Abstract
The retina is a stratified layer of sensory neurons lining the posterior portion of the eye. In humans, fine detail and color vision are enabled by the macula, a central region of the retina dense in cone photoreceptors (PRs). Achromatic low light and peripheral vision are facilitated by rod PRs found with increasing density outside the macula in the peripheral retina. The outer retina is nourished by choroidal blood flow regulated by a single layer of intervening retinal pigment epithelial (RPE) cells. Existing human retinal transcriptome projects have been critical for studying aspects of retinal development and disease however, there are currently no publicly available data sets accurately describing the aging human central retina, peripheral retina, and supporting RPE/choroid. Here we used Illumina RNA sequencing (RNA-seq) analysis to characterize the mRNA transcriptome of rod and cone PR-enriched human retina as well as supporting macular RPE/choroid tissue. These data will be valuable to the vision research community for characterizing global changes in gene expression in clinically relevant ocular tissues.
Measurement(s) | RNA • transcriptome |
Technology Type(s) | RNA sequencing |
Factor Type(s) | type of ocular tissue |
Sample Characteristic - Organism | Homo sapiens |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.12464540
Similar content being viewed by others
Background & Summary
Since the emergence of commercial Next Generation Sequencing (NGS) technology ~13 years ago, genomics data represents one of the most proliferative Big Data domains with unprecedented growth projected by the year 20251. Once made publicly available, genome-wide experiments provide the research community with valuable data that can be subsequently mined to further scientific knowledge. Beyond submission to public repositories, detailed curation of these datasets is critical for accurate interpretation and repurposing of NGS data. NGS technology has allowed for extensive eukaryotic transcriptome analysis using a wide range of tissues and single cells harvested from a variety of species2,3. These RNA-seq analyses have become the gold standard for in depth characterization of global differences in transcript expression as well as the accumulation of novel tissue and cell-specific transcript isoforms.
The vision research community has greatly benefited over the past decade from a deluge of RNA-seq and associated NGS data describing the neural retina and its supporting tissues4,5. Non-mammalian models such as zebrafish and chickens have been successfully employed to characterize global changes in chromatin organization6,7,8 and associated transcriptional networks9,10,11 during vertebrate retinal development. Mammalian models, particularly the mouse, have been more extensively used to integrate retinal transcriptome and chromatin organization NGS data sets12,13,14,15. Collectively, these data have been critical for the determination of chromatin states required for cell-type specific transcription in vertebrate retinal neurons as well as for modeling rare retinal degenerative diseases. Ultimately though, detailed aspects of human retinal development as well as genetically complex human retinal diseases such as age-related macular degeneration (AMD) have proven difficult to model in non-human animals.
The experiment described here is part of a larger ongoing project within the James Madison University’s Center for Genome & Metagenome Studies (CGEMS) investigating transcriptional regulation in the developing, mature, aging, and diseased vertebrate retina. Within the human retina, fine detail and color vision required for reading, facial recognition, and many other day-to-day activities are enabled by the macula, a 5.5 mm central region dense in cone photoreceptors (PRs). Achromatic low light and peripheral vision are facilitated by rod PRs found with sharply increasing density outside of the macula moving toward the peripheral regions of the retina16. Rod and cone PR neurons orchestrate cell-specific transcriptional networks critical for differentiation and proper function17. Farkis and colleagues completed the first comprehensive RNA-seq analysis of whole retina tissue collected from three adult human donors18. This analysis provided initial insight into the complexity of the whole human retinal transcriptome including novel exons and novel transcripts expressed in whole retinal tissue. A subsequent study sampling central and peripheral regions of eight adult human retinas hinted at transcriptional networks controlling rod and cone-specific PR function19. However, a 8 mm region of the central retina was collected in this study extending to an area 1.5X beyond the anatomical macular circumference and thereby resulting in rods representing the majority PR cell type in both sample groups. Additionally, Li and colleagues report limited read quality and sampling metrics in their study.
