Abstract
An increasing burden of neurological diseases (NDs) has been a public health challenge in an aging society. Age, especially biological age, is the most important risk factor for NDs. Identification of biomarkers of aging to capture NDs might lead to a better understanding of the underlying mechanisms of pathological brain aging and the implementation of effective intervention. We conducted a comprehensive two-sample Mendelian Randomization (MR) study to investigate the association between various biomarkers of aging and three leading causes of NDs: Alzheimer’s disease (AD), vascular dementia (VaD), and ischemic stroke. Publicly available GWAS summary statistics on people from European ancestry were obtained for six molecular biomarkers, two physiological biomarkers, and eight functional biomarkers, and three NDs. Genetic variants serving as instrumental variables (IVs) were identified for each biomarker. The MR analysis included inverse variance weighted (IVW), weighted median, MR-Egger, and MR-PRESSO. We found that short telomere length and decrease in appendicular lean mass were associated with an increased risk for AD (OR IVW = 1.12 per 1SD decrease, 95% confidence interval 1.02–1.22, and OR IVW = 1.11, 1.06–1.16, respectively), whereas high frailty index showed a protective effect for AD. Accelerated BioAge appeared to be associated with increased risk for ischemic stroke (OR IVW = 1.3 per year in BioAge acceleration, 95% CI 1.19–1.41). Our findings implied a causal association of short telomere length and a decrease in appendicular lean mass with an increased risk for AD, while BioAge appeared to be a good biomarker for ischemic stroke. Further studies are needed to validate these associations and explore underlying mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The world has experienced an unprecedented demographic shift toward an aging society with a dramatic increase in life expectancy. Despite this increase in life expectancy, the improvements in health span, defined as lifespan without chronic diseases and disability, have lagged behind. This hinders us from fully enjoying extended lifespans [1, 2]. Therefore, expanding the health span is a public health priority.
Age is the most important risk factor for various age-related chronic diseases and disability [3, 4]. However, the aging process varies widely in individuals, and chronological age often fails to capture underlying molecular damage and functional decline accurately. Thus, the evaluation of biological age, which reflects age-related biological changes, is becoming more important [5]. Unlike chronological age, biological age could be modifiable, providing tools to identify possible anti-aging interventions and treatments. Therefore, biomarkers to assess biological age have been intensely sought out in the past decades [6].
Numerous biomarkers of aging have been proposed. According to the Biomarkers of Aging Consortium’s proposal, these biomarkers can be broadly categorized into three groups based on what they measure: molecular biomarkers based on omics data, physiological biomarkers to measure functional performance, and digital biomarkers [5]. Particularly, the molecular-based group constitutes the largest group due to the recent advancement of molecular biology. At the molecular level, 12 hallmarks of aging have been identified as underlying mechanisms of the aging process [7]. Despite the increasing number of biomarkers, there is no gold standard biomarker of aging. Many aging researchers think that no single biomarker can capture all aspects of aging due to the multi-dimensional and complicated nature of the aging process. It is more likely that different biomarkers reflect distinct aging aspects across various tissues and organs.
Among age-related chronic diseases, dementia and other neurological diseases (NDs) are some of the most common and feared ones, consisting of 19.5% of total deaths in the EU [8]. Eight hallmarks of ND have been proposed as drivers of pathological neurological aging [9]. These hallmarks largely overlap with those of aging but have some distinct characteristics [7]. For example, genetic instability and DNA/RNA damage are considered primary triggers for all other hallmarks, driving both aging and neurodegeneration. Furthermore, the impairment of mitochondria, crucial for cellular energy production, appears to accelerate both aging and neurogenerative aging [7, 10, 11]. Among ND hallmarks, pathological protein accumulation and synaptic function are crucial, characterizing various NDs. This implies that ND has its own biomarkers that most accurately capture pathological neurological aging. Recently, some observational studies showed biomarkers of aging, such as the epigenetic clock based on DNA methylation data and biological age derived from routinely measured clinical biomarkers, predict NDs, including dementia [12,13,14]. However, as a recent systematic review shows, observational studies, especially large-scale prospective studies, are limited and have inconsistent results [15]. Besides, observational studies are susceptible to confounding and bias. Moreover, most studies use one type of biomarker at a time, which makes the comparison of various biomarkers difficult in terms of their ability to predict NDs. Mendelian randomization (MR) is an analytical approach to investigate the association using genetic variants as instrumental variables, mitigating concerns over confounding and reverse causation [16]. The use of MR may strengthen existing evidence through triangulation of evidence.
