Abstract
We investigated the prevalence of antibiotic resistant staphylococci and detection of resistant, virulence, and Spa genes in a South African wastewater treatment plant. Species identified were Staphylococcus aureus, S. lentus, S. arlettae, S. cohnii, S. haemolyticus, S. nepalensis, S. sciuri (now Mammaliicoccus sciuri), and S. xylosus. Isolates showed high resistance to methicillin (91%), ampicillin (89%), ciprofloxacin (86%), amoxycillin (80%), ceftazidime (74%), and cloxacillin (71%). Multiple antibiotic resistance (MAR) index for the isolates exceeded 0.2 (0.50–0.70). Among the isolates, 77% were mecA-positive. All S. aureus strains were positive for nuc and 7 Spa gene types. The present study highlights possibility of treated wastewaters being potential reservoir for antibiotic-resistant staphylococci. This is a cause for concern as wastewater effluents are decanted into environmental waters and these are, in many cases, used for various purposes including recreation (full contact), religious (full body submersion), and drinking water for some rural communities and water for livestock.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Due to increasing human population and urbanization as well as changing climatic conditions, the challenge of water scarcity has heightened in many developed and developing countries (du Plessis 2019a). South Africa is no exception, and extended droughts in the catchment areas of reservoir dams in the Western and Eastern Cape, KwaZulu Natal, and Northern Cape were responsible for major metropolitan cities to implement water restrictions (Botai et al. 2019; du Plessis 2019b). Alternative water sources, such as the reclamation of wastewater treatment plant (WWTP) effluent for various purposes, become crucial (Salgot and Folch 2018).
Treated wastewater effluents are usually discharged into receiving waters and reused for various purposes such as preparing drinking water, agricultural irrigation and livestock water, recreation, and industrial purposes (Angelakis et al. 2018). Effectively treated wastewaters are determined by their quality which are dependent on the physico-chemical properties (pH, temperature, electrical conductivity, various chemical constituents such as phosphates, nitrogen containing compounds, and organic load metals) and microbial indicators of fecal contamination, mostly E. coli (Jordaan & Bezuidenhout 2013). Indiscriminate discharge of poorly treated or untreated wastewater effluents are major contributors to surface water pollution (Malassa et al. 2013; Amirsoleimani et al. 2019; Kiliça et al. 2023).
According to Börjesson et al. (2010), Said et al. (2017) and Azuma et al. (2022) wastewaters are potential sources for the dissemination of antibiotic resistant bacteria (ARB) such as staphylococcal species into natural water environments. These staphylococcal species rank high among the bacteria causing diseases. In addition, they have been incriminated for many human infections such as skin and soft tissue infections, surgical site/wound infections, pneumonia, septicemia, and bone infections (Nanoukona et al. 2017; Oladipo et al. 2019). Several studies have detected staphylococcal species in wastewaters (Börjesson et al. 2009; Goldstein et al. 2012; Gómez et al. 2016). Porrero et al. (2016) reported the presence of Staphylococcus aureus in WWTP in Madrid, Spain, while Faria et al. (2009) and Čuvalova et al. (2015) reported the survival of coagulase negative staphylococci (CoNS) in treated effluents and drinking water from Portugal and Slovak Republic, respectively. Specifically, Gómez et al. (2016) and Said et al. (2017) detected 5 coagulase negative staphylococci (CoNS) — S. lentus, S. cohnii, S. sciuri, S. haemolyticus, and S. xylosus in wastewaters in Spain and Tunisia. Borjesson et al. (2009) also identified S. lentus, S. sciuri, S. cohnii, and S. haemolyticus in a municipal wastewater treatment plant in Sweden.
Staphylococcus aureus may be associated with severe infection, hence the need to distinguish it from the opportunistic coagulase negative staphylococci. In routine laboratory practice, the production of coagulase is frequently used as the sole criterion to distinguish S. aureus from other staphylococci. The coagulase test is therefore an important distinguishing characteristic of staphylococci (Cheesbrough 2006). Those that are coagulase positive are generally regarded as S. aureus and are potential pathogens that are flagged for further diagnostic tests (Cheesbrough 2006), while the CoNS are generally regarded as non-pathogenic and are routinely disregarded in the clinical diagnostic sphere (Okwara et al. 2004).
