Abstract
Purpose
Proglacial terrains provide an ideal environment to determine soil formation and microbial colonization, where autotrophic microbes could play a role as primary producers. We investigated the succession and drivers of autotrophic microorganisms harboring the cbbL gene in deglaciated soils across a 14-year deglaciation chronosequence on the Tibetan Plateau.
Materials and methods
Surface soils were collected in triplicate at 15 points beginning at the glacier terminus. The abundance and community composition of autotrophic microbes were determined using quantitative PCR and sequencing of clone libraries, respectively.
Results and discussion
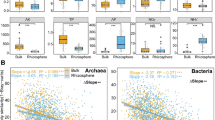
The abundance of form IC and ID cbbL increased gradually across deglaciation chronosequence, while form IA/B cbbL increased during the first 5 years since deglaciation and then remained stable. The succession of form IA/B and ID autotrophs occurred in a similar manner over three deglaciation stages, early (0-year-old), transitional (1–7-year-old), and late stage (8–14-year-old), whereas form IC autotrophs displayed distinct succession in the early (0–3-year-old) and transitional stage (4–7-year-old). Autotrophic microbial succession was prominently driven by soil moisture, pH, total carbon, and deglaciation age. Proteobacteria dominated young deglaciated soils (4–5-year-old), while Stramenopiles dominated old deglaciated soils (13-year-old). These findings revealed that autotrophic microbes follow clear successional patterns across recently deglaciated chronosequences and are primarily influenced by physicochemical parameters.
Conclusions
Autotrophic microbes quickly colonized deglaciated soils following glacier retreat, and their population increased with increasing deglaciation age. Physicochemical parameters played an important role in driving autotrophic microbes in the early, transitional, and late stages. Chemoautotrophic Proteobacterial-dominated young deglaciated soils to photoautotrophic algae-dominated old deglaciated soils.





Similar content being viewed by others
Data availability
Data are available upon request to the first author.
References
Alfreider A, Vogt C, Geiger-Kaiser M, Psenner R (2009) Distribution and diversity of autotrophic bacteria in groundwater systems based on the analysis of RubisCO genotypes. Sys Appl Microbiol 32:140–150. https://doi.org/10.1016/j.syapm.2008.11.005
Bowers RM, Lauber CL, Wiedinmyer C, Hamady M, Hallar AG, Fall R, Knight R, Fierer N (2009) Characterization of airborne microbial communities at a high-elevation site and their potential to act as atmospheric ice nuclei. Appl Environ Microbiol 75:5121–5130. https://doi.org/10.1128/AEM.00447-09
Bradley JA, Singarayer JS, Anesio AM (2014) Microbial community dynamics in the forefield of glaciers. Proc Royal Soc B 281:20140882. https://doi.org/10.1098/rspb.2014.0882
Cheng SM, Foght JM (2007) Cultivation-independent and -dependent characterization of Bacteria resident beneath John Evans Glacier. FEMS Microbiol Ecol 59:318–330. https://doi.org/10.1111/j.1574-6941.2006.00267.x
Corredor JE, Wawrik B, Paul JH et al (2004) Geochemical Rate-RNA integration study: ribulose-1,5-bisphosphate carboxylase/oxygenase gene transcription and photosynthetic capacity of planktonic photoautotrophs. Appl Environ Microbiol 70:5559–5568. https://doi.org/10.1128/AEM.70.9.5459-5468.2004
Del Moral Á, Garrido-Benavent I, Durán J, Lehmann JR, Rodríguez A, Heiðmarsson S, de Los RA (2021) Are recently deglaciated areas at both poles colonised by the same bacteria? FEMS Microbiol Lett 368:fnab011. https://doi.org/10.1093/femsle/fnab011
Elsaied HE, Hayashi T, Naganuma T (2004) Molecular analysis of deep-sea hydrothermal vent aerobic methanotrophs by targeting genes of 16S rRNA and particulate methane monooxygenase. Mar Biotechnol 6:503–509. https://doi.org/10.1007/Epub2004Aug24.s10126-004-3042-0