The aim of our study is to investigate the associations of various biomarkers of aging, including molecular, physiological, and functional biomarkers, with three common NDs: Alzheimer’s disease (AD), vascular dementia (VaD), and ischemic stroke, using the MR approach to provide evidence for putative causal mechanisms of biological aging effects on NDs. Our study seeks to strengthen existing evidence and shed some light on the underlying mechanism.
Methods
Study design
We used publicly available GWAS summary statistics for biomarkers of aging and NDs. Ethical approval was obtained in all original studies. A summary of study data sources is provided in Table 1. The paper follows reporting according to the STROBE-MR guidelines [17].
We conducted a two-sample MR to investigate the association between multiple biomarkers of aging and three NDs. As exposures, we used six molecular biomarkers, two physiological biomarkers, and eight functional biomarkers. Genetic variants that are used as instrumental variables (IVs) were identified in the GWAS dataset for each biomarker. To obtain unbiased causal estimates in MR, IVs need to satisfy three key assumptions. First, IVs need to be associated with the exposure of interest (the relevance assumption). Secondly, IVs should not share a common cause with the outcome (the independent assumption). Finally, IVs should not affect the outcome except through the exposure (the exclusion restriction assumption) [18]. A framework of our study design is presented in Fig. 1.
Genetic instruments for biomarkers of aging
We obtained publicly available GWAS summary statistics for various biomarkers of aging. We categorized biomarkers into three groups: molecular biomarkers based on omics, physiological biomarkers based on clinical data, and functional biomarkers based on functional performance. Molecular biomarkers include telomere length [19], four types of epigenetic clocks derived from DNA methylation (HannumAge,
IEAA, PhenoAge, GrimAge) [20], and mitochondria DNA copy number [21]. As physiological biomarkers, we obtained BioAge acceleration and PhenoAge acceleration [22]. Functional biomarkers included handgrip, appendicular lean mass, two lung function measures (FVC, forced vital capacity; FEV1, forced expiratory volume in 1 s), cognitive performance [26], retinal eye clock [25], brain age gap [27], and frailty index [28]. The short explanation of each biomarker is presented in Table 2.
For each biomarker, we identified variants with genome-wide significance (p < 5 × 10−8), and retained those independent variants as IVs (linkage disequilibrium r2 < 0.001) based on the European reference panel from the 1000 Genomes Project Phase 3 (Supplementary Table 1). The plausibleness of the relevant assumption was assessed using F statistics [18]. The proportion of trait variance explained by genetic instruments R2 was estimated using the formulae:
\({R}^{2}=(2{\beta }^{2}\text{ MAF (1-MAF))}/(2{\beta }^{2} MAF \left(1-MAF\right)+2N MAF \left(1-MAF\right) S{E}^{2})\) and the F statistic was calculated using the following formulae: \(F=({R}^{2} (N-2)/(1-{R}^{2})\) (where MAF = effect allele frequency, N = sample size, β = beta coefficient of the SNP, SE = standard error).
Study outcomes
We obtained publicly available GWAS summary statistics for each ND outcome. Only genetic data on people with European ancestry was included. We obtained the summary statistics for AD from the largest AD GWAS data, with 39,106 clinical cases, 46,828 proxy cases, and 401,577 controls [30]. In this GWAS data, the results from the European Alzheimer & Dementia Biobank consortium with clinical AD cases were meta-analyzed with the UK Biobank results with proxy AD cases. Clinical AD cases were mainly ascertained through clinical criteria and/or diagnosis from memory clinics, in spite of some variations among studies in the consortium [30]. Proxy AD cases in the UK Biobank were identified when participants had at least one parent or sibling affected by dementia. For VaD, summary statistics were taken from FinnGen, comprising 881 cases and 221,508 controls. Cases were identified from nationwide health registries using ICD-10 codes [32]. The summary statistics of all ischemic strokes were obtained from the MEGASTROKE consortium, including 40,585 cases and 406,111 controls in the European population (Table 1) [33]. The identification of ischemic stroke cases was mainly based on clinical symptoms and CT/MRI imaging in spite of variations among the included studies. Details of AD and stroke case identification can be found in the original articles [30, 33].
Positive and negative control analysis
To assess the validity of selected SNPs, we conducted an MR analysis using parental lifespan as a positive control outcome, where statistically significant associations with biomarkers of aging are expected. The absence of significant association may indicate invalid IVs or low power. Furthermore, to detect potential population stratification and invalidity of IVs, we repeated the analysis using skin color as a negative control outcome, where no associations with biomarkers of aging are expected. Any statistically significant associations with this negative control would suggest potential population stratification or invalid IVs [41]. GWAS data for parental lifespan were sourced from the University of Edinburgh data share site (http://dx.doi.org/https://doi.org/10.7488/ds/2463), and that for skin color was obtained using the University of Bristol’s IEU OpenGWAS API (ukb-b-19560).