The RNA-seq experiment described here characterizes several aspects of the human retina mRNA transcriptome. To investigate transcriptional networks specific to retinal neurons, whole corneas were analyzed as a non-neuronal ocular control tissue (Fig. 1A) for comparison to retinal samples (Fig. 1B). Additionally, to more accurately investigate cone and rod-specific cell type-restricted transcriptional regulation, 3 mm and 6 mm samples of the central and peripheral retina were analyzed respectively (Fig. 1C). Finally, this study also analyzes retinal pigment epithelia (RPE)/choroid tissue adjacent to the 3 mm central retina (Fig. 1C). The RPE is a single cell layer that functions as the barrier between the outer retina and the choroid retinal supply blood20. Together, the macular retina and adjacent RPE are the primary sites of pathology associated with the complex retinal disorder AMD. RNAs extracted from these tissues were subjected to a rigorous workflow for robust and accurate analysis of mRNA transcriptional networks in clinically relevant ocular cell and tissue types that will be valuable to our research group’s future studies as well as the vision research community (Fig. 1D).
Overview of human donor eye dissection, sample collection and downstream analysis. Pairs of whole globe human donor eyes were procured and dissected so that whole corneas were collected (A) prior to making radial cuts down the side of the globe to expose the retina (B). The central retina and adjacent RPE/choroid layers were collected using a 3 mm biopsy punch in the center of each retina while the rod-rich peripheral retina was collected using a 6 mm biopsy punch (C). A rigorous experimental workflow was applied to analyze the mRNA transcriptome of these human ocular samples (D).
Methods
Tissue collection and processing
This study conformed to Institutional Review Board regulations for use of human tissues at James Madison University (JMU). Informed consent from the donor next-of-kin was collected prior to tissue collection. Left and right pairs of whole globe human donor eyes were curated from the National Disease Research Interchange (NDRI; Philadelphia, PA; Fig. 1a; Table 1, Online-only Table 1). Corneas from each eye were collected by making circular cuts along the limbus. Liberated corneal tissue was rinsed in HBBS -Ca, -Mg, blotted dry and immediately flash frozen and ground into a fine powder using a mortar and pestle super cooled with dry ice. Retinas were exposed by making 4 radial cuts down the side of globes from anterior to posterior, laying the dissected eye flat, and removing the vitreous with forceps (Fig. 1b). A central region of the retina was collected using a 3 mm biopsy punch centered on the macula (Integra Miltex, Rietheim-Weilheim, Germany). After peeling off retinal tissue from 3 mm biopsies, the RPE/choroid layer immediately beneath the central retina was separately collected from each specimen herein referred to as RPE/choroid. Rod-rich retinas were similarly collected from regions of the peripheral retina using a 6 mm biopsy punch (Fig. 1c). Tissues were briefly rinsed in cold HBSS -Ca, -Mg and transferred to 1.5 mL tubes containing RLT + lysis buffer (Qiagen; AllPrep kit) supplemented with 2-Mercaptoethanol (Sigma) and vortexed vigorously to dissociate and lyse the tissue. Ground corneas were similarly transferred to RLT+/BME lysis buffer solution and vortexed. Samples were stored in lysis buffer at −80 °C. Similar tissues from left and right eyes from each donor were pooled into single samples. Whole globes and dissected eyes were imaged using a Leica M80 high performance stereomicroscope equipped with an IC80HD camera. Tissues were collected from donors between the ages of 68–95 years and processed within 50 hours of donor death (Online-only Table 1).
Total RNA isolation
Total RNA was extracted from 37 human ocular tissues using a Qiagen AllPrep Mini Kit (Hilden, Germany) with an on column DNaseI treatment step per the manufacturer’s instructions (Table 1). Isolated RNAs were eluted in nuclease free water, validated for quality and quantity using UV spectrophotometry, and stored at −80 °C. RNAs with a OD260/280 ratio between 1.9 and 2.1 were deemed high quality and used for downstream analysis.
RNA preparation and sequencing
Total RNA samples were submitted to the Genewiz commercial sequencing facility (South Plainfield, NJ) for Bioanalyzer quality control analysis (Agilent, Santa Clara, CA) and Illumina Next Generation Sequencing. All submitted samples had an RNA integrity number (RIN) > 8. Stranded TruSeq cDNA libraries with poly dT enrichment were prepared from total RNA from each sample according to the manufacture’s protocol. Libraries for the 37 cDNA samples were sequenced using the Illumina HiSeq sequencing platform yielding 23–34.3 million 150 bp paired end (PE) sequence reads per sample (Table 1). 74 PE FASTQ files received back from Genewiz were analyzed using a customized bioinformatics workflow (Fig. 1d).
