Abstract
Heme is an essential prosthetic group that serves as a co-factor and a signaling molecule. Heme levels decline with age, and its deficiency is associated with multiple hallmarks of aging, including anemia, mitochondrial dysfunction, and oxidative stress. Dysregulation of heme homeostasis has been also implicated in aging in model organisms suggesting that heme may play an evolutionarily conserved role in controlling lifespan. However, the underlying mechanisms and whether heme homeostasis can be targeted to promote healthy aging remain unclear. Here, we used Saccharomyces cerevisiae as a model to investigate the role of heme in aging. For this, we have engineered a heme auxotrophic yeast strain expressing a plasma membrane-bound heme permease from Caenorhabditis elegans (ceHRG-4). This system can be used to control intracellular heme levels independently of the biosynthetic enzymes by manipulating heme concentration in the media. We observed that heme supplementation leads to a significant extension of yeast replicative lifespan. Our findings revealed that the effect of heme on lifespan is independent of the Hap4 transcription factor. Surprisingly, heme-supplemented cells had impaired growth on YPG medium, which requires mitochondrial respiration to be used, suggesting that these cells are respiratory deficient. Together, our results demonstrate that heme homeostasis is fundamentally important for aging biology, and manipulating heme levels can be used as a promising therapeutic target for promoting longevity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Heme is an essential iron-containing tetrapyrrole that serves both as a co-factor and a signaling molecule. As a protein prosthetic group, it is involved in different processes including electron transfer [1], chemical catalysis [2], and oxygen transport [3, 4]. It also serves as an important signaling molecule that regulates transcription factors controlling oxidative phosphorylation, antioxidant defense, and proliferation [5, 6]. Although heme is essential for health, it may also cause toxicity if present in excess [7]. Therefore, body heme levels are tightly regulated. Dysregulation of heme homeostasis has been implicated in the development of several diseases. In humans, defects in heme synthesis result in the accumulation of its intermediates leading to porphyria [8]. In addition, previous studies have shown that heme synthesis declines with age, and its deficiency leads to mitochondrial dysfunction and iron accumulation leading to oxidative stress [9, 10]. On the other hand, deficiency of heme oxygenases, enzymes that are necessary for heme degradation, has been implicated in the development of several age-related disorders, such as Alzheimer’s disease, cancer as well as cardiovascular and metabolic diseases [11]. Moreover, recent genome-wide association studies (GWAS) have identified several genetic variants in genes involved in heme metabolism that are associated with human aging [12]. However, the molecular mechanisms by which heme regulates aging remain unclear.
Yeast Saccharomyces cerevisiae has proven to be a useful model for understanding the basic mechanisms by which dysregulation of heme homeostasis contributes to aging. Previous studies have shown that heme levels decrease with aging in yeast, which is associated with the decline of mitochondrial function [13]. The effects of heme on yeast lifespan were attributed to its direct functions as a co-factor in the electron transport chain in mitochondria and activating the heme activator protein (HAP) transcriptional complex which induces genes for maintaining mitochondrial biogenesis and function [14, 15]. Consistently, overexpression of Hap4, a component of the HAP complex, significantly extends lifespan [16]. Although increasing evidence suggests an important link between heme homeostasis and aging, the underlying mechanisms and causes of the decline in heme levels during aging are not completely understood.
In the present study, we developed a model to investigate the impact of heme on the yeast replicative lifespan. To distinguish the effects of endogenously produced heme from the effects of exogenous heme acquired from the media, we generated a heme auxotroph yeast strain, hem1Δ, which is unable to synthesize heme. Additionally, we introduced a membrane-bound heme permease from C. elegans into the yeast genome. This approach enabled us to directly control intracellular heme levels, independently of biosynthetic enzymes, by manipulating heme concentration in the media. Our findings demonstrate that supplementing heme, but not its precursor 5-aminolevulinic acid (ALA), is sufficient to extend replicative lifespan in S. cerevisiae. Conversely, overexpression of the Hmx1 heme oxygenase, which is involved in heme degradation, results in a shortened replicative lifespan. Furthermore, we observed that heme supplementation extends lifespan independently of the Hap4 transcription factor, resulting in the inability of yeast cells to grow on respiratory carbon sources, indicating an adverse effect on mitochondrial function. Taken together, our findings reveal a crucial role of heme in the regulation of yeast replicative lifespan, laying the foundation for the development of heme-based therapeutic strategies for age-dependent diseases in humans.
Methods
Yeast strains and culture conditions
The yeast strains used in this study and their genotypes are listed in Table S1. One-step polymerase chain reaction (PCR)-mediated gene disruption was performed using standard techniques to delete genes of interest. The genotypes of the resulting strains were verified using colony PCR.
