Abstract
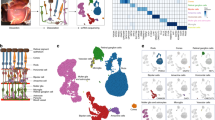
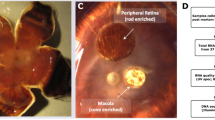
The human retina serves as a light detector and signals transmission tissue. Advanced insights into retinal disease mechanisms and therapeutic strategies require a deep understanding of healthy retina molecular events. Here, we sequenced the mRNA of over 0.6 million single cells from human retinas across six regions at nine different ages. Sixty cell sub-types have been identified from the human mature retinas with unique markers. We revealed regional and age differences of gene expression profiles within the human retina. Cell-cell interaction analysis indicated a rich synaptic connection within the retinal cells. Gene expression regulon analysis revealed the specific expression of transcription factors and their regulated genes in human retina cell types. Some of the gene’s expression, such as DKK3, are elevated in aged retinas. A further functional investigation suggested that over expression of DKK3 could impact mitochondrial stability. Overall, decoding the molecular dynamic architecture of the human retina improves our understanding of the vision system.
Similar content being viewed by others
References
Aibar, S., González-Blas, C.B., Moerman, T., Huynh-Thu, V.A., Imrichova, H., Hulselmans, G., Rambow, F., Marine, J.C., Geurts, P., Aerts, J., et al. (2017). SCENIC: single-cell regulatory network inference and clustering. Nat Methods 14, 1083–1086.
Baffet, A.D., Hu, D.J., and Vallee, R.B. (2015). Cdk1 activates pre-mitotic nuclear envelope dynein recruitment and apical nuclear migration in neural stem cells. Dev Cell 33, 703–716.
Bonnel, S., Mohand-Said, S., and Sahel, J.A. (2003). The aging of the retina. Exp Gerontol 38, 825–831.
Boycott, B.B., and Wässle, H. (1991). Morphological classification of bipolar cells of the primate retina. Eur J Neurosci 3, 1069–1088.
Clark, B.S., Stein-O’Brien, G.L., Shiau, F., Cannon, G.H., Davis-Marcisak, E., Sherman, T., Santiago, C.P., Hoang, T.V., Rajaii, F., James-Esposito, R.E., et al. (2019). Single-cell RNA-Seq analysis of retinal development identifies NFI factors as regulating mitotic exit and late-born cell specification. Neuron 102, 1111–1126.e5.
Correia-Melo, C., Marques, F.D.M., Anderson, R., Hewitt, G., Hewitt, R., Cole, J., Carroll, B.M., Miwa, S., Birch, J., Merz, A., et al. (2016). Mitochondria are required for pro-ageing features of the senescent phenotype. EMBO J 35, 724–742.
Cowan, C.S., Renner, M., De Gennaro, M., Gross-Scherf, B., Goldblum, D., Hou, Y., Munz, M., Rodrigues, T.M., Krol, J., Szikra, T., et al. (2020). Cell types of the human retina and its organoids at single-cell resolution. Cell 182, 1623–1640.e34.
Deng, Y., Qiao, L., Du, M., Qu, C., Wan, L., Li, J., and Huang, L. (2022). Age-related macular degeneration: epidemiology, genetics, pathophysiology, diagnosis, and targeted therapy. Genes Dis 9, 62–79.
Fan, Z., Beresford, P.J., Zhang, D., and Lieberman, J. (2002). HMG2 interacts with the nucleosome assembly protein SET and is a target of the cytotoxic T-lymphocyte protease granzyme A. Mol Cell Biol 22, 2810–2820.
Fletcher, A.E. (2010). Free radicals, antioxidants and eye diseases: evidence from epidemiological studies on cataract and age-related macular degeneration. Ophthal Res 44, 191–198.
Harman, A., Abrahams, B., Moore, S., and Hoskins, R. (2000). Neuronal density in the human retinal ganglion cell layer from 16–77 years. Anat Rec 260, 124–131.
Harman, D. (1981). The aging process. Proc Natl Acad Sci USA 78, 7124–7128.