Statistical analysis
Then, we conducted a two-sample MR analysis to investigate the associations between biomarkers of aging and three NDs. The IV dataset was merged with the outcome GWAS summary. For an absent SNP in ND GWAS, a proxy SNP was sought using the European panel from the 1000 Genome Project as the reference panel (R2 > 0.8). Palindromic SNPs with minor allele frequency > 0.42 were excluded. As the main analysis, we used the inverse variance-weighted (IVW) method to estimate the association of biomarkers of aging with each outcome. The Cochran’s Q test was used to assess heterogeneity across the IVs.
Sensitivity analysis
To obtain more robust MR estimates and explore violations of the exclusion restriction assumption, we conducted several sensitivity analyses, including the weighted median regression which allows up to 50% of invalid IVs [42], MR-Egger regression which allows the presence of pleiotropy [43], and MR-PRESSO which can detect and correct horizontal pleiotropy [44]. Similar effect estimates across various methods make the observed association more likely [45]. The funnel plots were visually inspected where asymmetry suggesting the presence of pleiotropy. Furthermore, due to sample overlap between the AD GWAS and GWAS for biomarkers of aging, we repeated the AD analysis by excluding the UK Biobank population from the AD GWAS.
All statistical analyses were performed in R version 4.2.3 using the TwoSampleMR package. For each MR analysis, post hoc power calculations were conducted using an online calculator tool (https://sb452.shinyapps.io/power/).
Results
F-statistics of genetic instruments ranged from 28.0 to 1105.8 (Supplementary Table 2). The MR analysis using the negative control outcome skin color did not show evidence of population stratification for most of the biomarkers, except for cognitive performance and frailty index (Supplementary Fig. 1). Regarding the analysis using the positive control outcome (parental lifespan), all molecular biomarkers showed statistically insignificant associations although the direction of association indicated that advanced biological age was associated with shorter parental lifespan, as expected (Supplementary Fig. 2). Most functional biomarkers showed statistically significant associations in the expected direction. Regarding appendicular lean mass, FEV1, and retinal eye clock, the associations were insignificant in spite of expected directions.
Biomarkers of aging and AD
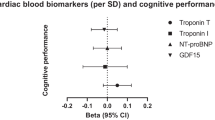
Overall, we observed a positive association between advanced biological age and increased risk of AD (Fig. 2A). This association was clear in functional biomarkers but not in molecular biomarkers. Among molecular biomarkers, only short TL was associated with an increased risk of AD (OR IVW = 1.12 per 1SD increase in TL, 95% CI 1.02–1.22). Physiological biomarkers showed rather a protective effect. BioAge acceleration was associated with decreased risk of AD (OR IVW = 0.87 per year in BioAge acceleration, 95% CI 0.82–0.93). Functional biomarkers appeared to increase the risk of AD, with some evidence for appendicular lean mass (OR IVW = 1.11 per 1SD decrease in lean mass, 95% CI 1.06–1.16) and FVC (OR IVW = 1.18 per 1SD decrease in FVC, 95% CI 1.07–1.29). On the contrary, the genetic liability for a higher frailty index showed a protective effect of AD (OR IVW = 0.67, 95% CI 0.49–0.87). Post hoc power calculation suggested AD GWAS sample size is enough for most biomarkers except for some biomarkers such as GrimAge, BioAge, retinal eye clock, brain age gap, and frailty index (Supplementary Table 3).
Inverse-variance weighted Mendelian randomization estimates for genetically predicted biomarkers of aging on A Alzheimer’s disease, B vascular dementia, and C ischemic stroke. AD, Alzheimer’s disease; CI, confidence interval; FEV1, forced expiratory volume; FVC, forced vital capacity; N_SNV, number of single-nucleotide variations; OR IVW, odds ratio inverse variance weighted; VaD, vascular dementia
Biomarkers of aging and VaD
We did not find a clear trend in the association between advanced biological age and VaD using the IVW (Fig. 2B). Biomarkers showed no evidence of association with the risk of VaD, and the confidence intervals were considerably wide. Post hoc power calculation suggests that the sample size is not enough to detect the association between biomarkers of aging and VaD with 80% power (Supplementary Table 3).
