Abstract
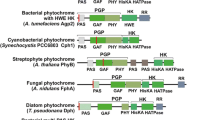
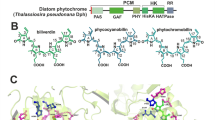
The phytochrome gene from Microcystis aeruginosa FACHB-912, isolated from algal blooms in Taihu Lake in China, was sequenced and found to code for a wavelength transition light receptor protein. The gene was composed of five distinct domains: a PAS fold domain, a GAF domain, a phytochrome domain, a His Kinase A (phosphoacceptor) domain, and a histidine kinase domain. Red-absorbing phytochrome, far-red-absorbing phytochrome, and photoconversion kinetics were monitored via the spectral characteristics of the protein. To detect positive selection, homologous sequences from cyanobacterium phytochrome genes were obtained from GenBank. Potential selective pressure was identified by phylogenetic analysis with maximum likelihood analyses. The frequency of the phytochrome gene being subjected to selective constraints suggests that positive selection is a potentially important mechanism that promotes the evolution of the cyanobacteria phytochrome gene. The sites identified in this study provide targets for further research on the structural-functional role of these residues, and on the correlation with the mechanism of algal blooms.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Quail P H, Boylan M T, Parks B M, et al. Phytochromes: Photosensory perception and signal transduction. Science, 1995, 268: 675–680
Smith H. Phytochromes and light signal perception by plants-an emerging synthesis. Nature, 2000, 407: 585–591
Ikeda H, Fujii N, Setoguchi H. Molecular evolution of phytochromes in Cardamine nipponica (Brassicaceae) suggests the involvement of PHYE in local adaptation. Genetics, 2009, 182: 603–614
Giraud E, Vermeglio A. Bacteriophytochromes in anoxygenic photosynthetic bacteria. Photosynth Res, 2008, 97: 141–153
Lamparter T. Evolution of cyanobacterial and plant phytochromes. FEBS Lett, 2004, 573: 1–5
Kevei E, Nagy F. Phytochrome controlled signalling cascades in higher plants. Physiol Plant, 2003, 117: 305–313
Muse S V, Gaut B S. A likelihood approach for comparing synonymous and nonsynonymous nucleotide substitution rates, with application to the chloroplast genome. Mol Biol Evol, 1994, 11: 715–724
Nielsen R, Yang Z. Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics, 1998, 148: 929–936
Wong W S, Yang Z, Goldman N, et al. Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics, 2004, 168: 1041–1051
Yang Z, Nielsen R, Goldman N, et al. Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics, 2000, 155: 431–449
Yang Z. PAML: A program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci, 1997, 13: 555–556
Yang Z, Nielsen R. Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol, 2002, 19: 908–917
Demarsac N T, Houmard J. Adaptation of cyanobacteria to environmental stimuli-new steps towards molecular mechanisms. Fems Microbiol Rev, 1993, 104: 119–189
Pichardo S, Jos A, Zurita J, et al. Toxic effects produced by microcystins from a natural cyanobacterial bloom and a Microcystis aeruginosa isolated strain on the fish cell lines RTG-2 and PLHC-1. Arch Environ Contam Toxicol, 2006, 51: 86–96
Carre-Mlouka A, Mejean A, Quillardet P, et al. A new rubisco-like protein coexists with a photosynthetic rubisco in the planktonic cyanobacteria Microcystis. J Biol Chem, 2006, 281: 24462–24471
Landgraf F T, Forreiter C, Hurtado Pico A, et al. Recombinant holophytochrome in Escherichia coli. FEBS Lett, 2001, 508: 459–462
Edgar R C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res, 2004, 32: 1792–1797
Edgar R C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics, 2004, 5: 113
Larkin M A, Blackshields G, Brown N P, et al. Clustal W and clustal X version 2.0. Bioinformatics, 2007, 23: 2947–2948
