Abstract
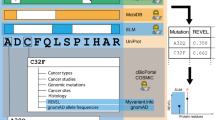
In this work, the most detrimental missense mutations of Mad1 protein that cause various types of cancer were identified computationally and the substrate binding efficiencies of those missense mutations were analyzed. Out of 13 missense mutations, IMutant 2.0, SIFTand PolyPhen programs identified 3 variants that were less stable, deleterious and damaging respectively. Subsequently, modeling of these 3 variants was performed to understand the change in their conformations with respect to the native Mad1 by computing their root mean squared deviation (RMSD). Furthermore, the native protein and the 3 mutants were docked with the binding partner Mad2 to explain the substrate binding efficiencies of those detrimental missense mutations. The docking studies identified that all the 3 mutants caused lower binding affinity for Mad2 than the native protein. Finally, normal mode analysis determined that the loss of binding affinity of these 3 mutants was caused by altered flexibility in the amino acids that bind to Mad2 compared with the native protein. Thus, the present study showed that majority of the substrate binding amino acids in those 3 mutants displayed loss of flexibility, which could be the theoretical explanation of decreased binding affinity between the mutant Mad1 and Mad2.
Similar content being viewed by others
References
Bava K A, Gromiha M M, Uedaira H, Kitajima K, Sarai A (2004). ProTherm, version 4.0: thermodynamic database for proteins and mutants. Nucleic Acids Res, 32(90001 Database issue): D120–D121
Berman HM, Battistuz T, Bhat T N, Bluhm WF, Bourne P E, Burkhardt K, Feng Z, Gilliland G L, Iype L, Jain S, Fagan P, Marvin J, Padilla D, Ravichandran V, Schneider B, Thanki N, Weissig H, Westbrook J D, Zardecki C (2002). The Protein Data Bank. Acta Crystallogr D Biol Crystallogr, 58(Pt 6 No 1): 899–907
Bharadwaj R, Yu H (2004). The spindle checkpoint, aneuploidy, and cancer. Oncogene, 23(11): 2016–2027
Boeckmann B, Bairoch A, Apweiler R, Blatter M C, Estreicher A, Gasteiger E, Martin M J, Michoud K, O’Donovan C, Phan I, Pilbout S, Schneider M (2003). The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res, 31(1): 365–370
Brady DM, Hardwick K G (2000). Complex formation between Mad1p, Bub1p and Bub3p is crucial for spindle checkpoint function. Curr Biol, 10(11): 675–678
Cahill D P, Lengauer C, Yu J, Riggins G J, Willson J K, Markowitz S D, Kinzler K W, Vogelstein B (1998). Mutations of mitotic checkpoint genes in human cancers. Nature, 392(6673): 300–303
Capriotti E, Fariselli P, Casadio R (2005). I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res, 33(Web Server Web Server issue): W306–10
Carlson H A, McCammon J A (2000). Accommodating protein flexibility in computational drug design. Mol Pharmacol, 57(2): 213–218
Chao W C, Kulkarni K, Zhang Z, Kong E H, Barford D (2012). Structure of the mitotic checkpoint complex. Nature, 484(7393): 208–213
Chen R H, Shevchenko A, Mann M, Murray A W (1998). Spindle checkpoint protein Xmad1 recruits Xmad2 to unattached kinetochores. J Cell Biol, 143(2): 283–295
Chen R H, Waters J C, Salmon E D, Murray AW (1996). Association of spindle assembly checkpoint component XMAD2 with unattached kinetochores. Science, 274(5285): 242–246
Chung E, Chen R H (2002). Spindle checkpoint requires Mad1-bound and Mad1-free Mad2. Mol Biol Cell, 13(5): 1501–1511