To generate a yeast heme auxotroph strain expressing ceHRG-4, the sequence containing ceHRG-4 was first PCR amplified from pYES-DEST52-ceHRG-4 plasmid (pPP96) [17] and integrated into p416-GPD [18] vector using BamHI and XhoI restriction sites to generate pPP97 plasmid. The sequence encoding ceHRG-4 under the control of the glyceraldehyde-3-phosphate dehydrogenase promoter (GPD1pr-ceHRG-4) along with the URA3 marker was then amplified from pPP97 and integrated into the hem1Δ strain using homologous recombination at URA3 locus. The resulting transformants were selected on SC-URA plates and verified by colony PCR and Sanger sequencing.
To generate the HMX1-OE strain expressing HMX1 under the control of the ADH1 promoter (ADH1pr-HMX1), a 705 bp sequence of the ADH1 promoter was amplified by PCR from yeast genomic DNA and integrated into the pRS306 [19] plasmid using NotI and XhoI restriction sites. Subsequently, the fragment encompassing the URA3 marker and the ADH1 promoter was amplified via PCR using the oPP399 and oPP400 oligonucleotides. These oligonucleotides have homology to the 3′ end of the HMX1 promoter and the 5′ end of the HMX1 ORF, respectively. The resulting PCR product was then utilized for genomic integration. To create the HAP4-OE strain, the fragment containing the URA3 marker and the ADH1 promoter was amplified using the oPP274 and oPP275 oligonucleotides and was integrated into the genome upstream of the HAP4 ORF. To create the yeast strain encoding an HA-tagged Hap4 protein, HA-tag sequence along with KanMX4 marker was amplified from pPP111 plasmid using the primers flanked on the 5′ side by sequence homologous to HAP4 ORF and the 3′ side by sequence homologous to HAP4 3′-UTR for genomic integration. The resulting transformants were selected on YPD plates containing 200 µg/mL G418 and verified by colony PCR and Sanger sequencing. The plasmids used in this study and sequences of primers used for generating yeast strains are listed in Table S2 and Table S3.
Yeast strains were cultured at 30 °C in standard YPD medium (1.0% yeast extract, 2.0% peptone, and 2.0% glucose) unless otherwise stated. hem1Δ cells were maintained in YPD supplemented with 250 µM 5-aminolevulinic acid (ALA). To test the effect of exogenous heme on replicative and chronological lifespan, the media was supplemented with hemin chloride. The stock hemin, chloride solution was prepared by dissolving hemin in 0.3 M NH4OH. The pH was adjusted to 8 with 6N HCl, and the mixture was filter-sterilized through a 0.2-μm filter.
Spot assays
The spot assays were used to assess the growth of strains under various growth conditions. Each strain to be tested was inoculated into a liquid culture medium and allowed to grow to the exponential phase. Once the cultures reached OD600 = 0.6, 10 × serial dilutions for each strain were spotted on YPD agar plates (containing 2% glucose) or YPG plates (contaminating 3% glycerol) supplemented with indicated concentrations of ALA or heme. The plates were incubated at 30 °C for 48 h before they were imaged.
Replicative lifespan analysis
Replicative lifespan assays were carried out as described previously [20]. Cells were cultured on freshly prepared YPD plates at 30 °C. Cells were monitored for cell divisions, and the subsequent daughter cells were removed using a micromanipulator. The replicative lifespan was determined by counting the number of divisions each mother cell underwent before it ceased dividing. The lifespan assays were conducted at least two times, and the data from separate biological replicates were combined. The number of cells assayed, and statistical analysis of the lifespan data are shown in Table S4.
Chronological lifespan assay
Yeast CLS assays were performed as described [21]. Briefly, single colonies for each strain were inoculated into 10 mL of synthetic complete dextrose media (SCD) and cultured for 4 days at 30 °C. Aliquots of each culture were serially diluted in SCD media until achieving a cell density of roughly 200 cells per 100 μL. Following this, 100 μL of the dilution was plated onto YPD agar plates, which were then kept at 30 °C for the indicated time, and colonies were manually counted. Colony forming units (CFU) were normalized relative to the CFUs counted on day 0. The averages and standard errors of the mean were calculated based on at least six biological replicates.