Quality validation, trimming, and read alignment
Between 23–34.3 million PE sequence reads per sample were delivered from Genewiz (Table 1). Trimmomatic software was used to filter and trim minority low quality sequencing reads from the data set21 (see Code availability 1). Figure 2a demonstrates that relatively few reads were filtered out of the data set. Quality of sequence reads in the 74 FASTQ files was evaluated using FastQC analysis22 (see Code availability 2), including per base (Fig. 2b) and per sequence (Fig. 2c) analysis which plots the Phred quality score distribution at each base and for each averaged full length read respectively for all reads in the data set. Collectively, Fig. 2 demonstrates that all 74 FASTQ sequencing files have an average per base Phred score >28, a conventional threshold denoting high quality NGS base calls. High quality sequence reads were aligned to the human hg38 reference transcriptome using the ultrafast Kallisto pseudoaligner23 (see Code availability 3). The percentage of aligned reads ranged from 73.9 to 91.1% (Table 1; Fig. 3a). Aggregate data visualizations for Trimmomatic, FastQC, and Kallisto were generated using MultiQC software24 (see Code availability 4).
Filtering and quality assessment of raw FASTQ sequencing data. RNA-seq FASTQ files representing the 37 PE samples used in this analysis were filtered and trimmed using Trimmomatic software (a). Following trimming and filtering, each of the 74 FASTQ files were assessed for average per base (b) and per sequence (c) quality as measured by Phred score.
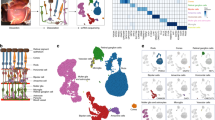
Quality assessment of read alignment and sample variance. The majority of high quality sequence reads successfully aligned to the human hg38 reference transcriptome using the Kallisto pseudoaligner (a). Kallisto transcript quantification of each sample was used for two dimensional principal component analysis (PCA) of sample variance with principle component 1 (PC1) and principle component 2 (PC2) accounting for the majority of each sample’s variance. PCA was used to visualize variance between the 4 distinct sample groups as well as similarity within sample replicates for all 37 samples (b). PCA was separately applied to 24 tightly clustered central retina and peripheral retina samples (c). Differentially expressed transcripts between central and peripheral retina samples plotted based on q-value [-log10(qval)] and fold change [b] demonstrates RHO, PDE6B, RCVRN, GUCY2F, and GNAT1 are preferentially expressed in the peripheral retina whereas CIB2, MAP1B, CRY2, NAPEPLD, DSCAM, CCDC66, and RPGR are representative transcripts preferentially expressed in the central retina (d).
Data transformation and downstream analysis
Transcript quantification of each sample was achieved using Kallisto pseudoalignments. Kallisto outputs were fed into the Sleuth statistical model25 for determination of differential transcript expression between samples (see Code availability 5). Sleuth is available as an R package and was used to generate a principal component analysis (PCA) plot demonstrating the variance between distinct sample groups as well as similarity within sample replicates for all 37 samples (Fig. 3b). To specifically highlight the utility of this data set for studying PR-specific cell type-restricted transcriptional networks, Sleuth was separately used for PCA of the tightly clustered 24 central and peripheral retina samples (Fig. 3c). This PCA plot demonstrates distinct clustering amongst 3 mm central retina and 6 mm peripheral retina samples respectively. Kallisto transcript quantification combined with Sleuth statistical analysis was further used to plot all differentially expressed transcripts between central and peripheral retina samples. This analysis demonstrates differential expression of rod-specific transcripts including RHO, PDE6B, RCVRN, GUCY2F, and GNAT1. These findings are similar to those observed in transcript analysis of the primate Macaca fascicularis central and peripheral retina supporting the practicality of our data set for studying PR cell type-restricted transcription26 (Fig. 3d). Curiously, cone-specific transcripts upregulated the central retina of M. fascicularis such as OPN1SW, OPN1MW, GNAT2, ARR3, and PDE6H were not differentially expressed in our study. Transcripts more abundantly expressed in human central retina involved in photoreceptor function include CIB2, MAP1B, CRY2, NAPEPLD, DSCAM, CCDC66, and RPGR (Fig. 3c). Discrepancies between human and non-human primate transcript expression patterns in the central retina are one of several areas of current investigation using this data set. Collectively, Fig. 3 demonstrates that the sampling strategy used in our study was effective for comparing differential transcript expression in clinically relevant ocular cell and tissue types.
