Abstract
Essential factors that control gene stability and expression are collectively known as epigenetics. Within the most well-studied epigenetic mechanisms are DNA methylation and histone modifications. A broad range of methods has been used for identifying differentially methylated regions, including biotechnological and enzymatic techniques. Nevertheless, in the last decade, there has been a proliferation of techniques called plasmonics which have emerged as an alternative to studying epigenetics. They take advantage of the different chemical composition of methylated compared to unmethylated histones and nucleotides to quantify their optical properties. Here, we introduce the basics of plasmonics and present a detailed description of how these techniques work. We also provide an outlook on the application of plasmonics in plant epigenetics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epigenetic regulation is an essential component of the transcriptional expression machinery [1]. Within the different epigenetic mechanisms, DNA methylation and histone modifications have been shown to be essential during plant development as well as under stressful conditions [2,3,4]. For instance, DNA methylation is fundamental for transcriptional repression of transposable elements (TEs) and regulation of gene expression [5, 6]. In plants, DNA methylation occurs at cytosine residue to produce 5-methylcytosine and it is found in CG, CHG, and CHH sequence context (where H represents A, T, or C). While methylation at CHG and CHH sites is predominantly found in TEs, CG methylation occurs in TEs and protein-encoding genes [7, 8].
To identify differentially methylated regions, bisulfite sequencing, methyl-sensitive cut counting (MSCC) using endonuclease digestion followed by sequencing, and enzymatic methyl-seq (EM-seq) have been the traditional strategies. Bisulfite sequencing is the most popular and widely used method in DNA methylation studies [9]. In this method, bisulfite-treated DNA samples undergo deamination of cytosine into uracil. During PCR-amplification and subsequent DNA sequencing analysis, uracil-converted residues will be read as thymine. However, methylated cytosine is resistant to uracil conversion. Thus, it remains read as cytosine. Comparative analysis of treated and untreated DNA samples enables the identification of methylated cytosines. MSCC uses methylation-sensitive endonucleases for initial digestion of unmethylated DNA sites followed by adaptor ligation that contains the site for another digestion enzyme that cuts outside of its recognized site. This strategy allows the generation of small fragments that can be sequenced. Interestingly, this strategy can be implemented using enzymes that cut either methylated or unmethylated DNA. There are a good number of methylation-dependent endonucleases that can use methylated DNA as a substrate. Examples of those include BisI, BlsI, GlaI, GluI, KroI, McrBC, MspI, MteI, PcsI, and PkrI. In plants, McrBC has been widely used in model and crop plant species [2, 10,11,12]. McrBC is an endonuclease able to cleave DNA containing methylcytosine between two half-sites of (G/A) mC that are located within 50–3000 bp from each other [13, 14]. Unlike the previously described techniques, EM-seq selectively uses a first enzymatic step to oxidize methylated cytosines and a second enzymatic step to convert unmethylated cytosines to uracils. After a subsequent PCR amplification step, oxidized methylcytosine forms base pairs with guanine, and uracil forms base pairs with adenine. EM-seq enzymatic reactions are non-destructive. Therefore, a higher yield and accuracy are obtained.
Even though these methods have been the gold standard for epigenetic studies, optical techniques, collectively known as plasmonics, have recently captured the interest of researchers in the epigenetic field. The use of plasmonics has been well implemented in chemistry, physics, and medicine, among other STEM fields. However, the area where these interactions have flourished in the last decade is on biological systems where both bare or functionalized metallic nanoprobes allow detecting and quantifying, targeting specific intracellular processes or as colorimetric biosensors for monitoring or detection objectives [15,16,17,18]. Plasmonics takes advantage of both light scattering and absorption processes and even can use the naked eye as a detector. When a process occurring close to nanoparticles in colloidal suspension causes its aggregation, a change in color and hence a shift of the absorption spectra and scattering profile is observed and can be measured. The heart of the application of all these techniques is the metallic nanostructured surface, either in the form of colloidal suspension, as a solid substrate for SERS spectroscopy, or in colloidal form for dark-field and colorimetric applications. Among all possibilities of bioanalysis, those related to chemical DNA modifications have recently captivated the interest in plasmonics with the additional advantage that its application in this field does not involve nucleotide-based amplifications as in other classical sensing procedures [19,20,21]. This is reinforced by the fact that DNA bases have high affinity towards metallic nanoparticles, mainly gold and silver nanoparticles [22]. Here, we present a comprehensive review of the strategies that exploit the possibilities of techniques based on plasmonics for studying processes associated with chemical modifications of DNA bases or epigenetics.
Fundamentals of Plasmonics
A specific condition is reached when light is confined either in between or at the proximities of a nanostructured metal surface where electromagnetic oscillations or surface plasmon polaritons permanently occur. When these oscillating electrons compelled to the geometry of a nanostructure are irradiated with an excitation wavelength bigger than the nanoparticle size, a resonant interaction starts, causing a localized oscillation with frequency known as localized surface plasmon resonance (LSPR). As a result, unique optical properties arise, being one of the most important the enhancement of the electromagnetic field. As these conditions allow manipulating light at levels below the diffraction limit, the intensity of signals coming from species adsorbed on the metallic surface is drastically intensified. Plasmonics embrace this junction between surface plasmons and light and it is the core in the development of analytical techniques in nano-optics [23].
One of the most interesting plasmonics techniques that have become well known and that take advantage of those optical properties is surface-enhanced Raman spectroscopy (SERS). SERS relies on the strong signal amplification effect of scattered light obtained when a molecule is close, or better, in the gap between two metallic nanoparticles, a region known as hotspots. The irradiation of a chemical species in this condition results in enhancement factors as high as 105–106 times, allowing the acquisition of spectra with a very high signal-to-noise (S/N) ratio. This is perhaps the main reason that has driven the technique from electrochemical and physical chemistry studies to an increasing amount of analytical and quantitative applications, many of them related to biological systems, even though reproducibility of the spectra has been continuously on the spot as its main disadvantage [24, 25].