Hoshino, A., Ratnapriya, R., Brooks, M.J., Chaitankar, V., Wilken, M.S., Zhang, C., Starostik, M.R., Gieser, L., La Torre, A., Nishio, M., et al. (2017). Molecular anatomy of the developing human retina. Dev Cell 43, 763–779.e4.
Hu, Y., Wang, X., Hu, B., Mao, Y., Chen, Y., Yan, L., Yong, J., Dong, J., Wei, Y., Wang, W., et al. (2019). Dissecting the transcriptome landscape of the human fetal neural retina and retinal pigment epithelium by single-cell RNA-seq analysis. PLoS Biol 17, e3000365.
Huang, L., Chen, Y., Lin, Y., Tam, P.O.S., Cheng, Y., Shi, Y., Gong, B., Lu, F., Yang, J., Wang, H., et al. (2019). Genome-wide analysis identified 17 new loci influencing intraocular pressure in Chinese population. Sci China Life Sci 62, 153–164.
Huang, L., Fang, L., Liu, Q., Torshizi, A.D., and Wang, K. (2022). Integrated analysis on transcriptome and behaviors defines HTT repeat-dependent network modules in Huntington’s disease. Genes Dis 9, 479–493.
Huang, L., Zhang, H., Cheng, C.Y., Wen, F., Tam, P.O.S., Zhao, P., Chen, H., Li, Z., Chen, L., Tai, Z., et al. (2016). A missense variant in FGD6 confers increased risk of polypoidal choroidal vasculopathy. Nat Genet 48, 640–647.
Isik, S., Zaim, M., Yildiz, M.T., Negis, Y., Kunduraci, T., Karakas, N., Arikan, G., and Cetin, G. (2015). DNA topoisomerase IIβ as a molecular switch in neural differentiation of mesenchymal stem cells. Ann Hematol 94, 307–318.
Jin, S., Guerrero-Juarez, C.F., Zhang, L., Chang, I., Ramos, R., Kuan, C.H., Myung, P., Plikus, M.V., and Nie, Q. (2021). Inference and analysis of cell-cell communication using CellChat. Nat Commun 12, 1088.
Kaneko, A. (1970). Physiological and morphological identification of horizontal, bipolar and amacrine cells in goldfish retina. J Physiol 207, 623–633.
Kolb, H., Fernandez, E., and Nelson, R. (2020). Webvision: The Organization of the Retina and Visual System. 1–1839.
Kolb, H., Linberg, K.A., and Fisher, S.K. (1992). Neurons of the human retina: a Golgi study. J Comp Neurol 318, 147–187.
Kummer, E., and Ban, N. (2021). Mechanisms and regulation of protein synthesis in mitochondria. Nat Rev Mol Cell Biol 22, 307–325.
Lake, B.B., Ai, R., Kaeser, G.E., Salathia, N.S., Yung, Y.C., Liu, R., Wildberg, A., Gao, D., Fung, H.L., Chen, S., et al. (2016). Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590.
Li, W., Cheng, H., Li, G., and Zhang, L. (2020). Mitochondrial damage and the road to exhaustion. Cell Metab 32, 905–907.
Lian, G., Wong, T., Lu, J., Hu, J., Zhang, J., and Sheen, V. (2019). Cytoskeletal associated filamin A and RhoA affect neural progenitor specification during mitosis. Cerebral Cortex 29, 1280–1290.
Liao, N., Li, C., Jiang, H., Fang, A., Zhou, S., and Wang, Q. (2016). Neovascular glaucoma: a retrospective review from a tertiary center in China. BMC Ophthalmol 16, 14.
Lin, M.T., and Beal, M.F. (2006). Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443, 787–795.
Lu, Y., Shiau, F., Yi, W., Lu, S., Wu, Q., Pearson, J.D., Kallman, A., Zhong, S., Hoang, T., Zuo, Z., et al. (2020). Single-cell analysis of human retina identifies evolutionarily conserved and species-specific mechanisms controlling development. Dev Cell 53, 473–491.e9.