Biomarkers of aging and ischemic stroke
The results from IVW MR suggested a positive association of physiological and functional biomarkers with risk of ischemic stroke (Fig. 2C). However, no single molecular biomarkers were associated with the risk of ischemic stroke. Regarding physiological biomarkers, both BioAge and PhenoAge acceleration were associated with an increased risk of ischemic stroke (OR IVW = 1.30 per year in BioAge acceleration, 95% CI 1.19–1.41; OR IVW = 1.02 per year in PhenoAge acceleration, 95% CI 1.00–1.05). Among functional biomarkers, a decrease in lung function, both FVC and FEV1, and lower appendicular lean mass increased the risk of ischemic stroke (OR IVW = 1.08, 1.12, and 1.16 per 1 SE decrease, respectively).
Sensitivity analysis
Cochran’s Q statistics suggested heterogeneity of effect estimates among SNPs for many biomarkers (Supplementary Table 2). However, the intercept of MR Egger did not indicate the presence of pleiotropy for most of the biomarkers (Supplementary Table 2). MR estimates obtained from more robust methods are presented in Supplementary Table 2. Although most of the associations became insignificant, TL, appendicular lean mass, and frailty remained significantly associated with AD in most sensitivity analyses. Regarding ischemic stroke, only the association with BioAge acceleration remained robust in sensitivity analyses.
The GWAS for biomarkers of aging we used were mostly derived from the UK Biobank participants. Because our AD GWAS includes the UKB participants, to address population overlap, we repeated the MR analysis by excluding the UKB participants from the AD GWAS (Supplementary Fig. 3). As for our main findings, appendicular lean mass stayed significantly associated with AD. Although the association of TL and frailty index became insignificant, the direction of effect remained unchanged.
Discussion
We investigated the association of multiple biomarkers of aging with three NDs using the two-sample MR framework, and had five main findings. Firstly, a positive association of advanced biological age with increased risk of AD and ischemic stroke was observed, particularly for functional biomarkers. Notably, no molecular marker was associated with ischemic stroke. Secondly, among molecular biomarkers, short TL appeared to be associated with an increased risk of AD but not ischemic stroke. Thirdly, physiological biomarker BioAge acceleration was associated with an increased risk of ischemic stroke. Fourthly, among functional biomarkers, reduced appendicular lean mass may increase the risk of AD. Finally, an increased frailty index showed a protective effect for AD.
Our MR findings suggested that short TL may be causally related to AD. This result aligns with previous observational studies and an MR study based on 16 SNPs from a small AD GWAS [46, 47]. Regarding other molecular biomarkers, no significant associations with NDs were observed. This was unexpected, given the accumulating evidence from observational studies suggesting the association between molecular biomarkers and NDs. The discrepancy could be partially attributed to insufficient statistical power. The number of instrumental variables tended to be lower for molecular biomarkers than for functional ones, except for TL. Moreover, the sample size for GWAS data on four epigenetic clocks was smaller in comparison to GWAS data for functional biomarkers. It is possible that we could not capture the true association between molecular biomarkers and neurological diseases due to the quality of instrumental variables. For an unbiased and precise MR estimate, further studies are necessary, utilizing more valid SNPs and enhanced GWAS data.
Regarding our MR finding of the association between appendicular lean mass and AD, prospective observational studies are limited and have mixed results. In a US study with 1175 community-dwelling older people, a decrease in skeletal muscle mass estimated using bioimpedance analysis was not associated with AD/mild cognitive impairment, while grip strength was associated with AD after a 5.6-year follow-up [48]. Conversely, a small US study with 344 old people showed that total lean mass, especially appendicular lean mass, was associated with a low risk for dementia, while handgrip was not after 15 years of follow-up [49]. Taken together, skeletal muscle appears to play an important role in the development of dementia. Further prospective or MR studies of high quality are needed to confirm this finding. A high frailty index, another functional biomarker, showed a protective effect for AD, which is counter intuitive. Previous observational studies in this area are relatively limited with mixed results. Some meta-analyses of cohort studies suggest that frailty is associated with dementia and cognitive decline, which contrasts with our findings [50,51,52]. However, most cohort studies measure frailty using the frailty phenotype, and the frailty index is dichotomized. Additionally, studies using a continuous frailty index are not included in these meta-analyses, which might explain differences from our findings. Regarding a protective effect for AD observed in our study, selective survival might to some extent explain this. Unlike dementia, the association between a high frailty index and increased mortality has been consistently shown in previous cohort studies [53]. High mortality rates among people with a high frailty index might preclude the occurrence of dementia, leading to an apparent protective effect. However, the frailty index showed a significant association with negative control outcome, raising some concerns over the presence of population stratification and the validity of the IVs. An MR study using more valid IVs, along with a prospective cohort study with a strong design, is needed to confirm our findings.