Scattered light by a metallic nanoparticle acting as a nanoprobe can also be detected through an optical camera and this possibility allows the study of processes happening at nanoparticle or single molecule level in real time. The optical properties that depend on the nanoparticle size, morphology, composition, and refractive index of the surroundings have been the basics for the relatively recent development of the dark-field spectroscopy technique that offers both spectrum and image as output. Using these properties, it is possible to obtain the scattering intensity and its location at the same time [26].
Contrary to the relatively recent development of these techniques, synthesis procedures of noble metal nanostructures have been widely studied, and a myriad of modifications for tuning their physical properties are available in the literature [27]. Nevertheless, solid nanostructured substrates can be prepared by simply dropping a colloidal suspension, or by generating a roughened electrode. They have also gained benefits from modern techniques such as sputtering or electron beam lithography, to mention a few. Syntheses of colloidal nanoparticles are available in all sorts of different procedures, where most of them require simple glassware and involve the use of a reduction agent for taking the metal to its zero-valent form. In the search for a highly reproducible and reliable colloid, many researchers have focused on understanding the mechanisms behind the formation of nanoparticles with a given size and morphology, and various experimental conditions have been studied and evaluated, from modifications in temperature, stirring speed or reaction time, to several reduction agents and stabilizers [24]. In the face of all available morphologies, perhaps the most popular are colloidal nanospheres, since their synthesis is relatively well-established, and reproducible enough, and plenty of applications have been already reported [28, 29].
Plasmonic Techniques
Colorimetric Sensors
Nanoparticles-based colorimetric sensing is perhaps the simpler approach of plasmonic techniques. This methodology is based on the same principle of conventional colorimetric analysis where a given procedure induces the appearance of color (or changes in it), and the detection step is performed by UV–Vis spectrophotometry or even by the naked eye [30, 31]. When introduced to colorimetric detection, the unique plasmon resonance properties of noble metal nanoparticles enable ultrasensitive detection and its absorbance (and hence, their color) can be tuned by modifying their morphology, size, metal, and chemical environment. The extraordinary optical properties come from the free electron oscillation on the metallic surface when stimulated by light. Specifically, the main phenomenon involved is the localized surface plasmon resonance (LSPR), characterized by its high confinement at the nanostructures and enhanced electromagnetic field. The absorption profile shows a peak in the visible and near-infrared regions [32, 33].
Two main experimental strategies can be adopted to perform colorimetric assays, both based on exploiting the intrinsic properties of nanoparticles by modifying interparticle distance and morphology, and size. Changes in the interparticle distance are related to nanoparticle aggregation that induces surface plasmon coupling between particles, accompanying the LSPR shift and color transition at the solution. For instance, the red-to-blue transitions in gold nanoparticles, AuNPs upon aggregation. This strategy is practical for sensing several targets, from small molecules to macromolecules or living cells. More specific modulations are driven by covalent and non-covalent forces such as hydrogen bonding or electrostatic interactions, dependent on pH and temperature. The electrostatic interactions have a direct influence on the nanoparticles assembly aggregation and its redispersion can be tuned by external changes in the surroundings, among them pH alterations, electrical field, temperature, mechanical stress, or presence of specific species (DNA, proteins, for example), and the result will be an LSPR shift, the key step where detections are realized. However, the main disadvantage of relying on electrostatic interactions is a diminished sensitivity, due in part to the low strength of these forces, although several procedures (including polymerization, physicochemical reactions, and catalysis) can be done to improve it [34,35,36].
Another strategy that can be successfully explored lies in the incorporation of functional groups on the nanoparticles surface through a stronger interaction named covalent bonding. This improves sensitivity and facilitates surface functionalization. Hence, more specific analyses can be performed. The hydrophobic character on NPs surfaces or in its surroundings have also an important role in building sensing strategies, even with suitable specificity [37]. Assemblies with highly specific recognition can be obtained by exploiting multiple intermolecular interactions and can be biologically specific. For instance, using nucleobases functionalized NPs to recognize a target species, although the sensitivity of these approaches directly depends on the aggregate size. Another interesting way of reaching specific bindings is based on the concept of host–guest molecules that guarantees, in the first place, high selectivity. It is worth mentioning that non-covalent interactions such as pi-stacking (in aromatic systems) or Van Der Waals forces are useful in plasmonic sensing methodologies. The predominant interaction exploited in its assembly will largely depend on the physical–chemical nature of the whole system [38, 39].
Tuning morphology and size of metal NPs is also an interesting strategy that induces changes in their optical properties, mainly their LSPR, and thus performs recognition. There are numerous factors associated with the NPs synthesis that controls its morphology and size, among them, precursors, capping, and reducing agents. Usually, small changes in it can give origin to quite different types of NPs and thereby shifts in their LSPR are observed. Thus, the species of interest can be used for example as the reducing agent and is the growth process itself that permits its quantification. Another approach uses the molecule target to carry out a controlled etching procedure that starts with an oxidation step of the NPs, until inducing detectable modifications in its composition of morphology. Then, the extent of the etching will be directly related to the number of species of interest leading to their quantification. When the target is used as the reducing agent, its quantification is performed through the controlled growth of NPs, reflected in the LSPR shift [40]. This methodology has advantages; likewise, its short time of analysis makes it suitable for rapid and simple applications. Various applications include the use of enzymes as reducing agents and hence control NPs growth for several applications [41]. Changes in morphology take advantage of the possibility of reshaping NPs in the presence of a given species (the target). This chemical etching process can be mediated by a variety of compounds, making possible the development of sensing procedures of targets such as hydrogen peroxide, heavy metals, and anionic species. Although currently, NPs for plasmonic colorimetric essays are synthesized from mainly noble metals gold and silver, copper NPs are also used, despite their application could be challenging due to the natural reactivity of this metal. The properties of plasmonic surfaces can also be modified by using different coating materials (silica, polymers) and thickness, or even NPs of metal oxides or a variety of composites.