RT-qPCR
Total RNA was extracted using hot acid phenol method followed by purification using Direct-zol RNA Miniprep Kit (Zymo Research). The RNA was treated with DNaseI, and 2 µg of RNA was used for cDNA synthesis using SuperScript III reverse transcriptase (Thermo Fisher Scientific) with random hexamer primers according to the manufacturer’s instructions. To analyze mRNA expression, real-time PCR was performed using SYBR Fast qPCR Master Mix (Kapa Biosystems) and the CFX-96 Touch Real-Time PCR Detection System (Bio-Rad Laboratories). To normalize the gene expression, ACT1 was used as the reference gene. The primers used for RT-qPCR are listed in Table S3. Results are represented as means ± SEM from at least three independent experiments.
Western blot analysis
A single colony was inoculated in 3 mL of YPD medium and incubated overnight at 30 °C. The next day, the cells were diluted to an OD600 of 1 in 10 mL of fresh medium and grown until they reached the log phase. The cells were collected by centrifugation and washed with sterile water, and the pellet was frozen in liquid nitrogen and stored at − 80 °C for later use. For protein extraction, the cell pellet was re-suspended in 300 µL of lysis buffer (50 mM Tris pH 7.5, 150 mM NaCl, 1 mM EDTA, and 1% DMSO) containing 1 mM phenylmethylsulfonyl fluoride (PMSF). The samples were homogenized with glass beads by vortexing at maximum speed for seven 30-s cycles and centrifuged for 5 min at 12,000 rpm at 4 °C to obtain the supernatant. Equal amounts of proteins were resolved in 10% SDS-PAGE gels and transferred to PVDF membrane. The membrane was then probed with horseradish peroxidase (HRP)-conjugated HA tag monoclonal antibody (2–2.2.14) (ThermoFisher) using 1:1000 dilution, and signal quantification was performed using ImageJ software. Mouse anti-Pgk1 monoclonal antibody (1:5000, Life Technologies) and HRP-conjugated secondary anti-mouse antibody (1:100,000, Santa Cruz Biotechnology) were used to detect Pgk1 protein levels as a loading control.
Heme quantification
Intracellular heme levels were assessed using the oxalic acid method as previously described with minor changes [22]. Briefly, overnight yeast cultures were diluted to OD600 = 0.2 units/mL, and grown until cells reached OD600 = 0.8 units/mL. Subsequently, 8 OD600 units of cells were harvested by centrifugation at 2500 × g and washed with distilled water, and the pellet was resuspended in 500 µL of 20 mM oxalic acid. An additional 500 µL of 2 M oxalic acid was added, and the suspension was divided equally into two tubes. One set of sample tubes was placed in a heating block at 100 °C for 45 min, while the other set of samples was kept at room temperature for the same duration as a baseline. Following incubation, samples were cooled to room temperature, and 100 µL of suspension was transferred per well into a black-well 96-well plate in duplicates. The fluorescence of porphyrin was measured using a BioTek plate reader with 400 nm excitation and 662 nm emission. Baseline values (from parallel unheated samples in oxalic acid) were subtracted, and the relative fluorescence intensity in arbitrary units (A.F.U.) was plotted.
Quantification and statistical analysis
Statistical analysis was performed using Prism 9.3.1 (GraphPad Software, Inc.). The statistical significance of the heme quantification and RT-qPCR data was determined by calculating p values using one-way ANOVA. Error bars represent standard errors of the mean (SEM). The statistical significance of the lifespan data was evaluated using the Wilcoxon Rank-Sum test [23].
Results
Expression of C. elegans HRG-4 enhances heme import in yeast
The ability of yeast cells to synthesize heme might complicate the examination of how externally supplemented heme regulates the lifespan of S. cerevisiae. In order to avoid the effect of endogenously produced heme, we first introduced heme auxotrophy in yeast by deleting the HEM1 gene encoding for the first enzyme of the heme biosynthesis pathway (Fig. 1A). Consistent with prior reports, cells lacking HEM1 exhibited impaired growth unless supplemented with 5-aminolevulinic acid (ALA) or an excess of exogenous heme (provided as hemin chloride) in the growth medium [24]. Because S. cerevisiae utilizes exogenous heme inefficiently, we introduced a gene encoding a plasma membrane-bound permease from C. elegans (ceHRG-4) into the yeast genome (Fig. 1B). The growth of hem1Δ cells expressing ceHRG-4 was rescued by a lower concentration of heme (10 µM) compared to the hem1Δ mutant, indicating a more efficient import of heme into the yeast cells in the presence of ceHRG-4 expression. Together, our data suggest that expressing ceHRG-4 increases heme uptake in yeast, allowing for precise control of intracellular heme levels by manipulating heme concentration in the media independently of heme biosynthetic enzymes.