Connolly M L (1983). Solvent-accessible surfaces of proteins and nucleic acids. Science, 221(4612): 709–713
Delarue M, Dumas P (2004). On the use of low-frequency normal modes to enforce collective movements in refining macromolecular structural models. Proc Natl Acad Sci USA, 101(18): 6957–6962
Duhovny D, Nussinov R, Wolfson H J (2002). Efficient unbound docking of rigid molecules. In: Proceedings of the 2nd Workshop on Algorithms in Bioinformatics (WABI) Lecture Notes in Computer Science, Rome, Italy, 2452: 185–200
Fava L L, Kaulich M, Nigg E A, Santamaria A (2011). Probing the in vivo function of Mad1:C-Mad2 in the spindle assembly checkpoint. EMBO J, 30(16): 3322–3336
Gemma A, Seike M, Seike Y, Uematsu K, Hibino S, Kurimoto F, Yoshimura A, Shibuya M, Harris C C, Kudoh S (2000). Somatic mutation of the hBUB1 mitotic checkpoint gene in primary lung cancer. Genes Chromosomes Cancer, 29(3): 213–218
Han J H, Kerrison N, Chothia C, Teichmann S A (2006). Divergence of interdomain geometry in two-domain proteins. Structure, 14(5): 935–945
Han S, Park K, Kim H Y, Lee M S, Kim H J, Kim Y D, Yuh Y J, Kim S R, Suh H S (2000). Clinical implication of altered expression of Mad1 protein in human breast carcinoma. Cancer, 88(7): 1623–1632
Hardwick K G, Weiss E, Luca F C, Winey M, Murray A W (1996). Activation of the budding yeast spindle assembly checkpoint without mitotic spindle disruption. Science, 273(5277): 953–956
Hinkle A, Tobacman L S (2003). Folding and function of the troponin tail domain. Effects of cardiomyopathic troponin T mutations. J Biol Chem, 278(1): 506–513
Hoyt M A, Totis L, Roberts B T S (1991). S. cerevisiae genes required for cell cycle arrest in response to loss of microtubule function. Cell, 66(3): 507–517
Hwang L H, Lau L F, Smith D L, Mistrot C A, Hardwick K G, Hwang E S, Amon A, Murray A W (1998). Budding yeast Cdc20: a target of the spindle checkpoint. Science, 279(5353): 1041–1044
Jallepalli P V, Lengauer C (2001). Chromosome segregation and cancer: cutting through the mystery. Nat Rev Cancer, 1(2): 109–117
Kim S H, Lin D P, Matsumoto S, Kitazono A, Matsumoto T (1998). Fission yeast Slp1: an effector of the Mad2-dependent spindle checkpoint. Science, 279(5353): 1045–1047
Li R, Murray AW (1991). Feedback control of mitosis in budding yeast. Cell, 66(3): 519–531
Lindahl E, Azuara C, Koehl P, Delarue M (2006). NOMAD-Ref: visualization, deformation and refinement of macromolecular structures based on all-atom normal mode analysis. Nucleic Acids Res, 34(Web Server Web Server issue): W52–6
Lopes C S, Sunkel C E (2003). The spindle checkpoint: from normal cell division to tumorigenesis. Arch Med Res, 34(3): 155–165
Michael S C, Gary J G (1995). Microinjection of mitotic cells with the 3F3/2 Anti-phosphoepitope antibody delays the onset of anaphase. J Cell Biol, 129(5): 1195–1204
Mimori K, Inoue H, Alder H, Ueo H, Tanaka Y, Mori M (2001). Mutation analysis of hBUB1, human mitotic checkpoint gene in multiple carcinomas. Oncol Rep, 8(1): 39–42
Ng P C, Henikoff S (2001). Predicting deleterious amino acid substitutions. Genome Res, 11(5): 863–874
Ng P C, Henikoff S (2003). SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res, 31(13): 3812–3814
Nomoto S, Haruki N, Takahashi T, Masuda A, Koshikawa T, Takahashi T, Fujii Y, Osada H, Takahashi T (1999). Search for in vivo somatic mutations in the mitotic checkpoint gene, hMAD1, in human lung cancers. Oncogene, 18(50): 7180–7183