Surface Plasmon Resonance-Based Techniques
Surface plasmon resonance (SPR) techniques rely on the anomalous diffraction obtained upon the excitation of surface plasmon waves, a phenomenon observed for the first time by Wood in 1902 and that boosted the development of various sensing approaches, from microscopy-based techniques to SPR imaging [42, 43].
SPR is defined as the charge-density oscillations that occur at the interface between two media (a metal and a dielectric) having dielectric constants of opposite signs. When the electric field vector from the incident light at a given angle, a part of it matches the free electron movement in the metal (constrained by boundary conditions coming from the material nature), excitation of surface plasma waves occurs, and because of this efficient coupling, a guided electromagnetic wave along the interface is generated. This is called plasmon and it propagates parallel to the metal surface. A typical SPR biosensor configuration includes a high-reflective glass prism as shown in Fig. 1, and it is known as Kretschmann geometry [44, 45].
Kretschmann geometry. A A typical surface plasmon resonance (SPR) setup includes a light source, a metal layer put at the base of the prism separating it from the dielectric medium containing the analytes or species of interest. As a detector, a charged-coupled device camera is usually used. B Functional coating with analyte-biorecognition species on the SPR sensor surface. C Reflected light spectra obtained upon changes in refractive index. D Changes in refractive index due to molecular interactions in the dielectric medium. Before (I) and after (II) binding
As the angle at which resonance is generated depends on the refractive index of the material close to the metal surface, when a small change in the reflective index of the sensing medium, there is no plasmon formation. Thus, the changes in the reflected light are measured at the detector, and resonance angle shifts or intensity can also be monitored and associated with quantitative information. In practice, an SPR biosensor works by first immobilizing a probe molecule on the sensor surface, and in a second stage, the analyte is put in contact with the modified surface where affinity interactions should occur. After the probe-analyte binding, there is an increase of the refractive index at the SPR surface, and the resulting signal change is measured in resonance or response units (RU). Each RU unit is equal to a critical angle shift of 10−4 degrees and the first RU value for measurement is the starting critical angle.
The analyte concentration for a layer of thickness h is related to the change in the refractive index delta (n) through Eq. 1 below:
where \({(\frac{dn}{dc})}_{vol}\) is the increase of refractive index, n; vol is the volume of the analyte of concentration c. In real time, when the incident light becomes coupled to the propagating surface plasmon, it allows the detection and quantification of the target molecules. The limit of detection (LOD) will depend on the binding affinity of the analyte to the probe molecule, molecular weight, and the coverage surface area of the immobilized molecule. Other particularly useful information as specificity and kinetic parameters can be determined by using SPR approaches. The bioanalytical field has taken great advantage of the information that SPR techniques offer and numerous applications on biological systems including proteins, enzymes, DNA, and virus have been widely studied.
Surface-Enhanced Raman Spectroscopy
The surface-enhanced Raman spectroscopy (SERS) technique is based on the surface-enhanced Raman scattering effect obtained when an incident beam of monochromatic light interacts with a given molecule attached to a noble metal nanostructured surface, mainly fabricated with gold, silver, or copper. The inelastic light scattered by the molecule can be intensified to a large extent (factors up to 108 or higher) being able to reach very low LOD values (up to femtomolar scale) [46]. This effect was accidentally observed when a pyridine molecule was adsorbed on gold electrodes, initially by Fleishmann and co-workers [47] and later by Albrecht and Creighton [48] which associated this phenomenon with plasmon excitation. Through approximately 40 years of SERS research, a remarkably high number of both applied and fundamental studies have been carried out [49,50,51,52] and although quantitative applications were at some point, difficult to perform, today SERS spectroscopy is for sure an exceptionally good option for quantitative purposes [53, 54].
A classical SERS experiment involves in the first step, synthesizing a well-ordered (ideally) nanostructured substrate either as a solid surface or as a colloidal suspension. Although the effect can be obtained on copper, silver, or gold surfaces, while silver offers higher intensification factors, gold nanostructures allow more reproducible signals and are more suitable for biological applications due to their low ability to cause damage to these systems. In the following stage, a procedure for an efficient binding molecule-nanoparticle is realized. For solid surfaces, it is usual to deposit the analyte of interest from a solution either manually or using an automated device.
When nanoparticles are in the form of colloidal suspensions, various possibilities are in the landscape: they could be used as probes itself inside complex systems as cells or tissues (for imaging, for example), or they can be functionalized with a Raman marker to probe a specific bond or to track its route or modifications. The simpler way for using colloidal nanostructures includes a main attaching step where the molecule is adsorbed onto the surface. This procedure can be modified or customized and this is usually done by functionalizing the surface according to the chemical nature of the species of interest and depending on the objective of analysis.
The phenomena behind the high signal enhancement factors that can be obtained with this technique start at nanometer-sized gaps also known as hotspots (typically in the 2–10 nm size range) where the electromagnetic field is intensified. They can be formed between two metallic nanoparticles, or within their junctions and a flat metal surface. Geometrical properties and the gap distance between other characteristics play an important role in the intensification effect. Two main mechanisms are considered responsible for the signal intensification; the first one relies on an enhanced electromagnetic field arising upon plasmon excitation. The second is related to the contributions of Raman scattering not associated with the electromagnetic environment but related to the electrons being transferred between the molecule and the surface or substrate. It is important to remark that in practice, these two effects cannot be rigorously separated [55].
Applications
Colorimetric Sensors
The relative simplicity of measurements using plasmonic colorimetric sensors has allowed the development of several methodologies for assessing epigenetic biomarkers. Studies on DNA methylation modification have shown a notoriously increasing trend over the last ~ 5 years [56]. Among the available metallic nanostructure surfaces, AuNPs have been perhaps the most popular, due to their unique sensitivity associated with changes in size and distance in NPs, high stability, and due to be more chemically inert, although their extinction coefficient is lower than the AgNPs’ [57].