Engineering a heme auxotroph yeast strain to study the role of heme in aging. A Cells lacking HEM1 have impaired growth in the absence of ALA or heme. The hem1Δ strain was cultured in YPD media containing 250 µM ALA overnight, and serial (10x) dilutions were spotted on YPD plates supplemented with the indicated concentrations of ALA or heme. Plates were incubated at 30 °C for 2 days prior to imaging. B Expression of ceHRG-4 enhances heme import in yeast. The hem1Δ and hem1Δ ceHRG-4 strains were cultured in YPD media containing 250 µM ALA overnight, and serial (10x) dilutions were spotted on YPD plates supplemented with the indicated concentrations of heme. A YPD plate without heme was used as a control. Plates were incubated at 30 °C for 2 days prior to imaging
Heme supplementation extends yeast replicative lifespan
To investigate how heme levels affect aging, we analyzed yeast replicative lifespan in hem1Δ cells expressing ceHRG-4 in the presence of different concentrations of heme in the media. Although hem1Δ-ceHRG-4 cells are unable to grow in the absence of heme, we found that increasing heme concentration from 10 to 100 µM extended lifespan by 67.8% (p < 0.001) (Fig. 2A). Similarly, wild-type cells expressing ceHRG-4 exhibited significant lifespan extension (46.3%; p < 0.0001) when supplemented with 100 µM heme (Fig. 2B). However, supplementing yeast with 5-aminolevulinic acid (ALA), a heme biosynthesis precursor, did not result in a statistically significant difference in lifespan (Fig. 2C). Additionally, we analyzed the effect of heme supplementation on chronological lifespan (CLS). Similar to replicative lifespan, CLS was significantly extended by heme supplementation (Supplementary Figure S1). To validate that the effect of heme on lifespan is mediated by increased heme levels, we measured intracellular heme concentration in cells supplemented with different concentrations of heme and ALA (Fig. 2D). We found that supplementing heme to the media resulted in a drastic increase in heme levels in cells expressing ceHRG-4. Conversely, ALA supplementation did not significantly alter intracellular heme levels, even at high concentrations.
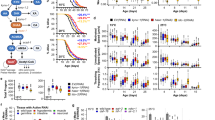
Heme supplementation extends yeast replicative lifespan. A Supplementing heme extends the replicative lifespan in hem1Δ cells expressing ceHRG-4. Lifespan of the hem1Δ ceHRG-4 mutant was analyzed on YPD plates supplemented with indicated concentrations of heme. The replicative lifespan assays were conducted at least two times, and the data from separate biological replicates were combined. The average lifespan is shown in parentheses. B Replicative lifespan of wild-type cells expressing ceHRG-4. C Supplementing 5-aminolevulinic acid (ALA) does not extend replicative lifespan in yeast. D Spectrofluorometric quantification of intracellular heme levels in wild-type cells expressing ceHRG-4 supplemented with indicated concentrations of heme and ALA. Error bars represent SEM of three biological replicates, each containing two technical replicates. **p < 0.01, ***p < 0.001 compared with ceHRG-4 control (one-way ANOVA). E Shortened lifespan in cells overexpressing HMX1 (HMX1-OE) can be rescued by heme supplementation. F Increased lifespan in yeast cells supplemented with heme correlates with intracellular heme concentration. Error bars represent SEM of three biological replicates, each containing two technical replicates. *p < 0.05, **p < 0.01 compared with wild-type control (one-way ANOVA)
Given the beneficial effect of heme supplementation on lifespan, we sought to investigate the effects of decreasing intracellular heme levels on lifespan. To this end, we overexpressed the HMX1 gene, which encodes heme oxygenase involved in intracellular heme degradation. HMX1 overexpression (HMX1-OE) decreased lifespan by 16.4% compared to wild-type cells (Fig. 2E). Furthermore, supplementing the HMX1-OE strain with heme was able to rescue the shortened lifespan caused by the HMX1 overexpression, and this increase in lifespan correlated with intracellular heme concentration (Fig. 2F). Together, these findings suggest that heme plays a critical role in the regulation of replicative lifespan in yeast.
Heme extends yeast lifespan independently of Hap4
Previous studies have shown that supplementation of yeast cells with heme or its precursors results in an increased expression of the Hap4 transcription factor [25]. Hap4 plays a key role in the regulation of mitochondrial function and respiration in yeast [26, 27]. Furthermore, overexpression of HAP4 has been shown to extend the yeast lifespan [16]. Based on these observations, we hypothesized that the extended lifespan resulting from heme supplementation could be attributed to the overexpression of Hap4. To test this hypothesis, we first asked whether heme supplementation would lead to increased Hap4 expression. For this, we quantified the levels of the Hap4 protein in the cells treated with different concentrations of heme using Western blot analysis (Fig. 3A). Contrary to expectations, we found that Hap4 protein levels were significantly decreased (p < 0.001) in the presence of heme supplementation compared to the control cells.