Data Records
Raw FASTQ files for the RNA-seq libraries were deposited to the NCBI Sequence Read Archive (SRA), and have been assigned the SRA study accession SRP222833 (Table 1)27. Additionally, processed Kallisto output data files for trimmed and pseudoaligned sequence reads are accessible from the Figshare repository for each of the 37 samples analyzed in our study28. Kallisto produces three output files per sample:
-
1.
An abundances.h5 HDF5 binary file containing run info, abundance estimates, bootstrap estimates, and transcript length information length. This file can be read in by the sleuth statistical analysis program.
-
2.
An abundances.tsv plaintext file of the abundance estimates. Use the–plaintext mode to output plaintext abundance estimates. Alternatively, kallisto h5dump can be used to output an HDF5 file to plaintext. The first line contains a header for each column, including estimated counts, TPM, effective length.
-
3.
A run_info.json file containing information about the run.
Technical Validation
Quality control-RNA integrity
Quality of total RNA fractions was assessed using an Agilent Bioanalyzer to calculate a RNA Integrity Number (RIN). The RIN algorithm determines the RNA quality of the samples with the highest quality having a score of 10. Conventional to NGS analysis, only RNA samples with a RIN > 8 were used for sequencing analysis.
RNA-Seq raw data quality and filtering
Trimmomatic was used to filter and trim minority low quality sequencing reads and bases from downstream analysis (Fig. 2a). FastQC per base and per sequence quality analysis demonstrates mean Phred quality scores are well within the acceptable range for downstream analysis (Fig. 2b,c). Between 23 and 34.3 million reads were mapped to the human hg38 transcriptome assembly (Fig. 3a; Table 1). PCA biplot analysis confirmed the similarity between biological replicates and variability between tissue samples respectively (Fig. 3b,c).
Usage Notes
The bioinformatics pipeline applied to our data set outlined in Fig. 1d was achieved using a collection of freely available, open access tools. These analyses however, are interchangeable with many other currently available tools for achieving different experimental outcomes. Our raw FASTQ data can be aligned to any available human reference genome or transcriptome, including the most recent 2013 hg38 reference assemblies using a variety of freely available aligners. In this study we used Kallisto, an alignment-free transcriptome pseudoaligner23, with the specific interest in expression quantification of previously characterized mRNA isoforms. An alignment-free pipeline significantly reduces the time of analysis as well as required computing power and file storage, which may be beneficial for some users. Other very fast alignment-free programs such as Sailfish29 and Salmon30 can be used to achieve similar expression quantification analysis with these data. Alignment-free programs however, are not suitable for novel isoform analysis. More traditional alignment-based RNA-seq pipelines such as the ‘new tuxedo’ suite can be used to analyze these data for novel isoforms31. Here our differential gene expression analysis was carried out using the Sleuth software25, however other publicly available packages such as edgeR32 or StringTie31 may also be used for similar analysis. Importantly, QC data presented in Fig. 2 and Fig. 3 demonstrate the high quality of sequencing reads and precision of sampling respectively making this data set compatible with alignment tools currently available as well as new alignment tools that may become available in the future.
Our data set will be useful for a variety of studies investigating cell type and tissue-specific patterns of gene expression in the human retina, RPE/choroid, and cornea as well as diseases that affect these tissues. In particular, this work will build on existing genomic data sets investigating the human retina and AMD in particular. Fritsche and colleague’s landmark GWAS identified 52 SNPs at 32 loci constituting the majority of AMD heritability33. A recent transcriptome study integrating AMD GWAS data with RNA-seq data from 453 postmortem human retina samples identified three novel candidate AMD-associated genes34. Notably, Ratnapriya and colleagues collected their RNA-seq data from whole retina. In contrast, the study presented here focuses more specifically on primary sites of AMD pathology thereby providing a unique data set to the vision research community.