Staphylococcal species may exhibit resistance towards beta-lactam antibiotics such as ampicillin, methicillin, and penicillin (Porrero et al. 2016; Thompson et al. 2012; Said et al. 2017). The World Health Organization (WHO 2017a) reported that in Africa, 80% of Staphylococcus aureus infections are methicillin resistant. Multi-drug antibiotic resistance traits and antibiotic resistance genes (ARGs) such as in MRSA and other CoNS species had been isolated from wastewaters (Börjesson et al. 2009; Thompson et al. 2012; Wan and Chou 2014; Boopathy 2017; Said et al. 2017). In Nigeria, more recent studies by Adekanmbi et al. (2019), Oladipo et al. (2019), and Adesoji et al. (2020) have also confirmed the presence of multi-drug antibiotic-resistant staphylococci and mecA gene from wastewater sources.
The confirmation of the presence of the mecA gene has been the “golden standard” for detection of methicillin-resistant S. aureus (MRSA) worldwide (Yang et al. 2009; Igbinosa et al. 2016). The nuc gene detection is a confirmatory test for S. aureus strains, while pvl is generally used as a marker for community acquired MRSA (Gillen et al. 2015). The PVL gene is a virulence factor, which can enhance the ability of the bacterium to cause severe infections in human and animal hosts. Spa-typing of S. aureus strains is an investigation which could provide useful insight and information into the virulence potentials and nature of S. aureus specie. This test may further assist in the grouping of isolates into clonal lineages and S. aureus populations (Kolawole et al. 2013).
Occurrence of MRSA and genes in wastewater effluents discharged into water environments has therefore raised public health concerns due to likely threats posed to the human communities which could lead to community acquired MRSA (CA-MRSA) infections (Börjesson et al. 2009, 2010; Plano et al. 2011; Rosenberg et al. 2012). In South Africa, there is limited data on the detection of staphylococcal species in WWTPs and whether these make it into receiving water bodies as well as their persistence in these waters (Chidamba et al. 2016).
The aim of this study was to determine (i) the prevalence of staphylococcal species that are resistant to methicillin and other related antibiotics and (ii) the presence of mecA, nuc, and luk-pvl genes and spa types in the resistant staphylococci isolated from a South African wastewater treatment plant using standard protocols.
Materials and methods
Description and treatment processes at the wastewater treatment plant sampled
Water samples were collected from a wastewater treatment plant (WWTP) in the North-West Province of South Africa. From this plant, four sites were sampled; these were influent, primary effluent, secondary effluent, and final effluent. The plant is a full scale-wastewater treatment plant which has a designed capacity of 45,000 m3 per day with the potential of receiving wastewater streams from domestic, industrial, agricultural, abattoirs, hospital, and storm water sources. The average flow to the works is 29,000 m3 per day. The influent receives raw sewage into the plant for treatment. The treatment plan employed at the WWTP for each of the four sites is influent — here, preliminary filtration/mechanical methods are used. For the secondary effluent, biological treatment option is utilized, while the final or tertiary effluent is the final stage of treatment where chemical treatment by chlorination is employed. Generally, chlorination at 5 mg/L is used to reduce E. coli levels to 0 cfu/100 ml and to reduce odor caused by microorganisms before discharge into receiving waters.
Sampling description
Samples were collected from each of the four sites in sterile 500 mL Schott glass bottles each. Grab sampling technique was used, and triplicate samples were collected from each of the four sampling points weekly for a period of four months. The wastewater samples were then transported in ice chested coolers and preserved under refrigeration conditions for microbiological analyses. The latter were conducted within 12 h of collection.