An unamplified genomic DNA (gDNA) nanosensor was developed by Baetsen-Young et al. [58] using dextrin-capped AuNPs. The study was focused on the detection of DNA sequence from Pseudoperonospora cubensis, a primary threat for cucumber production. Another instance is the one-step procedure (dextrin and sodium carbonate were used as capping and reducing agents, respectively) that generates 13 nm glycol-AuNPs [59]. Using gDNA target isolated from Pseudoperonospora cubensis, and non-target DNA, from cucumber leaves, the authors successfully achieved a highly specific methodology, and even differences were visually distinguished. Regarding sensitivity in the detection of extracted and crude (non-extracted) DNA, values in the order of 3 fM for the former and 18 sporangia uL−1 for the latter were reported. Overall, this assay is not expensive, it is rapid (~ 30 min visual assay), does not require specific equipment, and it is becoming a suitable option to currently available diagnostic methods such as ELISA and PCR.
Although so far, studies focused on plant cells are still starting to be developed, an important number of applications in human cells showed how these strategies are very promising for identifying epigenetic modifications in eukaryotic genomes in general [60]. In a recent application, the formaldehyde obtained from HDMs-catalyzed demethylation was used as a probe [61]. This approach explores the fact that the formaldehyde can quantitatively inhibit the oxidation reaction of o-phenylenediamine Ag+-triggered, causing a decrease in fluorescence and absorbance and allowing a dual-mode reading. The associated mechanism behind both signal modes is shown in Fig. 2.
A Demethylation reaction catalyzed by HDMS with the release of formaldehyde. B In absence of formaldehyde, the o-phenylenediamine oxidation reaction (Ag + -triggered) is not inhibited allowing measuring fluorescence. C The formaldehyde inhibits the o-phenylenediamine oxidation and absorbance signals can be obtained
SERS
Being one of the most important epigenetic mechanisms, DNA methylation has been extensively studied through techniques such as HPLC–UV or PCR [62]. Large amounts of DNA samples are required for the former and a careful design and selection of primers is mandatory for the latter, not to mention the time-consuming procedures of sample preparation, as a common disadvantage. In these aspects, SERS is showing to be an attractive option, especially because DNA bases have a particularly high affinity towards noble metal nanostructured surfaces, evidenced through their rich-information spectra (Fig. 3) [22, 63]. In addition, SERS is a label-free technique, allowing reaching limit of detection values as low as 10−7 mol L−1 and sample conditioning is typically easy to perform.
A A typical architecture of the DNA nucleobases adenine and thymine attached to gold and silver nanoparticles, along with their macroscopic appearance in colloidal form. B Their SERS spectra with the main Raman frequencies: bands from in-plane symmetric ring breathing modes (665–802 cm−1), -C = O stretching for guanine and thymine (1640–1657 cm−1)
A procedure for the detection of DNA methylation combining SERS and chemometrics has been already implemented [64] where a combined Au@Ag core–shell nanopillar structure was used as a plasmonic substrate. Samples of guanine adducts were analyzed in concentrations between 1 nM and 10 µM. Chemometric modeling showed that the methodology allows clearly distinguishing non-methylated (guanine) and methylated guanine bases, at different positions (O6-methylguanine, N7-methylguanine, O6-hydroxyethyl guanine, N7-2hydroxyethyl guanine, O6-benzyl guanine, and O6-methyl-d3- guanine).
Using a plasmonic gold nanohole array and SERS, the methylation of cytosine was studied [65]. As a control system, a 9-base CpG sequence was used. Three main spectral peaks responded to methylation: a lowering in the intensity of ring breathing (785 cm−1), and ring stretching (1225 cm−1) modes of cytosine and increasing of -CH3 group vibration band (1365 cm−1). In the work by Luo et al. [66], a solid gold SERS substrate was used to evaluate the activity of M.SssI (CpG Methyltransferase). Specifically, the sequence 5′-CCGG-3′ contained in a gene fragment was chosen as the target DNA. A probing DNA, carrying both a recognition unit (a target-DNA-recognized sequence) and an extending spacer for triggering the hybridization chain reaction, was included. The probe, containing an assembled -SH at the 3′-end position, was immobilized through an Au–S bond. Moreover, its 5′-end position was labeled with Cy5, chosen as a Raman reporter that is expected to be absent when M.SssI is missing in the sample, becoming an indicator of methylation. Still, its signal intensity increases with M.SssI activity, allowing its quantitative determination (LOD was 2 10−4 U mL−1), a sensitivity comparable to fluorescence-based assays. Another advantage of the methodology reported in this work is the selective detection of M.SssI in human serum, from other MTase interferences, making this essay promising for methylation-related diseases and also plant-related research.
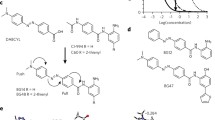
Mirajkar et al. [67] showed how SERS spectroscopy could be used to distinguish genomic DNA (gDNA) from two plants (Brassica juncea and Arabidopsis thaliana), identifying the ratio of adenine and deoxyribose as the sites of the molecule that interacts to the surface of metallic NPs. Extraction of gDNA was carried out following the CTAB extraction method, and Ag- and AuNPs were used as enhancer substrates, synthesized by well-known methods (hydroxylamine, borohydride, and citrate reduction [68, 69]), showing the potential of this technique. Interestingly, histone demethylation in plants has not been extensively studied using SERS. A work showed a relatively easy essay where the activity of two histone demethylases (HDM) could be assessed by taking advantage of the fact that both of them, through different routes, release formaldehyde as a by-product, probed using SERS [19]. Initially, a thiolated probe reacts with formaldehyde forming an adduct, and in a second stage, it covalently binds to a previously peptide-stabilized AuNPs as shown in Fig. 4. This procedure has an important potential as the probed by-product is also formed from the oxidative demethylation of proteins, nucleic acids, and other biological molecules.