Several considerations must be taken into account when using these data for downstream analysis. First, RNAs were extracted from retinal tissue enriched in rod and cone photoreceptors, RPE/choroid, and whole cornea without any further enrichment for cell type. Therefore, resulting downstream analysis will be representative of heterogeneous mixtures of differing cell types within these tissues. In particular, 3 mm biopsies of the central retina sample a subsection of the 5.5 mm diameter macula encompassing the fovea, parafovea and portions of the perifovea. Though this cone-rich region contains some rod PRs, our data demonstrates that rod-specific transcripts are present at much lower counts compared to adjacent peripheral retina (Fig. 3d). These data suggest that cones are the predominant PR cell type in these samples, though notably, several canonical cone-specific genes (OPN1SW, OPN1MW, GNAT2, ARR3, PDE6H) were not differentially expressed in our study. Second, cDNA libraries were prepared using a poly dT primer, thus the data set is representative of only polyadenylated transcripts and does not represent many non-coding RNA or other non-polyadenylated cellular transcripts. Additionally, usage of poly dT priming introduces a bias towards overrepresentation of the 3′ end of transcripts, particularly in the case of relatively large transcripts. Finally, the quantity of sequenced and mapped reads per sample in this study (Table 1; Fig. 3a) is sufficient for robust differential transcript/gene expression analysis, however, is below the conventional threshold for thorough differential isoform analysis35. Taking these considerations into account, these data will be a useful resource for the vision research community for robust and accurate analysis of polyadenylated transcriptional networks in clinically relevant ocular cell and tissue types.
Code availability
The following open access software and versions were used for quality control and data analysis as described in the main text:
Trimmomatic, version 0.36 was used to filter and trim low quality reads and bases from FASTQ sequencing data files: http://www.usadellab.org/cms/?page=trimmomatic
FastQC, version 0.11.5 was used for quality analysis of raw FASTQ sequencing data: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Kallisto, version 0.42.3 was used to index and psedudoalign sequencing reads to the human hg38 transcriptome as well as to quantify transcripts in each samples: https://pachterlab.github.io/kallisto/
MultiQC, was used to aggregate and visualize FastQC, Trimmomatic, and Kallisto data outputs: https://multiqc.info/
Sleuth, was used to analyze transcript abundances quantified by Kallisto as well as to calculate and summarize differential transcript expression: https://pachterlab.github.io/sleuth/about
All code and walkthroughs used for quality assessment and data analysis in this study is available at: https://github.com/enkera/Schumacker2019_Sci_Data
References
Stephens, Z. D. et al. Big Data: Astronomical or Genomical? PLOS Biol. 13, e1002195 (2015).
Van den Berge, K. et al. RNA Sequencing Data: Hitchhiker’s Guide to Expression Analysis. Annu. Rev. Biomed. Data Sci. 2, 139–173 (2019).
Stark, R., Grzelak, M. & Hadfield, J. RNA sequencing: the teenage years. Nat. Rev. Genet. 1–26, https://doi.org/10.1038/s41576-019-0150-2 (2019).
Farkas, M. H., Au, E. D., Sousa, M. E. & Pierce, E. A. RNA-Seq: Improving Our Understanding of Retinal Biology and Disease. Cold Spring Harb. Perspect. Med. 5, a017152 (2015).
Yang, H.-J., Ratnapriya, R., Cogliati, T., Kim, J.-W. & Swaroop, A. Vision from next generation sequencing: Multi-dimensional genome-wide analysis for producing gene regulatory networks underlying retinal development, aging and disease. Prog. Retin. Eye Res. 46, 1–30 (2015).
Powell, C., Grant, A. R., Cornblath, E. & Goldman, D. Analysis of DNA methylation reveals a partial reprogramming of the Müller glia genome during retina regeneration. Proc. Natl. Acad. Sci. USA 110, 19814–9 (2013).
Lee, I. et al. Data Descriptor: Whole genome DNA methylation sequencing of the chicken retina, cornea and brain. Sci. Data 4 (2017).
Murphy, D. P., Hughes, A. E., Lawrence, K. A., Myers, C. A. & Corbo, J. C. Cis-regulatory basis of sister cell type divergence in the vertebrate retina. Elife 8 (2019).
Enright, J. M., Lawrence, K. A., Hadzic, T. & Corbo, J. C. Transcriptome profiling of developing photoreceptor subtypes reveals candidate genes involved in avian photoreceptor diversification. J. Comp. Neurol. 523, 649–668 (2015).