Lukowski, S.W., Lo, C.Y., Sharov, A.A., Nguyen, Q., Fang, L., Hung, S.S., Zhu, L., Zhang, T., Grünert, U., Nguyen, T., et al. (2019). A single-cell transcriptome atlas of the adult human retina. EMBO J 38, e100811.
Macneil, M.A., Heussy, J.K., Dacheux, R.F., Raviola, E., and Masland, R. H. (1999). The shapes and numbers of amacrine cells: matching of photofilled with Golgi-stained cells in the rabbit retina and comparison with other mammalian species. J Comp Neurol 413, 305–326.
MacNeil, M.A., and Masland, R.H. (1998). Extreme diversity among amacrine cells: implications for function. Neuron 20, 971–982.
Macosko, E.Z., Basu, A., Satija, R., Nemesh, J., Shekhar, K., Goldman, M., Tirosh, I., Bialas, A.R., Kamitaki, N., Martersteck, E.M., et al. (2015). Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214.
Masland, R. (2001). Neuronal diversity in the retina. Curr Opin Neurobiol 11, 431–436.
Masland, R.H. (2012). The neuronal organization of the retina. Neuron 76, 266–280.
Mellough, C.B., Bauer, R., Collin, J., Dorgau, B., Zerti, D., Dolan, D.W.P., Jones, C.M., Izuogu, O.G., Yu, M., Hallam, D., et al. (2019). An integrated transcriptional analysis of the developing human retina. Development 146.
Nomura, Y., Mulavara, A.P., Richards, J.T., Brady, R., and Bloomberg, J.J. (2005). Optic flow dominates visual scene polarity in causing adaptive modification of locomotor trajectory. Cogn Brain Res 25, 624–631.
Novichkov, P.S., Rodionov, D.A., Stavrovskaya, E.D., Novichkova, E.S., Kazakov, A.E., Gelfand, M.S., Arkin, A.P., Mironov, A.A., and Dubchak, I. (2010). RegPredict: an integrated system for regulon inference in prokaryotes by comparative genomics approach. Nucl Acids Res 38, W299–W307.
Peng, Y.R., Shekhar, K., Yan, W., Herrmann, D., Sappington, A., Bryman, G.S., van Zyl, T., Do, M.T.H., Regev, A., and Sanes, J.R. (2019). Molecular classification and comparative taxonomics of foveal and peripheral cells in primate retina. Cell 176, 1222–1237.e22.
Reichenbach, A., and Bringmann, A. (2020). Glia of the human retina. Glia 68, 768–796.
Shen, Y., Shen, H., Guo, D., Sun, X., Sun, Y., Hong, N., Wang, X., Xie, C., Zhao, Y., He, Q., et al. (2020). Recent developments in regenerative ophthalmology. Sci China Life Sci 63, 1450–1490.
Stephenson, J., Nutma, E., van der Valk, P., and Amor, S. (2018). Inflammation in CNS neurodegenerative diseases. Immunology 154, 204–219.
Szél, A., Lukáts, A., Fekete, T., Szepessy, Z., and Röhlich, P. (2000). Photoreceptor distribution in the retinas of subprimate mammals. J Opt Soc Am A 17, 568–579.
Szklarczyk, D., Franceschini, A., Kuhn, M., Simonovic, M., Roth, A., Minguez, P., Doerks, T., Stark, M., Muller, J., Bork, P., et al. (2011). The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucl Acids Res 39, D561–D568.
Takahashi, K., and Yamanaka, S. (2006). Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676.
Ugrinova, I., Pashev, I.G., and Pasheva, E.A. (2009). Nucleosome binding properties and Co-remodeling activities of native and in vivo acetylated HMGB-1 and HMGB-2 proteins. Biochemistry 48, 6502–6507.
Ungvari, Z., Tarantini, S., Kiss, T., Wren, J.D., Giles, C.B., Griffin, C.T., Murfee, W.L., Pacher, P., and Csiszar, A. (2018). Endothelial dysfunction and angiogenesis impairment in the ageing vasculature. Nat Rev Cardiol 15, 555–565.