Frey B, Rieder SR, Brunner I, Plötze M, Koetzsch S, Lapanje A, Brandl H, Furrer G (2010) Weathering-associated bacteria from the Damma glacier forefield: physiological capabilities and impact on granite dissolution. Appl Environ Microbiol 76:4788–4796. https://doi.org/10.1128/AEM.00657-10
Gordon DA, Priscu J, Giovannoni S (2000) Origin and phylogeny of microbes living in permanent Antarctic lake ice. Microb Ecol 39:197–202. https://doi.org/10.1007/s002480000016
Guo G, Kong W, Liu J, Zhao J, Du H, Zhang X, Xia P (2015) Diversity and distribution of autotrophic microbial community along environmental gradients in grassland soils on the Tibetan Plateau. Appl Microbiol Biotechnol 99:8765–8776. https://doi.org/10.1007/s00253-015-6723-x
Guo G-X, Deng H, Qiao M, Yao H-Y, Zhu Y-G (2013) Effect of long-term wastewater irrigation on potential denitrification and denitrifying communities in soils at the watershed scale. Environ Sci Technol 47:3105–3113. https://doi.org/10.1021/es304714a
Gyeong H, Hyun CU, Kim SC, Tripathi BM, Yun J, Kim J, Kim M (2021) Contrasting early successional dynamics of bacterial and fungal communities in recently deglaciated soils of the maritime Antarctic. Mol Ecol 30:4231–4244. https://doi.org/10.1111/mec.16054
Hell K, Edwards A, Zarsky J, Podmirseg SM, Girdwood S, Pachebat JA, Insam H, Sattler B (2013) The dynamic bacterial communities of a melting high Arctic glacier snowpack. ISME J 7:1814–1826. https://doi.org/10.1038/ismej.2013.51
Heywood JL, Sieracki ME, Bellows W, Poulton NJ, Stepanauskas R (2011) Capturing diversity of marine heterotrophic protists: one cell at a time. ISME J 5:674–684. https://doi.org/10.1038/ismej.2010.155
Jangid K, Whitman WB, Condron LM, Turner BL, Williams MA (2013) Soil bacterial community succession during long-term ecosystem development. Mol Ecol 22:3415–3424. https://doi.org/10.1111/mec.12325
John DE, Wang ZA, Liu X, Byrne RH, Corredor JE, Lopez JM, Cabrera A, Bronk DA, Tabita FR, Paul JH (2007) Phytoplankton carbon fixation gene (RuBisCO) transcripts and air-sea CO2 flux in the Mississippi River plume. ISME J 1:517–531. https://doi.org/10.1038/ismej.2007.70
Kabala C, Zapart J (2012) Initial soil development and carbon accumulation on moraines of the rapidly retreating Werenskiold Glacier, SW Spitsbergen, Svalbard archipelago. Geoderma 175:9–20. https://doi.org/10.1016/j.geoderma.2012.01.025
Kastovska K, Elster J, Stibal M, Santruckova H (2005) Microbial assemblages in soil microbial succession after glacial retreat in Svalbard (high Arctic). Microb Ecol 50:396–407. https://doi.org/10.1007/s00248-005-0246-4
Kazemi S, Hatam I, Lanoil B (2016) Bacterial community succession in a high-altitude subarctic glacier foreland is a three-stage process. Mol Ecol 25:5557–5567. https://doi.org/10.1111/mec.13835
Khan A, Kong W, Ji M, Yue L, Xie Y, Liu J, Xu B (2020) Disparity in soil bacterial community succession along a short time-scale deglaciation chronosequence on the Tibetan Plateau. Soil Ecol Lett 2:83–92. https://doi.org/10.1007/s42832-020-0027-5
Khan A, Kong W, Muhammad S, Wang F, Zhang G, Kang S (2019) Contrasting environmental factors drive bacterial and eukaryotic community successions in freshly deglaciated soils. FEMS Microbiol Lett 366:fnz229. https://doi.org/10.1093/femsle/fnz229
Kim M, Jung JY, Laffly D, Kwon HY, Lee YK (2017) Shifts in bacterial community structure during succession in a glacier foreland of the High Arctic. FEMS Microbiol Ecol. https://doi.org/10.1093/femsec/fiw213
Kong W, Ream DC, Priscu JC, Morgan-Kiss RM (2012) Diversity and expression of RubisCO genes in a perennially ice-covered Antarctic lake during the polar night transition. Appl Environ Microbiol 78:4358–4366. https://doi.org/10.1128/AEM.00029-12
