Abstract
Background
Small antral follicles as the final reserve of folliculogenesis, are existed throughout the reproductive life span of sheep. However, the ovarian cycles of ewe cease in the anestrus phase. This study in thus was aimed to elucidate the ovarian small antral follicles transcriptome in the ewe’s anestrus phase.
Methods
Granulosa cells of small antral follicles (#3 mm) were collected from ovaries of anestrous ewes under long days of summer in the non-breeding season as anestrus phase. Transcriptome profiling of these granulosa cells were obtained using the RNA-Seq technology. An integrative analysis was utilized to identify key regulatory genes whose may have potential impacts on intra-ovarian molecular activities.
Results
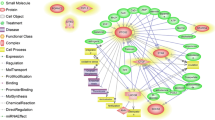
Globally, 14506 genes were expressed whose higher expressions were belonged to genes that encoded ribosomal proteins. Top significant terms of gene ontology were pertained to protein translational processes. Apart of this, most of highly significant terms were also relevant to apoptotic process through extracellular vesicles, including apoptotic bodies and exosomes. Regarding to node effect property, UBA52 (ubiquitin A-52 residue ribosomal protein fusion product 1) and RPS5 (ribosomal proteins S5) contained in highest out-degree and in-degree, respectively.
Conclusion
Our data suggest that ribosomal mRNA/proteins could make the granulosa cells undergo a lot of changes from the point of view that ovarian activities are ceased in the anestrus phase.
Similar content being viewed by others
References
Albert R (2005). Scale-free networks in cell biology. J Cell Sci, 118(21): 4947–4957
Bader G D, Hogue C W (2003). An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics, 4(1): 2
Bahrami A, Miraie-Ashtiani S R, Sadeghi M, Najafi A (2017). miRNAmRNA Network involved in folliculogenesis interactome: Systems biology approach. Reproduction, 154(1): 51–65
Barnard G F, Mori M, Staniunas R J, Begum N A, Bao S, Puder M, Cobb J, Redman K L, Steele G D Jr, Chen L B (1995). Ubiquitin fusion proteins are overexpressed in colon cancer but not in gastric cancer. Biochim Biophys Acta, 1272(3): 147–153
Bartlewski P M, Baby T E, Giffin J L (2011). Reproductive cycles in sheep. Anim Reprod Sci, 124(3–4): 259–268
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman W H, Pagès F, Trajanoski Z, Galon J (2009). ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics, 15(8): 1091–1093
Bonnet A, Cabau C, Bouchez O, Sarry J, Marsaud N, Foissac S, Woloszyn F, Mulsant P, Mandon-Pepin B (2013). An overview of gene expression dynamics during early ovarian folliculogenesis: specificity of follicular compartments and bi-directional dialog. BMC Genomics, 14(1): 904
Bonnet A, Lê Cao K A, Sancristobal M, Benne F, Robert-Granié C, Law-So G, Fabre S, Besse P, De Billy E, Quesnel H, Hatey F, Tosser-Klopp G (2008). In vivo gene expression in granulosa cells during pig terminal follicular development. Reproduction, 136(2): 211–224
Chen F W, Ioannou Y A (1999). Ribosomal proteins in cell proliferation and apoptosis. Int Rev Immunol, 18(5-6): 429–448
Cocucci E, Racchetti G, Meldolesi J (2009). Shedding microvesicles: artefacts no more. Trends Cell Biol, 19(2): 43–51
Cohen B D, Bariteau J T, Magenis L M, Dias J A (2003). Regulation of follitropin receptor cell surface residency by the ubiquitin-proteasome pathway. Endocrinology, 144(10): 4393–4402