Vasiljevic, A., Champier, J., Figarella-Branger, D., Wierinckx, A., Jouvet, A., and Fèvre-Montange, M. (2013). Molecular characterization of central neurocytomas: potential markers for tumor typing and progression. Neuropathology 33, 149–161.
Wang, S., Poli, S., Liang, X., and Peng, G.H. (2021a). Longitudinal single-cell RNA-seq of hESCs-derived retinal organoids. Sci China Life Sci 64, 1661–1676.
Wang, S., Zheng, Y., Li, Q., He, X., Ren, R., Zhang, W., Song, M., Hu, H., Liu, F., Sun, G., et al. (2021b). Deciphering primate retinal aging at single-cell resolution. Protein Cell 12, 889–898.
Wei, Y.H., Ma, Y.S., Lee, H.C., Lee, C.F., and Lu, C.Y. (2001). Mitochondrial theory of aging matures-roles of mtDNA mutation and oxidative stress in human aging. Chin Med J 64, 259–270.
Whitmore, S.S., Wagner, A.H., DeLuca, A.P., Drack, A.V., Stone, E.M., Tucker, B.A., Zeng, S., Braun, T.A., Mullins, R.F., and Scheetz, T.E. (2014). Transcriptomic analysis across nasal, temporal, and macular regions of human neural retina and RPE/choroid by RNA-Seq. Exp Eye Res 129, 93–106.
Yang, Z.R., Dong, W.G., Lei, X.F., Liu, M., and Liu, Q.S. (2012). Overexpression of Dickkopf-3 induces apoptosis through mitochondrial pathway in human colon cancer. World J Gastroenterol 18, 1590–1601.
Yaqoob, P. (2017). Ageing alters the impact of nutrition on immune function. Proc Nutr Soc 76, 347–351.
Zepeda-Romero, L.C., Vazquez-Membrillo, M., Adan-Castro, E., Gomez-Aguayo, F., Gutierrez-Padilla, J.A., Angulo-Castellanos, E., Barrera de Leon, J.C., Gonzalez-Bernal, C., Quezada-Chalita, M.A., Meza-Anguiano, A., et al. (2017). Higher prolactin and vasoinhibin serum levels associated with incidence and progression of retinopathy of prematurity. Pediatr Res 81, 473–479.
Zhang, F., Kurokawa, K., Lassoued, A., Crowell, J.A., and Miller, D.T. (2019). Cone photoreceptor classification in the living human eye from photostimulation-induced phase dynamics. Proc Natl Acad Sci USA 116, 7951–7956.
Zhao, B., Liu, P., Fukumoto, T., Nacarelli, T., Fatkhutdinov, N., Wu, S., Lin, J., Aird, K.M., Tang, H.Y., Liu, Q., et al. (2020). Topoisomerase 1 cleavage complex enables pattern recognition and inflammation during senescence. Nat Commun 11, 908.
Zhou, S., Zhu, Y., Mashrah, M., Zhang, X., He, Z., Yao, Z., Zhang, C., Guo, F., Hu, Y., and Zhang, C. (2017). Expression pattern of DKK3, dickkopf WNT signaling pathway inhibitor 3, in the malignant progression of oral submucous fibrosis. Oncol Rep 37, 979–985.
Zhou, Y., Zhou, B., Pache, L., Chang, M., Khodabakhshi, A.H., Tanaseichuk, O., Benner, C., and Chanda, S.K. (2019). Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10, 1523.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81790643, 81970839, 82271105, 82121003), the Sichuan Science and Technology Program (2021YFS0033, 2021YFS0369, 2021YFS0404, 2021JDGD0036) and the Chinese Academy of Medical Sciences (CAMS) Innovation Fund for Medical Sciences (2019-I2M-5-032).
Author information
Authors and Affiliations
Corresponding author
Additional information
Compliance and ethics
The author(s) declare that they have no conflict of interest.
Supporting Information
Rights and permissions
About this article
Cite this article
Huang, L., Li, R., Ye, L. et al. Deep Sc-RNA sequencing decoding the molecular dynamic architecture of the human retina. Sci. China Life Sci. 66, 496–515 (2023). https://doi.org/10.1007/s11427-021-2163-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-021-2163-1