Microbiological analysis of wastewater samples and preliminary identification of staphylococcal species
About 100 mL of water samples collected were filtered using sterile 0.45 µm membrane filters. These filters were afterwards enriched in Bacto tryptic soy broth (soybean-casein digest medium; Becton Dickinson, USA) and were later placed onto Mannitol salt agar (Biotec Laboratories, Kentford, UK). The resulting yellow colonies were presumed to be S. aureus. These were afterwards confirmed by culturing on MRSA CHROMagar base (CHROMagar™ MRSA-ITK Diagnostics BV, Uithoorn, The Netherlands). The putative MRSA produced characteristic purple color on the chromogenic agar. Gram staining was used to ensure that isolates were Gram-positive cocci and have characteristic clusters. Staphylococcus aureus and other staphylococcal species were later confirmed by the coagulase and catalase tests using standard protocols (Cheesbrough 2006; Igbinosa et al. 2016). All isolated and identified staphylococci were subjected to antibiotic susceptibility testing using 13 antibiotics and the standard Kirby-Bauer’s disk diffusion technique (CLSI 2014).
16S rRNA gene–based identification of staphylococcal isolates
The Nucleospin® tissue extraction kit (Macherey–Nagel, Düren, Germany) was used, according to the manufacturer’s manual, to isolate genomic DNA. Briefly, 2 mL overnight broth cultures were centrifuged at 8000 rpm for 5 min at room temperature to harvest the cells. The supernatant was discarded, and pelleted cells were then resuspended in 100 µL T1 buffer. Quality and integrity of extracted DNA products were verified by micro-spectrophotometry and gel electrophoresis as described by Oladipo et al. (2018a).
PCR amplification
The C1000TM thermal cycler (Bio-Rad, Hercules, CA, USA) was used to perform PCR reactions. 16S rRNA gene amplification was conducted using primer sets and PCR conditions detailed in Table 1. Each PCR reaction included positive and negative controls as described by Oladipo et al. (2018b).
Sequencing of 16S rRNA genes
Purified PCR products were sequenced using the Big Dye terminator V. 3.1 cycle sequencing kit (Applied Biosystems, Warrington, UK) on a SeqStudio genetic analyzer and related software (Life Technologies, Holdings Pte Ltd, Singapore). Generated sequence electropherograms were inspected and then manually edited as described by Oladipo et al. (2018a). Edited sequences were aligned and compared against other sequences on the Basic Local Alignment Search Tool (BLAST) program alignment tool of the GenBank on the National Center for Biotechnology Information (NCBI) database (https://www.ncbi.nlm.nih.gov/). Phylogenetic sequence dendogram was constructed with closely related sequences obtained from GenBank by the neighbor-joining tree method using the Tamura–Nei substitution model in MEGA (Fig. 1). The partial 16S rRNA sequences obtained from this study are available in the GenBank with assigned accession numbers: MF409347–MF409381.
Unrooted neighbor-joining tree of Staphylococcus spp. isolated from a wastewater treatment plant in South Africa. Sequences obtained in this study are indicated as shaded circles. Accession numbers are indicated in bold. Neighbor-joining tree was constructed in MEGA (v. 6) using the Tamura-Nei substitution model replications. Bootstrap values below 50 are not shown
PCR amplification of mecA, nuc and luk-pvl genes in staphylococcal species
To differentiate MRSA from other staphylococci the PCR amplification of the mecA gene (encoding for methicillin resistance), nuc gene and the luk-pvl gene that encode for virulence in staphylococcal species were conducted. Each of the PCR reaction contained 12.5 µL, 2 × PCR Master mix (Thermos Scientific Technologies, Waltham, MA, USA), 50 ng DNA template, 5 µM each of the primers (forward and reverse), and nuclease-free water added to a final volume of 25 µL. Detailed information on the primers and conditions used are presented in Table 1. To determine if the PCRs worked, electrophoresis of the amplicons were performed using a 1% w/v agarose gel and conditions described in Oladipo et al. (2018a). Previously known positive genes of mecA, nuc, and pvl and positive Staphylococcus aureus isolates were used as control strains.
DNA amplification and sequencing of the protein A (spa)
For amplification of the Staphylococcus repeat region, a PCR was performed in a total volume of 50 μl containing cleaned DNA, 200 μM deoxynucleoside triphosphates (dATP, dCTP, dGTP, and dTTP), 10 pmol of each primer, 5 μl of tenfold concentrated PCR Buffer II (Applied Biosystems), MgCl2 1.5 mM, and 1.25 U of AmpliTaq DNA polymerase (Applied Biosystems, Hitachi, Tokyo, Japan). Detailed information on the primers and conditions used are presented in Table 1.