SERS strategy for HDM assay. A Formaldehyde obtention from a diMe H3K4 peptide substrate catalyzed by flavin-dependent amine oxidase (LDS1). B The reaction of formaldehyde with Purpald, a thiolated reactive probe, forming an adduct. The adduct binds through covalent interactions to the surface of peptide-stabilized AuNPs for SERS spectrum measurement
In another application, duplex SERS nanotags (silica-coated dimer and trimer AuNPs with a 4-mercapto-3-nitrobenzoic acid and 4-mercaptobenzoic acid as Raman reporters) for simultaneous monitoring of DNA methylation levels and quantitative assessment of total gDNA methylation have been used [70]. In this approach, gDNA is enzymatically digested into DNA fragments. Biotin-dNTPs were filled into the ends of the DNA fragments with DNA polymerase, followed by incubation with streptavidin magnetic beads and SERS nanotags. The beads capture all available biotinylated DNA and the streptavidin-SERS nanotags (SERS-SA) represent the DNA contained in the sample. Thus, the SERS-SA works as an internal control of the gDNA amount. At the same time, nanotags labeled with methyl binding domain proteins (SERS-MBD) are used for specific binding methyl groups in DNA. With the information coming from the Raman intensity of the spectra, the % methylation could be quantified.
SPR Sensors
The use of SPR biosensors has shown to be especially suitable for studying biomolecular interactions, and applications in DNA sensing have grown notoriously in the last 15–20 years [71]. In brief, an ssDNA probe is attached to the sensor surface and a fast, real-time monitoring of hybridization with the target DNA is possible. Moreover, this methodology allows tuning the oligonucleotide probe for increased specificity. Although commonly used methods such as the well-known polymerase chain reaction (PCR) offers high selectivity, specificity, and great versatility, it requires a DNA amplification procedure, making it time-consuming and expensive. Progress in SPR methodologies has made the technique simple, fast, and reliable. In this type of application, it is worth mentioning the work of Plácido et al. [72] and Grześkowiak et al. [73] which used biotinylated ssDNA captured probes coupled to SPR biosensor chips used for GMO detection or quantification of gene expression. Plácido et al. [72] targeted the transgenic construct of Roundup Ready (RR) in soybean and the lectin (the taxon-specific soybean gene). For the sensing probe, a universal ssDNA molecule conjugated with streptavidin (SA) binds to the SPR chip with a complementary sequence previously immobilized onto it. The system captures the biotinylated probe, and then the analyte could be studied. In this case, genomic DNA (from soybean bagasse, soybean seeds, soybean extract, textured soybean, and soy protein) was extracted and amplified, and after analysis, the SPR biosensor surface was regenerated and can be reused. Although the amplification procedure is perhaps the most time- and reagent-consuming step, the amplicons do not need further steps of purification or denaturation. In the work by Grześkowiak et al. [73], a gold NPs-based biosensor was used in the detection of foreign nucleic acids in genomic DNA without amplification, in transgenic tobacco plants expressing a Streptococcus mutans antigen. The target DNA was detected following a genomic DNA isolation procedure from roots, stems, and leaves of tobacco transgenic plants. The sensing strategy included the functionalization of AuNPs with streptavidin and a biotinylated DNA probe [74], and a streptavidin-coated SPR sensor where a sixteen-nucleotide-long biotinylated DNA probe was immobilized (Fig. 5).
Measurements here were performed using the coated SPR chips, where positive control and DNA samples had sequences complementary to the biotinylated DNA probes, and functionalized AuNPs were injected as a strategy for signal enhancing, achieving a significant increase in sensitivity. In addition, this methodology is shown to be rapid and does not require a DNA amplification step as in conventional diagnostic methods or in similar SPR applications [75]. In the work by Nguyen and Sim [45], a simultaneous detection of both hot-spot mutation and epigenetic changes was carried out in circulating tumor DNA (ctDNA), a promising biomarker for assessing cancer in a noninvasive way. To accomplish this, a biosensing platform or nanoplasmonic biosensor AuNPs-based was built, and immunogold colloids were prepared. The LSPR shifts were analyzed with a dark-field spectrometer.
The immunogold colloids (prepared by treating the gold surface with (1) thiol-oligo ethylene glycol and (2) 5-methylcytosine (5-mC) monoclonal antibody (mAb)), were able to detect methylation at two methylcytosine (mCpG) sites on the ctDNA with increased sensibility when compared to measurements in absence of treatment. When applied to commercial human serum samples, the nanoplasmonic biosensor detected ctDNA and also, this label-free methodology, along with the minimal amount of sample required for analysis, overcomes common issues of PCR or targeted deep sequencing techniques, classic strategies for quantification of tumor-specific mutations.
Imaging
Plasmonic-based imaging is without a doubt a very promising methodology for accessing epigenetic mechanisms. For instance, Wang et al. [76] and Zhao et al. [77, 78] showed the development of an SPR gold chip with a 3D polyrotaxane surface for detecting peptide interactions, then used for profiling histone “modification-reader” pairs in Arabidopsis thaliana, identifying several unique histone “mark-readers” by analyzing nine putative histone reader domains in this species. The comparison and analysis of histone post-transcriptional modifications (PTMs) in this study showed similarities in histone reader conservation between humans and A. thaliana. Still, the identification of plant-specific histone PTMs suggested an epigenetic adaptation or signaling in the studied species during evolution.