Regarding ischemic stroke, BioAge showed a robust association with an increased risk. This finding is consistent with two recent cohort studies. In the SATSA study conducted in Sweden, participants with high BioAge acceleration at baseline were associated with a higher risk of ischemic stroke over a 20-year follow-up [12]. Similarly, the analysis of 325,870 UK Biobank participants showed that BioAge acceleration at baseline was associated with ischemic stroke, along with all-cause dementia, and vascular dementia [13]. Taken together, these findings suggest that BioAge could serve as a valuable biomarker for predicting vascular brain aging. Conversely, BioAge appeared to have a protective effect for AD in our analysis. Previous studies show that high BioAge derived using Klemera and Doubal method was associated with high mortality [38, 54, 55]. Furthermore, BioAge is derived from several cardiovascular risk factors, such as systolic blood pressure and total cholesterol, and it reflects cardiovascular health. Similar to the discussion about frailty above, this apparent protective effect might be attributed to high mortality and cardiovascular risk in people with high BioAge.
The exact biological mechanism linking telomere length with AD remains unclear. Human telomeres, located at the end of chromosomes, protect them from DNA damage and regulate gene expression changes. Progressive telomere shortening with age leads not only to DNA damage and cell senescence but also to changes in gene expression and epigenetic alterations [56]. Because DNA instability induced by telomere attrition is upstream of all aging hallmarks, it can drive all remaining hallmarks [57]. This might explain the potential association between telomere shortening and brain aging. Regarding the link between appendicular lean mass and AD, a lot of attention has been paid to the role of myokines recently. Evidence suggests that loss of proteostasis, deregulated nutrient sensing, and mitochondria dysfunction are important drivers of skeletal muscle aging and serve as aging hallmarks [58]. Skeletal muscle, which comprises 40% of the human body, is the main source of myokines, including inflammatory cytokines, brain-derived neurotrophic factor, and fibroblast growth factor 21. Myokines play various roles in maintaining bodily homeostasis, including the control of inflammation and insulin resistance. Recent evidence suggests myokines might play a vital role in the cross-talk between skeletal muscle and the brain, helping to maintain cognitive function [59]. Skeletal muscle aging leads to decreased muscle mass and strength, resulting in a lower level of myokines. This reduction in myokines may contribute to cognitive dysfunction through changes in autocrine, paracrine, and endocrine function [60]. The link between BioAge acceleration and ischemic stroke is understandable because BioAge includes some cardiovascular risk factors such as systolic blood pressure and total cholesterol [22].
We conducted a comprehensive MR analysis to investigate the association of various biomarkers of aging with three neurological diseases, identifying several potential biomarkers. Some of our findings lend support to previous observational studies through triangulation of evidence. Nevertheless, our study has several limitations. First, there was a large population overlap between our exposure GWAS and the AD GWAS, attributed to both studies including participants from the UK Biobank participants. Overlapping samples can generate biased effect estimates. However, a recent simulation study suggests that bias introduced by population overlap might not be as significant as previously anticipated [61]. Besides, eliminating overlapping samples could lead to weak statistical power [62]. Consequently, to preserve statistical power, we investigated the association between biological age and AD using overlapping samples in the main analysis and conducted a sensitivity analysis excluding overlapping samples, which did not materially alter our main findings. Second, we cannot rule out the potential presence of population stratification, which introduces confounding. Indeed, a recent research study showed geographical clustering of genetic traits in the UK, most notably educational attainment [63]. Nevertheless, our analyses using negative control outcomes did not show a strong suggestion of the presence of population stratification for the majority of the biomarkers, except for cognitive performance and frailty index. Third, some biomarkers of aging had a small number of IVs, and post hoc power calculation showed a lack of outcome GWAS sample size, especially for VaD GWAS. Due to inadequate statistical power, our findings might not accurately reflect a true association. Further MR studies are required to confirm our findings. Finally, our analysis included only participants of European ancestry due to sample size availability. GWAS studies with more diverse populations are needed to enhance our understanding of the risk mechanism of NDs.
From a clinical perspective, our findings suggest that people with short TL and decreased appendicular lean mass are at an increased risk of AD and that BioAge acceleration increases the risk of ischemic stroke. If further studies confirm causal relationships, they might help identify high-risk populations and lead to potential intervention targets. Particularly, the appendicular lean mass would be an attractive target because bioimpedance analysis is noninvasive, widely available, and inexpensive, and an intervention to prevent lean mass loss is possible.