Copois V, Bibeau F, Bascoul-Mollevi C, Salvetat N, Chalbos P, Bareil C, Candeil L, Fraslon C, Conseiller E, Granci V, Mazière P, Kramar A, Ychou M, Pau B, Martineau P, Molina F, Del Rio M (2007). Impact of RNA degradation on gene expression profiles: assessment of different methods to reliably determine RNA quality. J Biotechnol, 127(4): 549–559
Cuiling L, Wei Y, Zhaoyuan H, Yixun L (2005). Granulosa cell proliferation differentiation and its role in follicular development. Chin Sci Bull, 50(23): 2665–2671
Di R, He J, Song S, Tian D, Liu Q, Liang X, Ma Q, Sun M,Wang J, Zhao W, Cao G, Wang J, Yang Z, Ge Y, Chu M (2014). Characterization and comparative profiling of ovarian microRNAs during ovine anestrus and the breeding season. BMC Genomics, 15(1): 899
Douville G, Sirard M A (2014). Changes in granulosa cells gene expression associated with growth, plateau and atretic phases in medium bovine follicles. J Ovarian Res, 7(1): 50
Edgar J R (2016). Q&A: What are exosomes, exactly? BMC Biol, 14(1): 46
Fortune J E (1994). Ovarian follicular growth and development in mammals. Biol Reprod, 50(2): 225–232
Goodman R L, Inskeep E K (2015). Control of the ovarian cycle of the sheep. In: Plant T M, Zeleznik A J, editors. Knobil and Neill’s Physiology of Reproduction 4. San Diego, CA, USA: Elsevier Inc, pp. 1259–1305
György B, Szabó T G, Pásztói M, Pál Z, Misják P, Aradi B, László V, Pállinger E, Pap E, Kittel A, Nagy G, Falus A, Buzás E I (2011). Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell Mol Life Sci, 68(16): 2667–2688
Hafez E (1952). Studies on the breeding season and reproduction of the ewe Part I. The breeding season in different environments Part II. The breeding season in one locality. J Agric Sci, 42(03): 189–231
Han X J, Lee M J, Yu G R, Lee Z W, Bae J Y, Bae Y C, Kang S H, Kim D G (2012). Altered dynamics of ubiquitin hybrid proteins during tumor cell apoptosis. Cell Death Dis, 3(1): e255
Hatzirodos N, Hummitzsch K, Irving-Rodgers H F, HarlandML, Morris S E, Rodgers R J (2014)a. Transcriptome profiling of granulosa cells from bovine ovarian follicles during atresia. BMC Genomics, 15(1): 40
Hatzirodos N, Irving-Rodgers H F, Hummitzsch K, HarlandML, Morris S E, Rodgers R J (2014)b. Transcriptome profiling of granulosa cells of bovine ovarian follicles during growth from small to large antral sizes. BMC Genomics, 15(1): 24
Hennet M L, Combelles C M (2012). The antral follicle: a microenvironment for oocyte differentiation. Int J Dev Biol, 56(10-11-12): 819–831
Holm S (1979). A simple sequentially rejective multiple test procedure. Scand J Stat, 6): 65–70
Kobayashi M, Oshima S, Maeyashiki C, Nibe Y, Otsubo K, Matsuzawa Y, Nemoto Y, Nagaishi T, Okamoto R, Tsuchiya K, Nakamura T, Watanabe M (2016). The ubiquitin hybrid gene UBA52 regulates ubiquitination of ribosome and sustains embryonic development. Sci Rep, 6(1): 36780
Kotni M K, Zhao M, Wei D Q (2016). Gene expression profiles and protein-protein interaction networks in amyotrophic lateral sclerosis patients with C9orf72 mutation. Orphanet J Rare Dis, 11(1): 148
Labrecque R, Fournier E, Sirard M A (2016). Transcriptome analysis of bovine oocytes from distinct follicle sizes: Insights from correlation network analysis. Mol Reprod Dev, 83(6): 558–569
Lai M D, Xu J (2007). Ribosomal proteins and colorectal cancer. Curr Genomics, 8(1): 43–49
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009). The sequence alignment/map format and SAMtools. Bioinformatics, 25(16): 2078–2079
Lin C Y, Lee T L, Chiu Y Y, Lin Y W, Lo Y S, Lin C T, Yang J M (2015). Module organization and variance in protein-protein interaction networks. Sci Rep, 5(1): 9386