Sequencing of the protein A gene (spa) was carried out using the Big Dye terminator V. 3.1 cycle sequencing kit (Applied Biosystems, Warrington, UK) on a 3130 Genetic analyzer (Applied Biosystems/Hitachi, Tokyo, Japan). The chromatograms obtained were analyzed with the Ridom Staph Type software version 1.4 (RidomGmbH, Sedanstr, Germany; http://spa.ridom.de/index.shtml). Spa types were deduced by the differences in number and sequence of spa repeats with the BURP algorithm (Ridom GmbH, Sedanstr, Germany) and the Ridom Spa Server database. Spa types with less than five or equal to 5 repeat units were excluded (Montanaro et al. 2016).
Antimicrobial susceptibility testing
All isolated and identified staphylococcal species were subjected to antibiotic susceptibility testing of 13 antibiotics using the standard Kirby-Bauer’s disk diffusion technique (CLSI 2014). The specific antibiotics selected are beta-lactam antibiotics, a class of antibiotic that contain a beta-lactam ring in their molecular structures, that usually acts by inhibiting the synthesis of bacterial cell walls. This includes ampicillin, cloxacillin, amoxicillin, and methicillin. Others were macrolide-erythromycin, azithromycin, aminoglycoside gentamycin, carbapenems (imipenem), and glycopeptides (vancomycin). Methicillin was used to determine the antibiotic sensitivity of Staphylococcus aureus to other penicillin facing β-lactam resistance.
Antimicrobial resistance data were analyzed using the WHONET 2017 software V 5.6 (WHO; http://www.whonet.org/software.html). The multiple antibiotic resistance (MAR) index for the dominant isolates (S. aureus, S. lentus, and other staphylococcal species) at the influent and effluent compartments of the WWTP was calculated and interpreted according to Krumperman (1983) using the formula:
*MAR index values > 0.2 indicate high risk source of contamination (Krumperman 1983).
Statistical analyses
Statistical difference of MAR index of the staphylococcal species was done using one-way analysis of variance (ANOVA) at 5% level of significance using IBM SPSS Statistics 25 (IBM Corporation, Armonk, NY, USA). Multiple sequence alignment was performed using MUSCLE (Edgar 2004) integrated into Molecular Evolutionary Genetics Analysis (MEGA) V. 7.0 (http://www.megasoftware.net/; Kumar et al. 2016).
Results
Prevalence and distribution of staphylococcal strains from the wastewater treatment plant
Thirty-five staphylococcal isolates belonging to eight (1 CoPS–S. aureus and 7 CoNS) species were identified. The most prevalent were Staphylococcus aureus (34.0%), S. lentus (29.0%), S. cohnii (11.0%), and S. sciuri (9.0%). Other isolates were S. haemolyticus and S. xylosus (6.0%) each and S. nepalensis and S. arlettae with 3% each. The phylogenic relatedness of the staphylococcus species alongside related GenBank sequences further confirmed these identities (Fig. 1). Twelve of the staphylococcal isolates {6 S. aureus, 2 S. lentus, 2 S.haemolyticus, 1 S. cohnii, and S. xylosus each} were from the influent, 8 of 35 (22.86%) comprising of S. aureus (1), S. lentus (4), S. cohnii (1), and S. scuiri (2) — {now reclassified as new genus, Mammaliicoccus sciuri} (Madhaiyan et al. 2020) from primary effluent. From the secondary effluent, 6 of 35 (17.14%) consisting of 3 S. aureus, 1 S. lentus, and 2 S. cohnii were identified, while 9 of 35 (25.71%) including S. aureus (2), S. lentus (3), S. scuiri (1), S. arlettae (1), S. xylosus (1), and S. nepalensis (1) were from the final effluent (Fig. 2). Furthermore, the distribution of the species across the four sampling points showed that S. aureus and S. lentus were isolated from all the sampling compartments. In addition, S. arlettae and S. nepalensis were exclusively isolated from the final effluent point while S. xylosus was isolated from the influent and the final effluent.