Hyperspectral dark-field imaging (HSDFI) has been applied with a quantitative approach, in single human cells (cell line MCF-7, primary glioblastoma multiforme cell line SF767, and cervical cancer cell line HeLa), as described in the work by Wang et al. [79]. In their work, a methodology along with the use of plasmonic nanoprobes was used to identify and locate key cytosine modifications. As plasmonic surfaces, Au- and AgNPs with 30 and 20 nm in size, respectively, were conjugated to secondary antibodies, and incubated with the cell samples. The results showed that it was possible not just to identify and quantify the most abundant form of cytosine modification (5mC), but those that account for < 0.01%, i.e., 5caC e 5fC, even at different stages of the cellular cycle. Additional advantages of applying HSDFI in this kind of sample are the high intensity of signals from the plasmonic nanoprobes and the absence of autofluorescence, a common issue in techniques such as fluorescence microscopy.
Availability of Data and Materials
All data generated or analyzed during this study are included in this published article.
References
Begcy K, Dresselhaus T (2018) Epigenetic responses to abiotic stresses during reproductive development in cereals. Plant Reprod 31:343–355
Folsom JJ, Begcy K, Hao X, Wang D, Walia H (2014) Rice fertilization-Independent Endosperm1 regulates seed size under heat stress by controlling early endosperm development. Plant Physiol 165:238–248
Enders L, Begcy K (2021) Unconventional routes to developing insect-resistant crops. Mol Plant 14:1439–1453
Lieberman-Lazarovich M, Kim T, Singh PK, Begcy K (2021) Epigenetics in horticultural crops: consequences and applications in abiotic stress tolerance. In: Stress Tolerance in Horticultural Crops. Elsevier, pp 75–90
Zhou W, Liang G, Molloy PL, Jones PA (2020) DNA methylation enables transposable element-driven genome expansion. Proc Natl Acad Sci 117:19359–19366
He L, Huang H, Bradai M, Zhao C, You Y, Ma J et al (2022) DNA methylation-free Arabidopsis reveals crucial roles of DNA methylation in regulating gene expression and development. Nat Commun 13:1335
Niederhuth CE, Schmitz RJ (2017) Putting DNA methylation in context: from genomes to gene expression in plants. Biochim Biophys Acta BBA - Gene Regul Mech 1860:149–156
Harris KD, Zemach A (2020) Contiguous and stochastic CHH methylation patterns of plant DRM2 and CMT2 revealed by single-read methylome analysis. Genome Biol 21:194
Feng S, Zhong Z, Wang M, Jacobsen SE (2020) Efficient and accurate determination of genome-wide DNA methylation patterns in Arabidopsis thaliana with enzymatic methyl sequencing. Epigenetics Chromatin 13:42
Rabinowicz PD, Citek R, Budiman MA, Nunberg A, Bedell JA, Lakey N et al (2005) Differential methylation of genes and repeats in land plants. Genome Res 15:1431–1440
Komiya R, Ikegami A, Tamaki S, Yokoi S, Shimamoto K (2008) Hd3a and RFT1 are essential for flowering in rice. Development 135:767–774
Wang M, Yuan J, Qin L, Shi W, Xia G, Liu S (2020) Ta CYP 81D5, one member in a wheat cytochrome P450 gene cluster, confers salinity tolerance via reactive oxygen species scavenging. Plant Biotechnol J 18:791–804
Steward N, Kusano T, Sano H (2000) Expression of ZmMET1, a gene encoding a DNA methyltransferase from maize, is associated not only with DNA replication in actively proliferating cells, but also with altered DNA methylation status in cold-stressed quiescent cells. Nucleic Acids Res 28:3250–3259
Sukackaite R, Grazulis S, Tamulaitis G, Siksnys V (2012) The recognition domain of the methyl-specific endonuclease McrBC flips out 5-methylcytosine. Nucleic Acids Res 40:7552–7562
Cathcart N, Chen JIL (2020) Sensing biomarkers with plasmonics. Anal Chem 92:7373–7381
Cialla-May D, Krafft C, Rösch P, Deckert-Gaudig T, Frosch T, Jahn IJ et al (2022) Raman spectroscopy and imaging in bioanalytics. Anal Chem 94:86–119
Ribeiro JA, Sales MGF, Pereira CM (2022) Electrochemistry combined-surface plasmon resonance biosensors: A review. TrAC Trends Anal Chem 157:116766
Turino M, Pazos-Perez N, Guerrini L, Alvarez-Puebla RA (2022) Positively-charged plasmonic nanostructures for SERS sensing applications. RSC Adv 12:845–859
Wang Y, Deng X, Liu J, Tang H, Jiang J (2013) Surface enhanced Raman scattering based sensitive detection of histone demethylase activity using a formaldehyde-selective reactive probe. Chem Commun 49:8489
Heck C, Michaeli Y, Bald I, Ebenstein Y (2019) Analytical epigenetics: single-molecule optical detection of DNA and histone modifications. Curr Opin Biotechnol 55:151–158
Li C, Wang Z, Wang L, Zhang C (2019) Biosensors for epigenetic biomarkers detection: a review. Biosens Bioelectron 144:111695
Barhoumi A, Halas NJ (2011) Detecting Chemically Modified DNA Bases Using Surface-Enhanced Raman Spectroscopy. J Phys Chem Lett 2:3118–3123
Le Ru EC, Etchegoin PG (2009) Principles of surface-enhanced Raman spectroscopy: and related plasmonic effects, 1st edn. Elsevier, Amsterdam; Boston
Fan M, Andrade GFS, Brolo AG (2011) A review on the fabrication of substrates for surface enhanced Raman spectroscopy and their applications in analytical chemistry. Anal Chim Acta 693:7–25