Ohshima K, Haraoka S, Yoshioka S, Hamasaki M, Fujiki T, Suzumiya J, Kawasaki C, Kanda M, Kikuchi M (2000). Mutation analysis of mitotic checkpoint genes (hBUB1 and hBUBR1) and microsatellite instability in adult T-cell leukemia/lymphoma. Cancer Lett, 158(2): 141–150
Peters J M (2006). The anaphase promoting complex/cyclosome: a machine designed to destroy. Nat Rev Mol Cell Biol, 7(9): 644–656
Peters J M (2008). Checkpoint control: the journey continues. Curr Biol, 18(4): R170–R172
Rajasekaran R, Priya Doss C G, Sudandiradoss C, Ramanathan K, Sethumadhavan R (2008). In silico analysis of structural and functional consequences in p16INK4A by deleterious nsSNPs associated CDKN2A gene in malignant melanoma. Biochimie, 90(10): 1523–1529
Ramensky V, Bork P, Sunyaev S (2002). Human non-synonymous SNPs: server and survey. Nucleic Acids Res, 30(17): 3894–3900
Reis R M, Nakamura M, Masuoka J, Watanabe T, Colella S, Yonekawa Y, Kleihues P, Ohgaki H (2001). Mutation analysis of hBUB1, hBUBR1 and hBUB3 genes in glioblastomas. Acta Neuropathol, 101(4): 297–304
Sato M, Sekido Y, Horio Y, Takahashi M, Saito H, Minna J D, Shimokata K, Hasegawa Y (2000). Infrequent mutation of the hBUB1 and hBUBR1 genes in human lung cancer. Jpn J Cancer Res, 91(5): 504–509
Schneidman-Duhovny D, Inbar Y, Nussinov R, Wolfson H J (2005). PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res, 33(Web Server issue): W363–7
Suhre K, Sanejouand Y H (2004). ElNemo: a normal mode web server for protein movement analysis and the generation of templates for molecular replacement. Nucleic Acids Res, 32( Web Server issue): W610–4
Tina K G, Bhadra R, Srinivasan N (2007). PIC: Protein Interactions Calculator. Nucleic Acids Res, 35(Web Server issue): W473–W476
Tsukasaki K, Miller C W, Greenspun E, Eshaghian S, Kawabata H, Fujimoto T, Tomonaga M, Sawyers C, Said JW, Koeffler H P (2001). Mutations in the mitotic check point gene, MAD1L1, in human cancers. Oncogene, 20(25): 3301–3305
Varfolomeev S D, Uporov I V, Fedorov E V (2002). Bioinformatics and molecular modeling in chemical enzymology. Active sites of hydrolases. Biochemistry (Mosc), 67(10): 1099–1108
Wang X, Jin D Y, Ng R W, Feng H, Wong Y C, Cheung A L, Tsao S W (2002). Significance of MAD2 expression to mitotic checkpoint control in ovarian cancer cells. Cancer Res, 62(6): 1662–1668
Yip Y L, Famiglietti M, Gos A, Duek P D, David F P, Gateau A, Bairoch A (2008). Annotating single amino acid polymorphisms in the UniProt/Swiss-Prot knowledgebase. Hum Mutat, 29(3): 361–366
Yip Y L, Scheib H, Diemand A V, Gattiker A, Famiglietti L M, Gasteiger E, Bairoch A (2004). The Swiss-Prot variant page and the ModSNP database: a resource for sequence and structure information on human protein variants. Hum Mutat, 23(5): 464–470
Yu H (2002). Regulation of APC-Cdc20 by the spindle checkpoint. Curr Opin Cell Biol, 14(6): 706–714
Zhang C, Vasmatzis G, Cornette J L, DeLisi C (1997). Determination of atomic desolvation energies from the structures of crystallized proteins. J Mol Biol, 267(3): 707–726
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lopus, M., Sethumadhavan, R., Chandrasekaran, P. et al. A computational approach to explore the functional missense mutations in the spindle check point protein Mad1. Front. Biol. 8, 618–625 (2013). https://doi.org/10.1007/s11515-013-1280-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11515-013-1280-0