Heme supplementation inhibits expression of Hap4. A Hap4 protein levels are repressed by heme. The expression of Hap4-HA in cells treated with different concentrations of heme was assessed by Western blotting using an anti-HA antibody. Pgk1 was used as a loading control. Quantification of protein expression was carried out using ImageJ software. Error bars represent SEM of three independent experiments. B Heme supplementation does not affect the proteasomal degradation of Hap4. pdr5Δ cells were treated with 200 µM heme in the presence or absence of 75 µM MG132 proteasome inhibitor. Hap4 protein expression was quantified using ImageJ software. Error bars represent SEM of three independent experiments. *p < 0.05; ***p < 0.001 (one-way ANOVA). C The expression of HAP4 and its targets in exponentially growing cells was assessed using RT-qPCR. Error bars represent SEM of three independent experiments. *p < 0.05; **p < 0.01 (one-way ANOVA). D Heme supplementation extends replicative lifespan independently of HAP4. The lifespan of the wild-type (WT) and hap4Δ cells was analyzed on YPD plates in the presence or absence of 200 µM heme. The replicative lifespan data from three biological replicates were combined. The average lifespan is shown in parentheses
It has been previously shown that Hap4 is susceptible to proteasome-mediated degradation [28]. To test whether decreased Hap4 protein levels can be attributed to its increased proteasomal degradation, we treated cells with heme in the presence or absence of MG132 proteasome inhibitor [29]. Because S. cerevisiae is resistant to MG132 [30], we employed the pdr5Δ strain with reduced drug efflux for experiments using proteasome inhibitors [31]. When pdr5Δ cells were supplemented with 75 µM MG132, degradation of Hap4 by proteasome was prevented (Supplementary Figure S2A). However, MG132 treatment did not counteract the negative impact of heme supplementation on Hap4 protein levels (Fig. 3B), suggesting that the decrease in Hap4 levels is not due to proteasomal degradation. Furthermore, the addition of the proteasome inhibitor was not able to rescue the growth of ceHRG-4-expressing cells treated with heme on glycerol-containing medium (YPG) (Supplementary Figure S2B). Additionally, we analyzed the abundance of HAP4 mRNA and expression of the Hap4 transcription factor targets, including KGD1, ACO1, and SDH1 (Fig. 3C) using RT-qPCR. We observed a significant decrease in HAP4 mRNA levels (p < 0.001) and diminished expression of the Hap4 targets when yeast cells were exposed to high heme concentrations.
To directly test if the increased lifespan in response to heme supplementation requires Hap4, we analyzed replicative lifespan in wild-type cells and the HAP4 deletion mutant (hap4Δ) in the absence and presence of heme supplementation (Fig. 3D). Although the deletion of HAP4 resulted in a shortened replicative lifespan (p < 0.001) compared to the wild-type strain, the supplementation of hap4Δ with heme was sufficient to significantly extend the lifespan. Taken together, our findings suggest that heme supplementation causes a reduction in HAP4 expression, and that the lifespan extension effect of heme cannot be attributed to the increased activity of the Hap4 transcription factor.
Heme supplementation negatively affects respiration
Since Hap4 is involved in regulating mitochondrial biogenesis in yeast [26, 27], we investigated whether the heme-induced reduction in HAP4 expression would result in decreased cell respiration. To explore this, we examined the growth of yeast cells supplemented with varying heme concentrations on YPG plates containing glycerol as a carbon source that requires respiration to be utilized (Fig. 4A). Notably, we observed a decline in yeast growth on YPG plates as heme concentration increased. Additionally, heme supplementation inhibited cellular growth when cells were cultured in media containing alternative carbon sources that rely on respiration, including ethanol and lactate (Fig. 4B). We further examined the impact of heme supplementation on population doubling time in glucose-containing medium (YPD) and glycerol-containing medium (YPG) that forces cells to use respiration. Our results show that ceHRG-4-expressing cells treated with heme display normal growth during the exponential phase (fermentative growth) on a YPD medium containing 2% glucose (Fig. 4C). However, once glucose was depleted during the diauxic shift, these cells were unable to switch metabolism from fermentation to respiration leading to impaired growth compared to untreated cells. Moreover, overexpression of HAP4 (HAP4-OE) was able to restore the growth of ceHRG-4-expressing cells in YPG media supplemented with heme (Fig. 4D and Supplementary Figure S3). Together, our findings indicate that heme supplementation negatively affects Hap4 expression and results in decreased respiratory capacity.