Kumar A, Mukhia S, Kumar R (2022) Microbial community dynamics from a fast-receding glacier of Western Himalayas highlight the importance of microbes in primary succession, nutrient recycling, and xenobiotics degradation. Ecol Indic 144:109565. https://doi.org/10.1016/j.ecolind.2022.109565
Liu J, Kong W, Zhang G, Khan A, Guo G, Zhu C, Wei X, Kang S, Morgan-Kiss RM (2016) Diversity and succession of autotrophic microbial community in high-elevation soils along deglaciation chronosequence. FEMS Microbiol Ecol 92:fiw160. https://doi.org/10.1093/femsec/fiw160
Montross S, Skidmore M, Christner B, Samyn D, Tison J-L, Lorrain R, Doyle S, Fitzsimons S (2014) Debris-rich basal ice as a microbial habitat, Taylor Glacier, Antarctica. Geomicrobiol 31:76–81. https://doi.org/10.1080/01490451.2013.811316
Nemergut DR, Anderson SP, Cleveland CC, Martin AP, Miller AE, Seimon A, Schmidt SK (2007) Microbial community succession in an unvegetated, recently deglaciated soil. Microb Ecol 53:110–122. https://doi.org/10.1007/s00248-006-9144-7
Norton JM, Klotz MG, Stein LY, Arp DJ, Bottomley PJ, Chain PS, Hauser LJ, Land ML, Larimer FW, Shin MW (2008) Complete genome sequence of Nitrosospira multiformis, an ammonia-oxidizing bacterium from the soil environment. Appl Environ Microbiol 74:3559–3572. https://doi.org/10.1128/AEM.02722-07
Nowak A, Hodson A (2014) Changes in meltwater chemistry over a 20-year period following a termal regime switch from polythermal to cold-based glaciation at Austre Broggerbreen. Svalbard Polar Res 33:22779. https://doi.org/10.3402/polar.v33.22779
Paul JH, Alfreider A, Wawrik B (2000) Micro- and macrodiversity in rbcL sequences in ambient phytoplankton populations from the southeastern Gulf of Mexico. Mar Ecol Prog Ser 198:9–18. https://doi.org/10.3354/meps198009
Pessi IS, Osorio-Forero C, Gálvez EJ, Simões FL, Simões JC, Junca H, Macedo AJ (2015) Distinct composition signatures of archaeal and bacterial phylotypes in the Wanda Glacier forefield, Antarctic Peninsula. FEMS Microbiol Ecol 91:1–10. https://doi.org/10.1093/femsec/fiu005
Philippot L, Tscherko D, Bru D, Kandeler E (2011) Distribution of high bacterial taxa across the chronosequence of two alpine glacier forelands. Microb Ecol 61:303–312. https://doi.org/10.1007/s00248-010-9754-y
Pothula SK, Adams BJ (2022) Community assembly in the wake of glacial retreat: a meta-analysis. Glob Change Biol 28:6973–6991. https://doi.org/10.1111/gcb.16427
Rola K, Rożek K, Chowaniec K, Błaszkowski J, Gielas I, Stanek M, Zubek S (2022) Vascular plant and cryptogam abundance as well as soil chemical properties shape microbial communities in the successional gradient of glacier foreland soils. Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2022.160550
Rull V (2020) Quaternary ecology, evolution, and biogeography. Academic Press
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Schmidt S, Nemergut D, Miller A, Freeman K, King A, Seimon A (2009) Microbial activity and diversity during extreme freeze–thaw cycles in periglacial soils, 5400 m elevation, Cordillera Vilcanota, Perú. Extremophiles 13:807–816. https://doi.org/10.1007/s00792-009-0268-9
Segawa T, Miyamoto K, Ushida K, Agata K, Okada N, Kohshima S (2005) Seasonal change in bacterial flora and biomass in mountain snow from the Tateyama Mountains, Japan, analyzed by 16S rRNA gene sequencing and real-time PCR. Appl Environ Microbiol 71:123–130. https://doi.org/10.1128/AEM.71.1.123-130.2005
Sheridan PP, Miteva VI, Brenchley JE (2003) Phylogenetic analysis of anaerobic psychrophilic enrichment cultures obtained from a Greenland glacier ice core. Appl Environ Microbiol 69:2153–2160. https://doi.org/10.1128/AEM.69.4.2153-2160.2003