Grys D, Chikkaraddy R, Kamp M, Scherman OA, Baumberg JJ, Nijs B (2021) Eliminating irreproducibility in SERS substrates. J Raman Spectrosc 52:412–419
Wang H, Zhang T, Zhou X (2019) Dark-field spectroscopy: development, applications and perspectives in single nanoparticle catalysis. J Phys Condens Matter 31:473001
Langer J, Jimenez de Aberasturi D, Aizpurua J, Alvarez-Puebla RA, Auguié B, Baumberg JJ et al (2020) Present and future of surface-enhanced Raman scattering. ACS Nano 14:28–117
Liu H, He Y, Cao K (2021) Flexible surface-enhanced Raman scattering substrates: a review on constructions, applications, and challenges. Adv Mater Interfaces 8:2100982
Liu X, Guo J, Li Y, Wang B, Yang S, Chen W et al (2021) SERS substrate fabrication for biochemical sensing: towards point-of-care diagnostics. J Mater Chem B 9:8378–8388
Reinhard I, Miller K, Diepenheim G, Cantrell K, Hall WP (2020) Nanoparticle design rules for colorimetric plasmonic sensors. ACS Appl Nano Mater 3:4342–4350
Sun J, Lu Y, He L, Pang J, Yang F, Liu Y (2020) Colorimetric sensor array based on gold nanoparticles: design principles and recent advances. TrAC Trends Anal Chem 122:115754
Zhang Y, McKelvie ID, Cattrall RW, Kolev SD (2016) Colorimetric detection based on localised surface plasmon resonance of gold nanoparticles: merits, inherent shortcomings and future prospects. Talanta 152:410–422
Wang X, Liu G, Hu R, Cao M, Yan S, Bao Y et al (2022) Principles of surface-enhanced Raman spectroscopy. In: Principles and Clinical Diagnostic Applications of Surface-Enhanced Raman Spectroscopy. Elsevier, pp 1–32
Zeng S, Yong K-T, Roy I, Dinh X-Q, Yu X, Luan F (2011) A review on functionalized gold nanoparticles for biosensing applications. Plasmonics 6:491–506
Polte J (2015) Fundamental growth principles of colloidal metal nanoparticles – a new perspective. CrystEngComm 17:6809–6830
Joudeh N, Linke D (2022) Nanoparticle classification, physicochemical properties, characterization, and applications: a comprehensive review for biologists. J Nanobiotechnology 20:262
Ma X, He S, Qiu B, Luo F, Guo L, Lin Z (2019) Noble metal nanoparticle-based multicolor immunoassays: an approach toward visual quantification of the analytes with the naked eye. ACS Sens 4:782–791
Zanoli LM, D’Agata R, Spoto G (2012) Functionalized gold nanoparticles for ultrasensitive DNA detection. Anal Bioanal Chem 402:1759–1771
Špringer T, Ermini ML, Špačková B, Jabloňků J, Homola J (2014) Enhancing sensitivity of surface plasmon resonance biosensors by functionalized gold nanoparticles: size matters. Anal Chem 86:10350–10356
Keunen R, Macoretta D, Cathcart N, Kitaev V (2016) Stable ligand-free stellated polyhedral gold nanoparticles for sensitive plasmonic detection. Nanoscale 8:2575–2583
Zhang Z, Wang H, Chen Z, Wang X, Choo J, Chen L (2018) Plasmonic colorimetric sensors based on etching and growth of noble metal nanoparticles: Strategies and applications. Biosens Bioelectron 114:52–65
Liedberg B, Nylander C, Lunström I (1983) Surface plasmon resonance for gas detection and biosensing. Sens Actuators 4:299–304
Souto DEP, Volpe J, de Oliveira DR (2022) SPR sensors: from configurations to bioanalytical applications. In: Kubota LT, da Silva JAF, Sena MM, Alves WA (eds) Tools and Trends in Bioanalytical Chemistry. Springer International Publishing, Cham, pp 223–239
Šípová H, Homola J (2013) Surface plasmon resonance sensing of nucleic acids: a review. Anal Chim Acta 773:9–23
Nguyen AH, Sim SJ (2015) Nanoplasmonic biosensor: detection and amplification of dual bio-signatures of circulating tumor DNA. Biosens Bioelectron 67:443–449
Shao H, Lin H, Guo Z, Lu J, Jia Y, Ye M et al (2019) A multiple signal amplification sandwich-type SERS biosensor for femtomolar detection of miRNA. Biosens Bioelectron 143:111616
Fleischmann M, Hendra PJ, McQuillan AJ (1974) Raman spectra of pyridine adsorbed at a silver electrode. Chem Phys Lett 26:163–166
Albrecht MG, Creighton JA (1977) Anomalously intense Raman spectra of pyridine at a silver electrode. J Am Chem Soc 99:5215–5217
Temperini MLA, Chagas HC, Sala O (1981) Raman spectra of pyridine adsorbed on a copper electrode. Chem Phys Lett 79:75–78
Sharma B, Frontiera RR, Henry A-I, Ringe E, Van Duyne RP (2012) SERS: materials, applications, and the future. Mater Today 15:16–25
dos Santos DP, Temperini MLA, Brolo AG (2019) Intensity fluctuations in single-molecule surface-enhanced Raman scattering. Acc Chem Res 52:456–464
Villa JEL, Afonso MAS, dos Santos DP, Mercadal PA, Coronado EA, Poppi RJ (2020) Colloidal gold clusters formation and chemometrics for direct SERS determination of bioanalytes in complex media. Spectrochim Acta A Mol Biomol Spectrosc 224:117380
Goodacre R, Graham D, Faulds K (2018) Recent developments in quantitative SERS: moving towards absolute quantification. TrAC Trends Anal Chem 102:359–368
Bell SEJ, Charron G, Cortés E, Kneipp J, Chapelle ML, Langer J et al (2020) Towards reliable and quantitative surface-enhanced Raman scattering (SERS): from key parameters to good analytical practice. Angew Chem Int Ed 59:5454–5462
Morton SM, Silverstein DW, Jensen L (2011) Theoretical studies of plasmonics using electronic structure methods. Chem Rev 111:3962–3994