Heme supplementation negatively affects respiration. A Supplementation of heme leads to delayed growth of wild-type and ceHRG-4 expressing cells on glycerol-containing medium (YPG). Yeast strains were cultured in YPD media overnight, and serial (10x) dilutions were spotted on YPD and YPG plates containing indicated concentrations of heme. Plates were incubated at 30 °C for 2 days prior to imaging. B Heme supplementation inhibits cellular growth in media containing alternative carbon sources that rely on respiration. Yeast strains were cultured in YPD media overnight, and serial (10x) dilutions were spotted on agar plates with YPD (2% glucose), YPG (3% glycerol), YPE (5% ethanol), and YPL (3% lactate) media in the presence or absence of 50 µM heme. Plates were incubated at 30 °C for 2 days prior to imaging. Representative images from two independent experiments are shown. C Heme supplementation leads to the growth inhibition of ceHRG-4-expressing cells during respiratory phase in YPD media. Representative growth curves from three independent experiments are shown. D HAP4 overexpression rescues the growth of ceHRG-4 expressing cells supplemented with heme on glycerol-containing medium (YPG). Representative growth curves from three independent experiments are shown
Discussion
Heme is an essential prosthetic group for enzymes involved in multiple biological functions. Defects in heme synthesis and transport have been associated with multiple human disorders, including anemia and mitochondrial dysfunction. However, the causes of the decline in heme levels during aging and mechanisms by which heme regulates lifespan are not understood. In this study, we used the S. cerevisiae model to investigate the role of heme in aging. To tightly regulate heme concentration, we induced heme auxotrophy in yeast through the deletion of the HEM1 gene. Since budding yeast utilizes exogenous heme inefficiently even in the absence of endogenous heme synthesis [24, 32], we employed a previously established strategy to manipulate heme uptake in yeast by overexpressing a plasma membrane-bound heme permease from C. elegans (ceHRG-4) [33].
How do intracellular heme levels affect lifespan in yeast? To distinguish the effects of endogenously produced heme from the effects of exogenous heme, we supplemented yeast cells with either ALA, a precursor of heme biosynthesis, or directly provided hemin in the media. Our findings revealed that heme supplementation significantly extends the yeast replicative lifespan. In contrast, supplementing ALA, which requires mitochondria to be utilized for the synthesis of intracellular heme, did not significantly affect lifespan. The observation that ALA supplementation did not extend lifespan may be attributed to low intracellular heme levels, possibly resulting from inefficient ALA transport or negative feedback regulation of heme biosynthesis enzymes.
Among the potential causes of the decline in heme levels during aging are increased heme degradation and dysregulation of heme biosynthesis. Using yeast as a model, we have recently shown that expression of HMX1, involved in heme degradation, is increased with aging whereas deletion of the HMX1 gene leads to lifespan extension [34]. The yeast HMX1 gene is regulated by the Aft1 transcription factor. During aging, increased activity of the Aft1 leads to increased expression of HMX1 resulting in decreased intracellular heme levels. Consistent with the role of Hmx1 in heme degradation, our data show that overexpression of HMX1 shortens lifespan (Fig. 2E).
Furthermore, the rate of heme production declines with aging. For example, the activity of the ALA synthase [35], a rate-limiting enzyme in heme biosynthesis, and ALA levels have been shown to decrease with aging [36, 37]. Notably, ALA supplementation was able to restore muscle function and extend the health span in Drosophila melanogaster [38]. Additionally, the mRNA-binding protein Cth2, which is involved in the regulation of heme biosynthesis enzymes, is aberrantly expressed with aging leading to the repression of heme synthesis [34]. On the other hand, mitochondria play a key role in heme biosynthesis. It is possible that reduced heme biosynthesis during aging is a consequence of mitochondrial dysfunction in aging cells.
The effects of heme on yeast lifespan can be attributed to its direct function as a cofactor in the electron transport chain (ETC) in mitochondria or serving as a signaling molecule. We hypothesized that heme could extend lifespan by activating the Hap4 transcription factor, which induces the expression of genes required for maintaining mitochondrial biogenesis and function. Surprisingly, we found that Hap4 levels were decreased in cells treated with heme. The decrease in Hap4 protein levels was not due to proteasomal degradation but instead was driven by changes in HAP4 mRNA abundance leading to the inhibition of Hap4 activity and the inability of cells to grow on respiratory carbon sources. We also found that heme supplementation extends lifespan independently of the Hap4 transcription factor. Since Hap4 expression is downregulated by heme and HAP4 is dispensable for its lifespan extension effects, these results suggest that heme supplementation may extend lifespan through alternative mechanisms, such as increased activity of enzymes requiring heme as a cofactor or activation of adaptive stress responses.
