Abstract
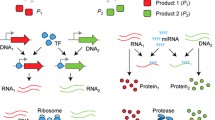
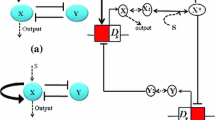
In gene regulatory networks, gene regulation loops often occur with multiple positive feedback, multiple negative feedback and coupled positive and negative feedback forms. In above gene regulation loops, auto-activation loops are ubiquitous regulatory motifs. This paper aims to investigate a two-component dual-positive feedback genetic circuit, which consists of a double negative feedback circuit and an additional positive feedback loop (APFL). We study effect of substrate concentration on gene expression in the single and the networked systems with APFLs, respectively. We find that substrate concentration can tune stochastic switch behavior in the signal system and then we explore relationship of substrate concentration with positive feedback strength in aspect of stochastic switch behavior. Furthermore, we also discuss gene expression and stochastic switch behavior in the networked systems with APFLs. Based on analysis in the networked systems, we discover that genes express in some specific cells and do not express in the other cells when the expression achieves its steady state. These results can be used to well explain the character of regionalization in the expression of genes and the phenomenon of gene differentiation.
Similar content being viewed by others
References
Alon U. An Introduction to Systems Biology: Design Principles of Biological Circuits. London: Chapman and Hall, CRC, 2006
Chen L, Wang L, Zhang X. Biomolecular Networks: Methods and Applications in Systems Biology. Hoboken, New Jersey: Wiley, 2009
Lei J. Systems Biology-Modeling, Analysis, Simulation. Shanghai: Shanghai Science and Technology Press, 2010
Zhou T. The Stochastic Dynamics of Biological Systems. Beijing: Science Press, 2009
Liu Z, Wang R, Yang L, et al. Construction and Analysis of Biomolecular Networks. Beijing: Science Press, 2012
Hasty J, Dolnik M, Rottschäfer V, et al. Synthetic gene network for entraining and amplifying cellular oscillations. Phys Rev Lett, 2002, 88: 148101
Wang P, Lü J, Wan L, et al. A stochastic simulation algorithm for biochemical reactions with delays. In: Proceedings of the 2013 IEEE International Conference on Systems Biology. Huangshan: IEEE, 2013. 109–114
Brandman O, Ferrell J E, Li R, et al. Interlinked fast and slow positive feedback loops drive reliable cell decisions. Science, 2005, 310: 496–498
Wiener N, Schade J P. Progress in Biocybernetics. New York: Elsevier Publishing Company, 1966
Pigliucci M, Murren C J. Perspective: Genetic assimilation and a possible evolutionary paradox: Can macroevolution sometimes be so fast as to pass us by? Evolution, 2013, 57: 1455–1464
Raser J M, O’Shea E K. Noise in gene expression: origins, consequences, and control. Science, 2005, 309: 2010–2013
Yuan Z, Zhang J, Zhou T. Noise induced continuous switching. Sci China Ser B-Chem, 2007, 51: 562–569
Zhang J, Wang J, Yuan Z, et al. Noise-induced synchronized switching of a multicellular system. Progr Biochem Biophys, 2008, 35: 929–939
Wartlick O, Kicheva A, González-Gaitán M. Morphogen gradient formation. Cold Spring Harbor Perspectives Biol, 2009, 1: a001255
Meinhardt H. Models for the generation and interpretation of gradients. Cold Spring Harbor Perspectives Biol, 2009, 1: a001362
Meinhardt H. Space-dependent cell determination under the control of a morphogen gradient. J Theor Biol, 1978, 74: 307–321
Meinhardt H. Models of Biological Pattern Formation. London: Academic Press, 1982
Lopes F J P, Vieira F M C, Holloway D M, et al. Spatial bistability generates hunchback expression sharpness in the drosophila embryo. PLoS Comput Biol, 2008, 4: e1000184
Freidlin M I, Wentzell A D. Random Perturbations of Dynamical Systems. 2nd ed. New York: Springer, 1998
Ferrell Jr J E. Self-perpetuating states in signal transduction: positive feedback, double-negative feedback and bistability. Curr Opin Cell Biol, 2002, 14: 140–148
Zhang L, Radtke K, Zheng L, et al. Noise drives sharpening of gene expression boundaries in the zebrafish hindbrain. Mol Syst Biol, 2012, 8: 613
Balázsi G, van Oudenaarden A, Collins J J. Cellular decision making and biological noise: From microbes to mammals. Cell, 2011, 144: 910–925
Hart Y, Reich-Zeliger S, Antebi Y E, et al. Paradoxical signaling by a secreted molecule leads to homeostasis of cell levels. Cell, 2014, 158: 1022–1032
Wang P, Zhang Y, Lü J, et al. Functional characteristics of additional positive feedback in genetic circuits. NOnlinear Dyn, 2015, 79: 397–408
Tessone C J, Mirasso C R, Toral R, et al. Diversity-induced resonance. Phys Rev Lett, 2006, 97: 194101
Giudicelli F, Taillebourg E, Charnay P, et al. Krox-20 patterns the hindbrain through both cell-autonomous and non cell-autonomous mechanisms. Genes Dev, 2001, 15: 567–580
Kampen N G. Stochastic Processes in Physics and Chemistry. New York: North Holland, 2007
Wang J, Zhang J, Yuan Z, et al. Noise-induced switches in network systems of the genetic toggle switch. BMC Syst Biol, 2007, 1: 50
White R J, Schilling T F. How degrading: Cyp26s in hindbrain development. Dev Dyn, 2008, 237: 2775–2790
Wang P, Lv J H. Control of genetic regulatory networks: Opportunities and challenges. Acta Autom Sin, 2013, 39: 1969–1979
Mitra S, Das R, Hayashi Y. Genetic networks and soft computing. IEEE/ACM Trans Comput Biol Bioinf, 2011, 8: 94–107
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, L., Wang, P. & Lü, J. Substrate concentration effect on gene expression in genetic circuits with additional positive feedback. Sci. China Technol. Sci. 61, 1175–1183 (2018). https://doi.org/10.1007/s11431-018-9301-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11431-018-9301-0