Stibal M, Gözdereliler E, Cameron KA, Box JE, Stevens IT, Gokul JK, Schostag M, Zarsky JD, Edwards A, Irvine-Fynn TD, Jacobsen CS (2015) Microbial abundance in surface ice on the Greenland Ice Sheet. Front Microbiol 6:225. https://doi.org/10.3389/fmicb.2015.00225
Stopnisek N, Bodenhausen N, Frey B, Fierer N, Eberl L, Weisskopf L (2014) Genus-wide acid tolerance accounts for the biogeographical distribution of soil Burkholderia populations. Environ Microbiol 16:1503–1512. https://doi.org/10.1111/1462-2920.12211
Strauss SL, Ruhland CT, Day TA (2009) Trends in soil characteristics along a recently deglaciated foreland on Anvers Island, Antarctic Peninsula. Polar Biol 32:1779–1788. https://doi.org/10.1007/s00300-009-0677-3
Tabita FR, Hanson TE, Satagopan S, Witte BH, Kreel NE (2008a) Phylogenetic and evolutionary relationships of RubisCO and the RubisCO-like proteins and the functional lessons provided by diverse molecular forms. Philos Trans R Soc Lond B Bio Sci 363:2629–2640. https://doi.org/10.1098/rstb.2008.0023
Tabita FR, Satagopan S, Hanson TE, Kreel NE, Scott SS (2008b) Distinct form I, II, III, and IV Rubisco proteins from the three kingdoms of life provide clues about Rubisco evolution and structure/function relationships. J Exp Bot 59:1515–1524. https://doi.org/10.1093/jxb/erm361
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tan KH (2005) Soil sampling, preparation, and analysis. CRC Press
Venkatachalam S, Kannan VM, Saritha VN, Loganathachetti DS, Mohan M, Krishnan KP (2021) Bacterial diversity and community structure along the glacier foreland of Midtre Lovénbreen, Svalbard, Arctic. Ecol Indic 126:107704. https://doi.org/10.1016/j.ecolind.2021.107704
Walker LR, Wardle DA, Bardgett RD, Clarkson BD (2010) The use of chronosequences in studies of ecological succession and soil development. J Ecol 98:725–736. https://doi.org/10.1111/j.1365-2745.2010.01664.x
Wu X, Zhang W, Liu G, Yang X, Hu P, Chen T, Zhang G, Li Z (2012) Bacterial diversity in the foreland of the Tianshan No. 1 glacier, China. Environ Res Lett 7:014038. https://doi.org/10.3389/fmicb.2016.01353
Xiao KQ, Bao P, Bao QL, Jia Y, Huang FY, Su JQ, Zhu YG (2013) Quantitative analyses of ribulose-1, 5-bisphosphate carboxylase/oxygenase (RubisCO) large-subunit genes (cbbL) in typical paddy soils. FEMS Microbiol Ecol 87:89–101. https://doi.org/10.1111/1574-6941.12193
Yao T, Thompson L, Yang W, Yu W, Gao Y, Guo X, Yang X, Duan K, Zhao H, Xu B (2012) Different glacier status with atmospheric circulations in Tibetan Plateau and surroundings. Nat Clim Chang 2:663–667. https://doi.org/10.1038/nclimate1580
Yuan H, Ge T, Chen C, O’Donnell AG, Wu J (2012) Significant role for microbial autotrophy in the sequestration of soil carbon. Appl Environ Microbiol 78:2328–2336. https://doi.org/10.1128/AEM.06881-11
Zumsteg A, Luster J, Goransson H, Smittenberg RH, Brunner I, Bernasconi SM, Zeyer J, Frey B (2012) Bacterial, Archaeal and Fungal Succession in the Forefield of a Receding Glacier. Microbiol Ecol 63:552–564. https://doi.org/10.1007/s00248-011-9991-8
Acknowledgements
The authors greatly thank the Muztagh Ata Station for Westerlies Environment Observation and Research, Chinese Academy of Sciences, for logistic support and sampling assistance.
Funding
This work was supported by the National Natural Science Foundation of China (32161123004 and 42177101) and the Chinese Academy of Sciences (XDA19070304).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Competing interests
The authors declare no competing interests.
Additional information
Responsible editor: Yuan Ge.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khan, A., Kong, W., Khan, S. et al. Community succession and drivers of CO2-fixing microbes in recently deglaciated soils on the Tibetan Plateau. J Soils Sediments 23, 1901–1912 (2023). https://doi.org/10.1007/s11368-023-03446-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-023-03446-6