Limitations of the study
Together, our results suggest the existence of a previously unanticipated role of heme in the regulation of lifespan in yeast and raise an exciting possibility that increased heme levels may extend lifespan in other organisms. Previous studies have shown that inhibition of mitochondrial function as well as components of the ETC and TCA cycle can extend the lifespan in several species [39], including yeast [40], worms [41], flies [42], and mice [43]. For example, the downregulation of aconitase and isocitrate dehydrogenase, two enzymes in the TCA cycle, leads to lifespan extension in worms [44]. Additionally, a genetic screen in yeast measuring the replicative lifespan of 4698 deletion mutants identified mitochondrial translation and components of TCA among the most enriched functional groups [45]. However, specific genes and stress response pathways that connect these changes to longevity remain unknown. Further studies analyzing age-dependent changes in transcriptional network of cells in response to heme treatment may provide additional insights into pathways associated with lifespan extension. It is also important to note that yeast do not depend on mitochondrial respiration in the same way that many metazoan cells do under physiological conditions. Therefore, further research is needed to fully understand the mechanisms underlying the effect of heme on lifespan in mammalian cells and its potential implications for human aging.
References
Schenkman JB, Jansson I. The many roles of cytochrome b5. Pharmacol Ther. 2003;97(2):139–52.
Chen H, Ikeda-Saito M, Shaik S. Nature of the Fe-O2 bonding in oxy-myoglobin: effect of the protein. J Am Chem Soc. 2008;130(44):14778–90.
Gell DA. Structure and function of haemoglobins. Blood Cells Mol Dis. 2018;70:13–42.
Ordway GA, Garry DJ. Myoglobin: an essential hemoprotein in striated muscle. J Exp Biol. 2004;207(Pt 20):3441–6.
Zhang L, Guarente L. Heme binds to a short sequence that serves a regulatory function in diverse proteins. EMBO J. 1995;14(2):313–20.
Ogawa K, et al. Heme mediates derepression of Maf recognition element through direct binding to transcription repressor Bach1. EMBO J. 2001;20(11):2835–43.
Dutt S, Hamza I, Bartnikas TB. Molecular mechanisms of iron and heme metabolism. Annu Rev Nutr. 2022;42:311–35.
Severance S, Hamza I. Trafficking of heme and porphyrins in metazoa. Chem Rev. 2009;109(10):4596–616.
Shetty T, et al. Heme synthesis inhibition blocks angiogenesis via mitochondrial dysfunction. iScience. 2020;23(8):101391.
Atamna H, et al. Heme deficiency may be a factor in the mitochondrial and neuronal decay of aging. Proc Natl Acad Sci U S A. 2002;99(23):14807–12.
Yachie A. Heme oxygenase-1 deficiency and oxidative stress: a review of 9 independent human cases and animal models. Int J Mol Sci. 2021;22:4.
Timmers P, et al. Multivariate genomic scan implicates novel loci and haem metabolism in human ageing. Nat Commun. 2020;11(1):3570.
Li Y, et al. A programmable fate decision landscape underlies single-cell aging in yeast. Science. 2020;369(6501):325–9.
Buschlen S, et al. The. S cerevisiae HAP complex, a key regulator of mitochondrial function, coordinates nuclear and mitochondrial gene expression. Comp Funct Genomics. 2003;4(1):37–46.
Feng MW, Adams PD. A new mechanistic insight into fate decisions during yeast cell aging process. Mech Ageing Dev. 2021;198:111542.
Lin SJ, et al. Calorie restriction extends Saccharomyces cerevisiae lifespan by increasing respiration. Nature. 2002;418(6895):344–8.
Yuan X, et al. Topologically conserved residues direct heme transport in HRG-1-related proteins. J Biol Chem. 2012;287(7):4914–24.
Mumberg D, Muller R, Funk M. Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene. 1995;156(1):119–22.
Sikorski RS, Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989;122(1):19–27.
Steffen KK, Kennedy BK, Kaeberlein M. Measuring replicative life span in the budding yeast. J Vis Exp. 2009(28):e1209. https://doi.org/10.3791/1209.
Murakami CJ, et al. A method for high-throughput quantitative analysis of yeast chronological life span. J Gerontol A Biol Sci Med Sci. 2008;63(2):113–21.
Sinclair PR, Gorman N, Jacobs JM. Measurement of heme concentration. Curr Protoc Toxicol. 2001;8:8.3.1–8.3.7. https://doi.org/10.1002/0471140856.tx0803s00.
Wilcoxon F. Individual comparisons of grouped data by ranking methods. J Econ Entomol. 1946;39:269.