Antibiotic resistance and susceptibility patterns of Staphylococcal species
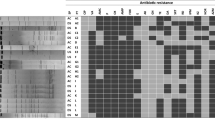
All thirty-five staphylococcal isolates from the wastewater treatment plant were subjected to 13 antibiotics at recommended concentrations for prove of resistance or susceptibility (Table 2). All the isolates were resistant to several of the 13 antibiotics tested. Resistance to the various antibiotics was in the following order: methicillin (91%), ampicillin (89%), ciprofloxacin (86%), amoxycillin (80%), ceftazidime (74%), and cloxacillin (71%). Other antibiotics are cefuroxime and azithromycin (43%), ofloxacin and vancomycin (40%), gentamycin (37%), imipenem (29%), and erythromycin (23%). About 70% (24 out of 35) of the isolates were resistant to at least 7 of the 13 antibiotics tested with S. aureus, S. lentus, and S. scuiri resistant to 10 of the 13 antibiotics (Table 2). It was observed that all the S. aureus strains isolated from the treatment plant regardless of their site of isolation were all resistant to methicillin and ampicillin (Table 2; Fig. 3). Staphylococcus aureus decreased from 50 to 17% as treatment progressed from influent to final effluent point in the WWTP. Furthermore, it was observed that 50% of S. aureus were resistant to imipenem and erythromycin.
Multiple antibiotic resistance (MAR) index and MAR phenotypes
The MAR index and phenotypes of the 2 dominant isolates (S. aureus and S. lentus) and other staphylococcal species comprising S. arlettae, S. haemolyticus, S. sciuiri, and S. cohnii are presented in Table 3. For S. aureus strains, the MAR index at the influent and effluent were 0.705 and 0.615, while for S. lentus, 0.577 and 0.692 were calculated. Furthermore, the MAR index of the other staphylococci — S. arlettae, S. haemolyticus, S. sciuiri, and S. cohnii (grouped as staphylococcus species) — were calculated as 0.519 for influents and 0.558 for effluents. However, there were no significant differences (p > 0.05) in the MAR index either within the sampling sites or among the different species. Also, the dominant phenotypes exhibited diverse patterns (Table 3). The MAR phenotype among S. aureus isolates from the influent showed diverse resistance to beta-lactam antibiotics and cephalosporins. The phenotypes — AMP-CLO-AMC-CAZ-FOX and AMP-CLO-AMC-CAZ-FOX-CFM — were dominant in S. aureus (16.7%), S. lentus (50.0%), and other staphylococci (33.3%) from the influent chamber of the WWTP. However, at the effluent chamber, the MAR phenotypes, AMP-CLO-AMC-CAZ-FOX-CFM-CIP-ERY-VAN (S. aureus), AMP-CLO-AMC-CAZ-FOX-CIP-VAN (S. lentus), and AMP-CLO-AMC-CAZ-FOX-CFM-AZM (Staph spp.), were observed in 50%, 25%, and 33.3% of the isolates, respectively.
Detection of resistance and virulence genes and in staphylococcal species
Seventy-seven percent of the isolates (27 of 35) were mecA positive. Among the 12 MRSA isolates, 11 were mecA positive. Other staphylococci that tested positive for mecA were S. lentus, S. scuiri (Mammaliicoccus sciuri), S. cohnii, S haemolyticus, and S. xylosus. This study revealed a higher number of isolates being recovered in the final effluent; however, 75% of these isolates did not carry the mecA resistance gene. Two (5.8%) S. aureus isolates were also positive for the luk-pvl gene, while the nuc gene was detected in all 12 S. aureus isolates (Table 4).
Detection of protein A (spa) types in Staphylococcus aureus strains
Seven different spa types were detected from the confirmed S. aureus strains recovered from the WWTP in this study (Table 4). Spa types t061, t6578, and t091 were detected at the influent, t447 from primary effluent, t7835 from secondary effluents, and spa types t091 and t5126 final effluent. The most frequent spa type t091 (16.7%) was observed from the influent and secondary effluent.