Lotia S, Montojo J, Dong Y, Bader G D, Pico A R (2013). Cytoscape app store. Bioinformatics, 29(10): 1350–1351
Malik A, Lee E J, Jan A T, Ahmad S, Cho K H, Kim J, Choi I (2015). Network Analysis for the Identification of Differentially Expressed Hub Genes Using Myogenin Knock-down Muscle Satellite Cells. PLoS One, 10(7): e0133597
Matragkou C N, Papachristou E T, Tezias S S, Tsiftsoglou A S, Choli-Papadopoulou T, Vizirianakis I S (2008). The potential role of ribosomal protein S5 on cell cycle arrest and initiation of murine erythroleukemia cell differentiation. J Cell Biochem, 104(4): 1477–1490
Mbayahaga J, Mandiki S N, Bister J L, Paquay R (1998). Body weight, oestrous and ovarian activity in local Burundian ewes and goats after parturition in the dry season. Anim Reprod Sci, 51(4): 289–300
Naora H, Nishida T, Shindo Y, Adachi M, Naora H (1995). Association of nbl gene expression and glucocorticoid-induced apoptosis in mouse thymus in vivo. Immunology, 85): 63–68
Noel B, Bister J L, Paquay R (1993). Ovarian follicular dynamics in Suffolk ewes at different periods of the year. J Reprod Fertil, 99(2): 695–700
Peluso J J, Steger R W (1978). Role of FSH in regulating granulosa cell division and follicular atresia in rats. J Reprod Fertil, 54(2): 275–278
Qi Y, Li X, Chang C, Xu F, He Q, Zhao Y, Wu L (2017). Ribosomal protein L23 negatively regulates cellular apoptosis via the RPL23/Miz-1/c-Myc circuit in higherrisk myelodysplastic syndrome. Sci Rep, 7(1): 2323
Ramsköld D, Wang E T, Burge C B, Sandberg R (2009). An abundance of ubiquitously expressed genes revealed by tissue transcriptome sequence data. PLOS Comput Biol, 5(12): e1000598
Raposo G, Nijman H W, Stoorvogel W, Liejendekker R, Harding C V, Melief C J, Geuze H J (1996). B lymphocytes secrete antigen-presenting vesicles. J Exp Med, 183(3): 1161–1172
Rodgers R J, Rodgers H F, Hall P F, Waterman M R, Simpson E R (1986). Immunolocalization of cholesterol side-chain-cleavage cytochrome P-450 and 17 alpha-hydroxylase cytochrome P-450 in bovine ovarian follicles. J Reprod Fertil, 78(2): 627–638
Romereim S M, Summers A F, Pohlmeier W E, Zhang P, Hou X, Talbott H A, Cushman R A, Wood J R, Davis J S, Cupp A S (2017). Gene expression profiling of bovine ovarian follicular and luteal cells provides insight into cellular identities and functions. Mol Cell Endocrinol, 439): 379–394
Shimizu T (2016). Molecular and cellular mechanisms for the regulation of ovarian follicular function in cows. J Reprod Dev, 62(4): 323–329
Skinner M K, Schmidt M, Savenkova M I, Sadler-Riggleman I, Nilsson E E (2008). Regulation of granulosa and theca cell transcriptomes during ovarian antral follicle development. Mol Reprod Dev, 75(9): 1457–1472
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, Simonovic M, Roth A, Santos A, Tsafou K P, Kuhn M, Bork P, Jensen L J, von Mering C (2015). STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res, 43(Database issue): D447–D452
Talebi R, Ahmadi A, Afraz F, Abdoli R (2016). Parkinson’s disease and lactoferrin: Analysis of dependent protein networks. Gene Rep, 4): 177–183
Talebi R, Ahmadi A, Afraz F (2018)a. Analysis of protein-protein interaction network based on transcriptome profiling of ovine granulosa cells identifies candidate genes in cyclic recruitment of ovarian follicles. J Anim Sci Technol, 60: 11
Talebi R, Ahmadi A, Afraz F, Sarry J, Plisson-Petit F, Genêt C, Fabre S (2018)b. Transcriptome analysis of ovine granulosa cells reveals differences between small antral follicles collected during the follicular and luteal phases. Theriogenology, 108): 103–117