In conclusion, our findings suggested underlying causal associations of short telomere length and decreased appendicular lean mass with an increased risk for AD, while BioAge appeared to be a good biomarker for ischemic stroke. Further studies are needed to validate these associations and explore underlying mechanisms.
References
Garmany A, Yamada S, Terzic A. Longevity leap: mind the healthspan gap. NPJ Regen Med. 2021;6(1):57.
Lee J, Lau S, Meijer E, Hu P. Living longer, with or without disability? A global and longitudinal perspective. J Gerontol Ser A. 2020;75(1):162–7.
Hou Y, Dan X, Babbar M, Wei Y, Hasselbalch SG, Croteau DL, et al. Ageing as a risk factor for neurodegenerative disease. Nat Rev Neurol. 2019;15(10):565–81.
Niccoli T, Partridge L. Ageing as a risk factor for disease. Curr Biol. 2012;22(17):R741–52.
Moqri M, Herzog C, Poganik JR, Justice J, Belsky DW, Higgins-Chen A, et al. Biomarkers of aging for the identification and evaluation of longevity interventions. Cell. 2023;186(18):3758–75.
Jylhävä J, Pedersen NL, Hägg S. Biological age predictors. EBioMedicine. 2017;21:29–36.
López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. Hallmarks of aging: an expanding universe. Cell. 2023;186(2):243–78.
Deuschl G, Beghi E, Fazekas F, Varga T, Christoforidi KA, Sipido E, et al. The burden of neurological diseases in Europe: an analysis for the Global Burden of Disease Study 2017. Lancet Public Health. 2020;5(10):e551–67.
Wilson DM, Cookson MR, Van Den Bosch L, Zetterberg H, Holtzman DM, Dewachter I. Hallmarks of neurodegenerative diseases. Cell. 2023;186(4):693–714.
Fang EF, Hou Y, Palikaras K, Adriaanse BA, Kerr JS, Yang B, et al. Mitophagy inhibits amyloid-β and tau pathology and reverses cognitive deficits in models of Alzheimer’s disease. Nat Neurosci. 2019;22(3):401–12.
Norat P, Soldozy S, Sokolowski JD, Gorick CM, Kumar JS, Chae Y, et al. Mitochondrial dysfunction in neurological disorders: exploring mitochondrial transplantation. NPJ Regen Med. 2020;5(1):22.
McMurran CE, Wang Y, Mak JKL, Karlsson IK, Tang B, Ploner A, et al. Advanced biological ageing predicts future risk for neurological diagnoses and clinical examination findings. Brain. 2023;146(12):4891–902.
Mak JKL, McMurran CE, Hägg S. Clinical biomarker-based biological ageing and future risk of neurological disorders in the UK Biobank. J Neurol Neurosurg Psychiatry. 2024;95(5):481–4. https://doi.org/10.1136/jnnp-2023-331917.
Sugden K, Caspi A, Elliott ML, Bourassa KJ, Chamarti K, Corcoran DL, et al. Association of pace of aging measured by blood-based DNA methylation with age-related cognitive impairment and dementia. Neurology. 2022;99(13):e1402–13.
Zhou A, Wu Z, Zaw Phyo AZ, Torres D, Vishwanath S, Ryan J. Epigenetic aging as a biomarker of dementia and related outcomes: a systematic review. Epigenomics. 2022;14(18):1125–38.
Smith GD, Ebrahim S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int J Epidemiol. 2003;32(1):1–22.
Skrivankova VW, Richmond RC, Woolf BAR, Davies NM, Swanson SA, VanderWeele TJ, et al. Strengthening the reporting of observational studies in epidemiology using Mendelian randomisation (STROBE-MR): explanation and elaboration. BMJ. 2021;375:n2233.
Davies NM, Holmes MV, Davey SG. Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. BMJ. 2018;362:k601.
Codd V, Wang Q, Allara E, Musicha C, Kaptoge S, Stoma S, et al. Polygenic basis and biomedical consequences of telomere length variation. Nat Genet. 2021;53(10):1425–33.
McCartney DL, Min JL, Richmond RC, Lu AT, Sobczyk MK, Davies G, et al. Genome-wide association studies identify 137 genetic loci for DNA methylation biomarkers of aging. Genome Biol. 2021;22:1–25.
Longchamps RJ, Yang SY, Castellani CA, Shi W, Lane J, Grove ML, et al. Genome-wide analysis of mitochondrial DNA copy number reveals loci implicated in nucleotide metabolism, platelet activation, and megakaryocyte proliferation. Hum Genet. 2022;141(1):127–46.