Langouet-Astrie, C. J., Meinsen, A. L., Grunwald, E. R., Turner, S. D. & Enke, R. A. RNA sequencing analysis of the developing chicken retina. Sci. Data 3 (2016).
Sun, C., Galicia, C. & Stenkamp, D. L. Transcripts within rod photoreceptors of the Zebrafish retina. BMC Genomics 19, 127 (2018).
Corbo, J. C. et al. CRX ChIP-seq reveals the cis-regulatory architecture of mouse photoreceptors. Genome Res. 20, 1512–1525 (2010).
Ruzycki, P. A., Zhang, X. & Chen, S. CRX directs photoreceptor differentiation by accelerating chromatin remodeling at specific target sites. Epigenetics Chromatin 11, 42 (2018).
Mo, A. et al. Epigenomic landscapes of retinal rods and cones. Elife 5 (2016).
Zelinger, L. & Swaroop, A. RNA Biology in Retinal Development and Disease. Trends Genet. 34, 341–351 (2018).
Xiao, M. & Hendrickson, A. Spatial and temporal expression of short, long/medium, or both opsins in human fetal cones. J. Comp. Neurol. 425, 545–559 (2000).
Swaroop, A., Kim, D. & Forrest, D. Transcriptional regulation of photoreceptor development and homeostasis in the mammalian retina. Nat. Rev. Neurosci. 11, 563–576 (2010).
Farkas, M. H. et al. Transcriptome analyses of the human retina identify unprecedented transcript diversity and 3.5 Mb of novel transcribed sequence via significant alternative splicing and novel genes. BMC Genomics 14, 486 (2013).
Li, M. et al. Comprehensive analysis of gene expression in human retina and supporting tissues. Hum. Mol. Genet. 23, 4001–4014 (2014).
Strauss, O. The Retinal Pigment Epithelium in Visual Function. Physiol. Rev. 85, 845–881 (2005).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Babraham Institute Bioinformatics Group. FastQC, version 0.11.5, http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Pimentel, H., Bray, N. L., Puente, S., Melsted, P. & Pachter, L. Differential analysis of RNA-seq incorporating quantification uncertainty. Nat. Methods 14, 687–690 (2017).
Mustafi, D. et al. Transcriptome analysis reveals rod/cone photoreceptor specific signatures across mammalian retinas. Hum. Mol. Genet. 25, ddw268 (2016).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP222833 (2019).
Schumacker, S. T. et al. RNA sequencing analysis of the human retina and associated ocular tissues. Figshare https://doi.org/10.6084/m9.figshare.c.4876785 (2020).
Patro, R., Mount, S. M. & Kingsford, C. Sailfish enables alignment-free isoform quantification from RNA-seq reads using lightweight algorithms. Nat. Biotechnol. 32, 462–464 (2014).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Fritsche, L. G. et al. A large genome-wide association study of age-related macular degeneration highlights contributions of rare and common variants. Nat. Genet. 48, 134–143 (2016).
Ratnapriya, R. et al. Retinal transcriptome and eQTL analyses identify genes associated with age-related macular degeneration. Nat. Genet. 51, 606–610 (2019).
Conesa, A. et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 17, 13 (2016).
Acknowledgements
The authors would like to thank David Micklos and Jason Williams at the Cold Spring Harbor Laboratory DNA Learning Center for providing bioinformatics consultation and support. This work was supported by NIH grant #1R15EY028725-01, Commonwealth Health Research Board grant #216-05-15a, JMU 4-VA Center for Genome & Metagenome Studies (CGEMS) Center Grant, and Burroughs Wellcome Fund Grant #1017506 awarded to R.A.E.
Author information
Authors and Affiliations
Contributions
S.T.S. conducted bioinformatics analysis and writing of the manuscript. K.R.C. assisted with bioinformatics analysis and writing of the manuscript. R.A.E. conceived and secured funding for the project as well as supervised all aspects of the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Online-only Table
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Schumacker, S.T., Coppage, K.R. & Enke, R.A. RNA sequencing analysis of the human retina and associated ocular tissues. Sci Data 7, 199 (2020). https://doi.org/10.1038/s41597-020-0541-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-020-0541-4
- Springer Nature Limited