Terenina E, Fabre S, Bonnet A, Monniaux D, Robert-Granié C, SanCristobal M, Sarry J, Vignoles F, Gondret F, Monget P, Tosser-Klopp G (2017). Differentially expressed genes and gene networks involved in pig ovarian follicular atresia. Physiol Genomics, 49(2): 67–80
Trapnell C, Williams B A, Pertea G, Mortazavi A, Kwan G, van BarenM J, Salzberg S L,Wold B J, Pachter L (2010). Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol, 28(5): 511–515
Vizirianakis I S, Papachristou E T, Andreadis P, Zopounidou E, Matragkou C N, Tsiftsoglou A S (2015). Genetic manipulation of RPS5 gene expression modulates the initiation of commitment of MEL cells to erythroid maturation: Implications in understanding ribosomopathies. Int J Oncol, 47(1): 303–314
Xu F, Stouffer R L, Müller J, Hennebold J D, Wright J W, Bahar A, Leder G, Peters M, Thorne M, Sims M, Wintermantel T, Lindenthal B (2011). Dynamics of the transcriptome in the primate ovulatory follicle. Mol Hum Reprod, 17(3): 152–165
Xu X, Zhao X, Lu L, Duan X, Qin H, Du X, Li G, Tao Z, Zhong S,Wang G (2016)a. Transcriptomic analysis of different stages of pigeon ovaries by RNA-Sequencing. Mol Reprod Dev, 83(7): 640–648
Xu Z, Zhou Y, Cao Y, Dinh T L, Wan J, Zhao M (2016)b. Identification of candidate biomarkers and analysis of prognostic values in ovarian cancer by integrated bioinformatics analysis. Med Oncol, 33(11): 130
Zhang J, Pan Z, Moloney S, Sheppard A (2014). RNA-Seq Analysis Implicates Detoxification Pathways in Ovine Mycotoxin Resistance. PLoS ONE9, 6: e99975
Zhang X, Huang L, Wu T, Feng Y, Ding Y, Ye P, Yin Z (2015). Transcriptomic analysis of ovaries from pigs with high and low litter size. PLoS One, 10(10): e0139514
Zhen X, Wu B, Wang J, Lu C, Gao H, Qiao J (2015). Increased Incidence of Mitochondrial Cytochrome C Oxidase 1 Gene Mutations in Patients with Primary Ovarian Insufficiency. PLoS One, 10(7): e0132610
Zhou X, Hao Q, Zhang Q, Liao J M, Ke J W, Liao P, Cao B, Lu H (2015). Ribosomal proteins L11 and L5 activate TAp73 by overcoming MDM2 inhibition. Cell Death Differ, 22(5): 755–766
Zitvogel L, Regnault A, Lozier A, Wolfers J, Flament C, Tenza D, Ricciardi-Castagnoli P, Raposo G, Amigorena S (1998). Eradication of established murine tumors using a novel cell-free vaccine: dendritic cell-derived exosomes. Nat Med, 4(5): 594–600
Acknowledgments
R.T. would like to special thanks to the Dr. Stéphane Fabre, PhD–HDR in INRA research center in Castanet-Tolosan, France, and the staff of the GeT-Genotoul genomic platform (https://doi.org/get/genotoul.fr) for the RNA sequencing, and Sarah Maman of the INRA Sigenae bioinformatics team for Galaxy support. R.T. also thanks Julien Sarry, technician assistance from INRA-GenPhySE research center in Castanet-Tolosan, France, for prepared the RNAseq libraries. Authors wanted to thank Dr. Abbas Farahavar, PhD in Bu-Ali Sina University of Hamedan, Iran, due to his assistances in collecting the mural granulosa cell. This work has been supported by a PhD grant from “Bu-Ali Sina University” and “Agricultural Biotechnology Research Institute” from Iran.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Talebi, R., Ahmadi, A. & Afraz, F. Construction of protein–protein interaction network based on transcriptome profiling of ovine granulosa cells during the sheep’s anestrus phase. Front. Biol. 13, 215–225 (2018). https://doi.org/10.1007/s11515-018-1499-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11515-018-1499-x