Tamura K, Dudley J, Nei M, et al. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol, 2007, 24: 1596–1599
Felsenstein J. Mathematics vs. evolution: Mathematical evolutionary theory. Science, 1989, 246: 941–942
Aravind L, Ponting C P. The GAF domain: An evolutionary link between diverse phototransducing proteins. Trends Biochem Sci, 1997, 22: 458–459
Karniol B, Wagner J R, Walker J M, et al. Phylogenetic analysis of the phytochrome superfamily reveals distinct microbial subfamilies of photoreceptors. Biochem J, 2005, 392: 103–116
Sharda S, Shah R, Gartner W. Domain interaction in cyanobacterial phytochromes as a prerequisite for spectral integrity. Eur Biophys J, 2007, 36: 815–821
Karniol B, Vierstra R D. The pair of bacteriophytochromes from Agrobacterium tumefaciens are histidine kinases with opposing photobiological properties. Proc Natl Acad Sci USA, 2003, 100: 2807–2812
Bhoo S H, Davis S J, Walker J, et al. Bacteriophytochromes are photochromic histidine kinases using a biliverdin chromophore. Nature, 2001, 414: 776–779
Yeh K C, Wu S H, Murphy J T, et al. A cyanobacterial phytochrome two-component light sensory system. Science, 1997, 277: 1505–1508
Giraud E, Vermeglio A. Bacteriophytochromes in anoxygenic photosynthetic bacteria. Photosynth Res, 2008, 97: 141–153
Taylor B L, Zhulin I B. PAS domains: Internal sensors of oxygen, redox potential, and light. Microbiol Mol Biol Rev, 1999, 63: 479–506
Evans K, Fordham-Skelton A P, Mistry H, et al. A bacteriophytochrome regulates the synthesis of LH4 complexes in Rhodopseudomonas palustris. Photosynth Res, 2005, 85: 169–180
Mathews S, Burleigh J G, Donoghue M J. Adaptive evolution in the photosensory domain of phytochrome A in early angiosperms. Mol Biol Evol, 2003, 20: 1087–1097
Balasubramanian S, Sureshkumar S, Agrawal M, et al. The PHYTOCHROME C photoreceptor gene mediates natural variation in flowering and growth responses of Arabidopsis thaliana. Nat Genet, 2006, 38: 711–715
Filiault D L, Wessinger C A, Dinnery J R, et al. Amino acid polymorphisms in Arabidopsis phytochrome B cause differential responses to lights (vol 105, pg 3157, 2008). Proc Natl Acad Sci USA, 2008, 105: 8482
Maloof J N, Borevitz J O, Dabi T, et al. Natural variation in light sensitivity of Arabidopsis. Nat Genet, 2001, 29: 441–446
Ingvarsson P K, Garcia M V, Hall D, et al. Clinal variation in phyB2, a candidate gene for day-length-induced growth cessation and bud set, across a latitudinal gradient in European aspen (Populus tremula). Genetics, 2006, 172: 1845–1853
Ingvarsson P K, Garcia M V, Luquez V, et al. Nucleotide polymorphism and phenotypic associations within and around the phytochrome B2 Locus in European aspen (Populus tremula, Salicaceae). Genetics, 2008, 178: 2217–2226
Mathews S, McBreen K. Phylogenetic relationships of B-related phytochromes in the Brassicaceae: Redundancy and the persistence of phytochrome D. Mol Phylogenet Evol, 2008, 49: 411–423
Mathews S, Sharrock R A. The phytochrome gene family in grasses (Poaceae): A phylogeny and evidence that grasses have a subset of the loci found in dicot angiosperms. Mol Biol Evol, 1996, 13: 1141–1150
White G M, Hamblin M T, Kresovich S. Molecular evolution of the phytochrome gene family in sorghum: Changing rates of synonymous and replacement evolution. Mol Biol Evol, 2004, 21: 716–723
Author information
Authors and Affiliations
Corresponding author
Additional information
These authors contributed equally to this work.
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Shi, F., Li, N., Liu, S. et al. Sequence analysis of the Microcystis aeruginosa FACHB-912 phytochrome gene supports positive selection in cyanobacteria. Chin. Sci. Bull. 57, 3270–3275 (2012). https://doi.org/10.1007/s11434-012-5238-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-012-5238-4