Adampourezare M, Hasanzadeh M, Seidi F (2022) Optical bio-sensing of DNA methylation analysis: an overview of recent progress and future prospects. RSC Adv 12:25786–25806
Sabela M, Balme S, Bechelany M, Janot J-M, Bisetty K (2017) A review of gold and silver nanoparticle-based colorimetric sensing assays. Adv Eng Mater 19:1700270
Baetsen-Young AM, Vasher M, Matta LL, Colgan P, Alocilja EC, Day B (2018) Direct colorimetric detection of unamplified pathogen DNA by dextrin-capped gold nanoparticles. Biosens Bioelectron 101:29–36
Anderson MJ, Torres-Chavolla E, Castro BA, Alocilja EC (2011) One step alkaline synthesis of biocompatible gold nanoparticles using dextrin as capping agent. J Nanoparticle Res 13:2843–2851
Su F, Wang L, Sun Y, Liu C, Duan X, Li Z (2015) Highly sensitive detection of CpG methylation in genomic DNA by AuNP-based colorimetric assay with ligase chain reaction. Chem Commun 51:3371–3374
Deng L, Liu Q, Lei C, Zhang Y, Huang Y, Nie Z et al (2020) Fluorometric and colorimetric dual-readout assay for histone demethylase activity based on formaldehyde inhibition of Ag +-triggered oxidation of O -phenylenediamine. Anal Chem 92:9421–9428
Kurdyukov S, Bullock M (2016) DNA methylation analysis: choosing the right method. Biology 5:3
Garcia-Rico E, Alvarez-Puebla RA, Guerrini L (2018) Direct surface-enhanced Raman scattering (SERS) spectroscopy of nucleic acids: from fundamental studies to real-life applications. Chem Soc Rev 47:4909–4923
Abid Hasan SM, He Y, Chang T-W, Wang J, Gartia MR (2019) Detecting DNA methylation using surface-enhanced Raman spectroscopy. J Phys Chem C 123:698–709
Luo X, Xing Y, Galvan DD, Zheng E, Wu P, Cai C et al (2019) Plasmonic gold nanohole array for surface-enhanced raman scattering detection of DNA methylation. ACS Sens 4:1534–1542
Luo X, Kang T, Zhu J, Wu P, Cai C (2020) Sensitivity-improved SERS detection of methyltransferase assisted by plasmonically engineered nanoholes array and hybridization chain reaction. ACS Sens 5:3639–3648
Mirajkar S, Dhayagude A, Maiti N, Suprasanna P, Kapoor S (2020) Distinguishing genomic DNA of Brassica juncea and Arabidopsis thaliana using surface-enhanced Raman scattering. J Raman Spectrosc 51:89–103
Turkevich J, Stevenson PC, Hillier J (1951) A study of the nucleation and growth processes in the synthesis of colloidal gold. Discuss Faraday Soc 11:55
Leopold N, Lendl B (2003) A new method for fast preparation of highly surface-enhanced Raman scattering (SERS) active silver colloids at room temperature by reduction of silver nitrate with hydroxylamine hydrochloride. J Phys Chem B 107:5723–5727
Wang Y, Wee EJH, Trau M (2016) Accurate and sensitive total genomic DNA methylation analysis from sub-nanogram input with embedded SERS nanotags. Chem Commun 52:3560–3563
Wang R (2004) Immobilisation of DNA probes for the development of SPR-based sensing. Biosens Bioelectron 20:967–974
Plácido A, Ferreira-da-Silva F, Leite JRSA, de-los-Santos-Álvarez N, Delerue-Matos C (2020) A convenient renewable surface plasmon resonance chip for relative quantification of genetically modified soybean in food and feed. PLOS ONE 15:e0229659
Grześkowiak BF, Tuśnio K, Woźniak A, Szalata M, Lipiński D, Jurga S et al (2019) Transgenic plant detection using an AuNPs based SPR biosensor. Biosensors 9:116
D’Agata R, Palladino P, Spoto G (2017) Streptavidin-coated gold nanoparticles: critical role of oligonucleotides on stability and fractal aggregation. Beilstein J Nanotechnol 8:1–11
Zhao Z, Chen Y, Xu W, Ma M (2013) Surface plasmon resonance detection of transgenic Cry1Ac cotton (Gossypium spp.). J Agric Food Chem 61:2964–9
Wang Y, Wang C, Cheng Z, Zhang D, Li S, Song L et al (2015) SPRi determination of inter-peptide interaction by using 3D supramolecular co-assembly polyrotaxane film. Biosens Bioelectron 66:338–344
Zhao S, Yang M, Zhou W, Zhang B, Cheng Z, Huang J et al (2017) Kinetic and high-throughput profiling of epigenetic interactions by 3D-carbene chip-based surface plasmon resonance imaging technology. Proc Natl Acad Sci 114
Zhao S, Zhang B, Yang M, Zhu J, Li H (2018) Systematic profiling of histone readers in Arabidopsis thaliana. Cell Rep 22:1090–1102
Wang X, Cui Y, Irudayaraj J (2015) Single-cell quantification of cytosine modifications by hyperspectral dark-field imaging. ACS Nano 9:11924–11932
Acknowledgements
The authors thank the Laboratory of Molecular Spectroscopy of IQ-USP – São Paulo – Brazil for the technical assistance with the SERS spectra used to generate Fig. 3 (FAPESP, grant 2016/21070-5).
Funding
This work was supported by the Brazilian agency CNPq (Grant 405087/2021–7) to MBML and the USDA National Institute of Food and Agriculture NIFA-AFRI program (Grant 2023–67013-39412) to KB.
Author information
Authors and Affiliations
Contributions
Mónica Benicia Mamián-López and Kevin Begcy conceived the idea, wrote and edited the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mamián-López, M.B., Begcy, K. Plasmonics: An Optical Approach to Study Plant Epigenetics. Plasmonics 19, 687–697 (2024). https://doi.org/10.1007/s11468-023-02000-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11468-023-02000-x