Kuo CL, Pilling LC, Liu Z, Atkins JL, Levine ME. Genetic associations for two biological age measures point to distinct aging phenotypes. Aging Cell. 2021;20(6):e13376.
Pei Y-F, Liu Y-Z, Yang X-L, Zhang H, Feng G-J, Wei X-T, et al. The genetic architecture of appendicular lean mass characterized by association analysis in the UK Biobank study. Communications biology. 2020;3(1):608.
Higbee DH, Granell R, Smith GD, Dodd JW. Prevalence, risk factors, and clinical implications of preserved ratio impaired spirometry: a UK Biobank cohort analysis. Lancet Respir Med. 2022;10(2):149–57.
Ahadi S, Wilson KA, Babenko B, McLean CY, Bryant D, Pritchard O, et al. Longitudinal fundus imaging and its genome-wide association analysis provide evidence for a human retinal aging clock. Elife. 2023;12. https://doi.org/10.7554/eLife.82364
Lee JJ, Wedow R, Okbay A, Kong E, Maghzian O, Zacher M, et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat Genet. 2018;50(8):1112–21.
Leonardsen EH, Vidal-Piñeiro D, Roe JM, Frei O, Shadrin AA, Iakunchykova O, et al. Genetic architecture of brain age and its causal relations with brain and mental disorders. Mol Psychiatry. 2023;28(7):3111–20.
Atkins JL, Jylhävä J, Pedersen NL, Magnusson PK, Lu Y, Wang Y, et al. A genome-wide association study of the frailty index highlights brain pathways in ageing. Aging Cell. 2021;20(9):e13459.
Timmers PR, Mounier N, Lall K, Fischer K, Ning Z, Feng X, et al. Genomics of 1 million parent lifespans implicates novel pathways and common diseases and distinguishes survival chances. Elife. 2019;8:e39856.
Bellenguez C, Küçükali F, Jansen IE, Kleineidam L, Moreno-Grau S, Amin N, et al. New insights into the genetic etiology of Alzheimer’s disease and related dementias. Nat Genet. 2022;54(4):412–36.
Elsworth B, Lyon M, Alexander T, Liu Y, Matthews P, Hallett J, et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv. 2020. https://doi.org/10.1101/2020.08.10.244293.
Kurki MI, Karjalainen J, Palta P, Sipilä TP, Kristiansson K, Donner KM, et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature. 2023;613(7944):508–18.
Malik R, Chauhan G, Traylor M, Sargurupremraj M, Okada Y, Mishra A, et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat Genet. 2018;50(4):524–37.
Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell. 2013;49(2):359–67.
Horvath S, Raj K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat Rev Genet. 2018;19(6):371–84.
Levine ME, Lu AT, Quach A, Chen BH, Assimes TL, Bandinelli S, et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (albany NY). 2018;10(4):573.
Lu AT, Quach A, Wilson JG, Reiner AP, Aviv A, Raj K, et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (albany NY). 2019;11(2):303.
Levine ME. Modeling the rate of senescence: can estimated biological age predict mortality more accurately than chronological age? J Gerontol Ser A Biomed Sci Med Sci. 2013;68(6):667–74.
Esteban-Cornejo I, Ho FK, Petermann-Rocha F, Lyall DM, Martinez-Gomez D, Cabanas-Sánchez V, et al. Handgrip strength and all-cause dementia incidence and mortality: findings from the UK Biobank prospective cohort study. J Cachexia Sarcopenia Muscle. 2022;13(3):1514–25.
Graham BL, Steenbruggen I, Miller MR, Barjaktarevic IZ, Cooper BG, Hall GL, et al. Standardization of spirometry 2019 update. An official American thoracic society and European respiratory society technical statement. Am J Respir Crit Care Med. 2019;200(8):e70–88.
Sanderson E, Richardson TG, Hemani G, Davey SG. The use of negative control outcomes in Mendelian randomization to detect potential population stratification. Int J Epidemiol. 2021;50(4):1350–61.
Bowden J, Davey Smith G, Haycock PC, Burgess S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol. 2016;40(4):304–14.
Bowden J, Davey Smith G, Burgess S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol. 2015;44(2):512–25.
Verbanck M, Chen C-Y, Neale B, Do R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet. 2018;50(5):693–8.
Burgess S, Smith GD, Davies NM, Dudbridge F, Gill D, Glymour MM, et al. Guidelines for performing Mendelian randomization investigations: update for summer 2023. Wellcome Open Res. 2019;4:186.