Protchenko O, et al. Role of PUG1 in inducible porphyrin and heme transport in Saccharomyces cerevisiae. Eukaryot Cell. 2008;7(5):859–71.
Zhang T, et al. Increased heme synthesis in yeast induces a metabolic switch from fermentation to respiration even under conditions of glucose repression. J Biol Chem. 2017;292(41):16942–54.
Forsburg SL, Guarente L. Identification and characterization of HAP4: a third component of the CCAAT-bound HAP2/HAP3 heteromer. Genes Dev. 1989;3(8):1166–78.
Bourgarel D, Nguyen CC, Bolotin-Fukuhara M. HAP4, the glucose-repressed regulated subunit of the HAP transcriptional complex involved in the fermentation-respiration shift, has a functional homologue in the respiratory yeast Kluyveromyces lactis. Mol Microbiol. 1999;31(4):1205–15.
Capps D, et al. Ubiquitin-conjugating enzymes Ubc1 and Ubc4 mediate the turnover of Hap4, a master regulator of mitochondrial biogenesis in Saccharomyces cerevisiae. Microorganisms. 2022;10:12.
Lee DH, Goldberg AL. Proteasome inhibitors: valuable new tools for cell biologists. Trends Cell Biol. 1998;8(10):397–403.
Lee DH, Goldberg AL. Proteasome inhibitors cause induction of heat shock proteins and trehalose, which together confer thermotolerance in Saccharomyces cerevisiae. Mol Cell Biol. 1998;18(1):30–8.
Fleming JA, et al. Complementary whole-genome technologies reveal the cellular response to proteasome inhibition by PS-341. Proc Natl Acad Sci U S A. 2002;99(3):1461–6.
Protchenko O, et al. A screen for genes of heme uptake identifies the FLC family required for import of FAD into the endoplasmic reticulum. J Biol Chem. 2006;281(30):21445–57.
Chen C, Hamza I. Notes from the underground: heme homeostasis in C. elegans. Biomolecules. 2023;13:7.
Patnaik PK, et al. Deficiency of the RNA-binding protein Cth2 extends yeast replicative lifespan by alleviating its repressive effects on mitochondrial function. Cell Rep. 2022;40(3):111113.
Heinemann IU, Jahn M, Jahn D. The biochemistry of heme biosynthesis. Arch Biochem Biophys. 2008;474(2):238–51.
Paterniti JR Jr, Lin CI, Beattie DS. delta-Aminolevulinic acid synthetase: regulation of activity in various tissues of the aging rat. Arch Biochem Biophys. 1978;191(2):792–7.
Bitar M, Weiner M. Modification of age-induced changes in heme and hemoproteins by testosterone in male rats. Mech Ageing Dev. 1983;23(3–4):285–96.
Nozawa N, et al. 5-Aminolevulinic acid and sodium ferrous citrate ameliorate muscle aging and extend healthspan in Drosophila. FEBS Open Bio. 2022;12(1):295–305.
Parkhitko AA, et al. Targeting metabolic pathways for extension of lifespan and healthspan across multiple species. Ageing Res Rev. 2020;64:101188.
Phua CZJ, et al. Genetic perturbation of mitochondrial function reveals functional role for specific mitonuclear genes, metabolites, and pathways that regulate lifespan. Geroscience. 2023;45(4):2161–78.
Lee SS, et al. A systematic RNAi screen identifies a critical role for mitochondria in C. elegans longevity. Nat Genet. 2003;33(1):40–8.
Copeland JM, et al. Extension of Drosophila life span by RNAi of the mitochondrial respiratory chain. Curr Biol. 2009;19(19):1591–8.
Liu X, et al. Evolutionary conservation of the clk-1-dependent mechanism of longevity: loss of mclk1 increases cellular fitness and lifespan in mice. Genes Dev. 2005;19(20):2424–34.
Hamilton B, et al. A systematic RNAi screen for longevity genes in C. elegans. Genes Dev. 2005;19(13):1544–55.
McCormick MA, et al. A comprehensive analysis of replicative lifespan in 4,698 single-gene deletion strains uncovers conserved mechanisms of aging. Cell Metab. 2015;22(5):895–906.
Acknowledgements
We would like to thank Dr. Iqbal Hamza for providing the pYES-DEST52-ceHRG-4 plasmid used in this study.
Funding
This work was supported by NIH Grants AG058713 and AG066704 to VML.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Patnaik, P.K., Nady, N., Barlit, H. et al. Lifespan regulation by targeting heme signaling in yeast. GeroScience 46, 5235–5245 (2024). https://doi.org/10.1007/s11357-024-01218-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11357-024-01218-9