
Garcia-Manero G, McCloskey J, Griffiths E, Yee K, et al. Pharmacokinetic exposure equivalence and preliminary efficacy and safety from a randomized cross over phase 3 study (ASCERTAIN study) of an oral hypomethylating agent ASTX727 (cedazuridine/decitabine) compared to IV decitabine. Oral Presented at: American Society of Hematology Annual Meeting. 2019; December 9, 2019.
Yang AS, Doshi KD, Choi SW, Mason JB, Mannari RK, Gharybian V, et al. DNA methylation changes after 5-aza-2′-deoxycytidine therapy in patients with leukemia. Cancer Res. 2006;66(10):5495–503. https://doi.org/10.1158/0008-5472.CAN-05-2385.
DiNardo CD, Pratz KW, Letai A, Jonas BA, Wei AH, Thirman M, et al. Safety and preliminary efficacy of venetoclax with decitabine or azacitidine in elderly patients with previously untreated acute myeloid leukaemia: a non-randomised, open-label, phase 1b study. Lancet Oncol. 2018;19(2):216–28. https://doi.org/10.1016/S1470-2045(18)30010-X.
Babiker HM, Milhem M, Aisner J, Edenfield W, Shepard D, Savona M, et al. Evaluation of the bioequivalence and food effect on the bioavailability of CC-486 (oral azacitidine) tablets in adult patients with cancer. Cancer Chemother Pharmacol. 2020;85(3):621–6. https://doi.org/10.1007/s00280-020-04037-9.
Laille E, Shi T, Garcia-Manero G, Cogle CR, Gore SD, Hetzer J, et al. Pharmacokinetics and pharmacodynamics with extended dosing of CC-486 in patients with hematologic malignancies. PLoS One. 2015;10(8):e0135520. https://doi.org/10.1371/journal.pone.0135520.
DiNardo CD, Pratz K, Pullarkat V, Jonas BA, Arellano M, Becker PS, et al. Venetoclax combined with decitabine or azacitidine in treatment-naive, elderly patients with acute myeloid leukemia. Blood. 2019;133(1):7–17. https://doi.org/10.1182/blood-2018-08-868752.
Wei AH, Strickland SA Jr, Hou JZ, Fiedler W, Lin TL, Walter RB, et al. Venetoclax combined with low-dose cytarabine for previously untreated patients with acute myeloid leukemia: results from a phase Ib/II study. J Clin Oncol. 2019;37(15):1277–84. https://doi.org/10.1200/JCO.18.01600.
Acknowledgements
The Vanderbilt-Ingram Cancer Center (VICC) Hematopoietic Malignancies Tissue Repository, the Vanderbilt University Medical Center Translational Pathology Shared Resource, and other VICC shared Core Services were critical in completion of this work.
Author information
Authors and Affiliations
Contributions
Conception and design: HR, AO, HK, MA, and MS. Development of methodology: HR, AO, and MA. Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): HR, AO, AG, LF, and MA. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, and computational analysis): HR, AO, AG, LF, MA, KB, HK, MA, and MS. Writing, review, and/or revision of the manuscript: HR, AO, HK, MA, and MS. Administrative, technical, or material support (i.e., reporting or organizing data and constructing database): HR, AO, AG, LF, and MA. Study supervision: AO, HK, MA, and MS.
Corresponding author
Ethics declarations
Funding
This study, including open-access publication costs, were supported by Astex Pharmaceuticals and the Biff Rittenberg Foundation. In addition, the Vanderbilt-Ingram Cancer Center (VICC) Hematopoietic Malignancies Tissue Repository, the Vanderbilt University Medical Center Translational Pathology Shared Resource, and other VICC shared Core Services were critical in completion of this work. This project has been funded in part with Federal funds from the National Cancer Institute. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government. The Vanderbilt-Ingram Cancer Center is supported by an NIH P30 CA068485-19. The REDCap database tool is supported by Grant UL1 TR000445 from NCATS/NIH. MS is a Leukemia and Lymphoma Society Clinical Scholar.
Conflict of interest
Aram Oganesian, Harold Keer, and Mohammad Azab are employees of Astex Pharmaceuticals. Michael R. Savona has equity ownership in Karyopharm; receives research funding from Astex, Incyte, Millennium, Sunesis, and TG Therapeutics; serves on consultancy/advisory board/monitoring committees for Abbvie, BMS, Celgene, Incyte, Karyopharm, Millennium, Sierra Oncology, Ryvu, and TG Therapeutics; and has patents and royalties with Boehringer-Ingelheim. Haley E. Ramsey, Agnieszka E. Gorska, Londa Fuller, Maria Arrate, and Kelli Boyd declare that they have no conflicts of interest that might be relevant to the contents of this manuscript.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Ramsey, H.E., Oganesian, A., Gorska, A.E. et al. Oral Azacitidine and Cedazuridine Approximate Parenteral Azacitidine Efficacy in Murine Model. Targ Oncol 15, 231–240 (2020). https://doi.org/10.1007/s11523-020-00709-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-020-00709-x