Forero DA, González-Giraldo Y, López-Quintero C, Castro-Vega LJ, Barreto GE, Perry G. Meta-analysis of telomere length in Alzheimer’s disease. J Gerontol Ser A Biomed Sci Med Sci. 2016;71(8):1069–73.
Haycock PC, Burgess S, Nounu A, Zheng J, Okoli GN, Bowden J, et al. Association between telomere length and risk of cancer and non-neoplastic diseases: a Mendelian randomization study. JAMA Oncol. 2017;3(5):636–51.
Beeri MS, Leugrans SE, Delbono O, Bennett DA, Buchman AS. Sarcopenia is associated with incident Alzheimer’s dementia, mild cognitive impairment, and cognitive decline. J Am Geriatr Soc. 2021;69(7):1826–35.
Cui C, Mackey RH, Shaaban CE, Kuller LH, Lopez OL, Sekikawa A. Associations of body composition with incident dementia in older adults: Cardiovascular Health Study-Cognition Study. Alzheimers Dement. 2020;16(10):1402–11.
Kojima G, Taniguchi Y, Iliffe S, Walters K. Frailty as a predictor of Alzheimer disease, vascular dementia, and all dementia among community-dwelling older people: a systematic review and meta-analysis. J Am Med Dir Assoc. 2016;17(10):881–8.
Borges MK, Canevelli M, Cesari M, Aprahamian I. Frailty as a predictor of cognitive disorders: a systematic review and meta-analysis. Front Med. 2019;6:26.
Zheng L, Li G, Gao D, Wang S, Meng X, Wang C, et al. Cognitive frailty as a predictor of dementia among older adults: a systematic review and meta-analysis. Arch Gerontol Geriatr. 2020;87:103997.
Kojima G, Iliffe S, Walters K. Frailty index as a predictor of mortality: a systematic review and meta-analysis. Age Ageing. 2018;47(2):193–200.
Murabito JM, Zhao Q, Larson MG, Rong J, Lin H, Benjamin EJ, et al. Measures of biologic age in a community sample predict mortality and age-related disease: the Framingham Offspring Study. J Gerontol Ser A. 2018;73(6):757–62.
Chan MS, Arnold M, Offer A, Hammami I, Mafham M, Armitage J, et al. A biomarker-based biological age in UK Biobank: composition and prediction of mortality and hospital admissions. J Gerontol Ser A. 2021;76(7):1295–302.
Shay JW. Telomeres and aging. Curr Opin Cell Biol. 2018;52:1–7.
López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013;153(6):1194–217.
Granic A, Suetterlin K, Shavlakadze T, Grounds MD, Sayer AA. Hallmarks of ageing in human skeletal muscle and implications for understanding the pathophysiology of sarcopenia in women and men. Clin Sci. 2023;137(22):1721–51.
Arosio B, Calvani R, Ferri E, Coelho-Junior HJ, Carandina A, Campanelli F, et al. Sarcopenia and cognitive decline in older adults: targeting the muscle–brain axis. Nutrients. 2023;15(8):1853.
Jo D, Yoon G, Kim OY, Song J. A new paradigm in sarcopenia: cognitive impairment caused by imbalanced myokine secretion and vascular dysfunction. Biomed Pharmacother. 2022;147:112636.
Jiang T, Gill D, Butterworth AS, Burgess S. An empirical investigation into the impact of winner’s curse on estimates from Mendelian randomization. Int J Epidemiol. 2023;52(4):1209–19.
Sadreev II, Elsworth BL, Mitchell RE, Paternoster L, Sanderson E, Davies NM, et al. Navigating sample overlap, winner’s curse and weak instrument bias in Mendelian randomization studies using the UK Biobank. MedRxiv. 2021:2021.06. 28.21259622. https://doi.org/10.1101/2021.06.28.21259622
Abdellaoui A, Hugh-Jones D, Yengo L, Kemper KE, Nivard MG, Veul L, et al. Genetic correlates of social stratification in Great Britain. Nat Hum Behav. 2019;3(12):1332–42.
Funding
Open access funding provided by Karolinska Institute. The research was conducted with support from the Swedish Research Council, Karolinska Institutet Foundation, Karolinska Institutet Joint NIH-grant for doctoral studies, and National Institute on Aging (NIA) R01AG067996.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Imahori, Y., Qin, C., Tang, B. et al. Comprehensive analysis of molecular, physiological, and functional biomarkers of aging with neurological diseases using Mendelian randomization. GeroScience (2024). https://doi.org/10.1007/s11357-024-01334-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11357-024-01334-6