Discussion
Several researchers in South Africa have investigated links between WWTP effluents and receiving waters. These have focused on physico-chemical properties (Agoro et al. 2018; Salvador-Oke et al. 2018) or microbial parameters (microorganisms) such as Vibrio (Okoh & Igbinosa 2010) and Aeromonas species (Igbinosa & Okoh 2012; Coetzee et al. 2017; Mann et al. 2019). The present study was designed to assess the prevalence of staphylococcal species (S. aureus — CoPs and CoNS), their antibiotic resistance patterns, and detection of resistance and virulence genes and Spa types in the recovered staphylococcal species from a WWTP in South Africa.
In the present study, eight staphylococcal species (1 CoPS and 7 CoNS) were isolated and identified across the 4 sampling points of a wastewater treatment plant in South Africa. These are Staphylococcus aureus, S. arlettae, S. cohnii, S. haemolyticus, S. lentus, S. nepalensis, S. sciuri, and S. xylosus. The Staphylococcus spp. demonstrated multidrug resistance (MDR), high MAR index (> 0.2), various MAR phenotypes, detection of mecA resistance gene, and the nuc and luk-pvl virulence gene and Spa types confirmed in S. aureus strains.
This study confirmed the presence of staphylococci in the final effluent after chlorination. Due to its simple management, low cost, and high efficiency in eliminating microorganisms in wastewater treatment plants, chlorination purification method at the final/tertiary effluent phase was considered an effective disinfection method (Wang et al. 2020; Collivignarelli et al. 2021). However, in recent times, chlorination has proved to transmit antibiotic resistance genes (ARGs) in treated wastewaters (Ghernaout & Elboughdiri 2020; Collivignarelli et al. 2021). Hence, this process may not lethally affect microorganisms including staphylococci in wastewaters (Liu et al. 2018; Collivignarelli et al. 2021). This may explain the detection of antibiotic-resistant staphylococcal species after chlorination in the wastewater treatment plant in this study.
WHO (2017b) had earlier reported chlorine resistance of staphylococci species. Previous studies (Huang et al. 2011; Shi et al. 2013; Mao et al. 2015) reported the presence of antibiotic-resistant bacteria that revealed resistance to chlorination. In addition, Gómez et al. (2016) confirmed the detection of multi-drug resistant staphylococci (S. aureus, S. lentus, S. cohnii, S. scuiri (Mammaliicoccus sciuri), and S. haemolyticus) in urban wastewater treatment plant in Spain at the final effluent phase. Similarly, Goldstein et al. (2012) and Maimon et al. (2014) recorded the occurrence of S. aureus and methicillin-resistant S. aureus (MRSA) in treated wastewater effluents from greywater, intended for reuse. Hence, in order to eliminate the presence of microorganisms such as staphylococci species from treated wastewaters, the use of ultraviolet (UV) radiation has been suggested as a promising and more effective treatment technology (Collivignarelli et al. 2021).
All S. aureus isolates in the present study were resistant to ampicillin and methicillin. This appears to be a constant observation amongst previous studies. Thapaliya et al. (2017) also recorded the prevalence of S. aureus and their antibiotic-resistance in wastewater treatment plant sites. Their study investigated the prevalence and molecular characteristics of S. aureus and MRSA in freshwater recreational beaches sand and water samples collected from 10 beaches in Northeast Ohio, USA. Results from their study revealed overall prevalence of S. aureus (22.8%) and PVL genes (21.4%), with 27 different spa types identified. In addition, 34.3% of the isolates showed oxacillin resistance while, all the isolates showed 100% resistance to penicillin. However, our present study revealed a higher prevalence of S. aureus (34.3%) with a prevalence of 5.71% for PVL genes and 7 spa types being identified among S. aureus isolates.
Results of Thompson et al. (2012) and Porrero et al. (2016) showed that 96% and 83% MRSA isolates, respectively, from urban effluents were resistant to ampicillin. The presence of MRSA and MSSA in river water and urban effluents was studied to analyze the S. aureus population and determine the genetic diversity. From their study, MRSA population in urban effluents and river water was 67.6% and 82.4%, while spa type t067 was the predominant MRSA genotype detected. This differs from our study in that we only recorded an MRSA prevalence of 35%, in a WWTP with spa type t091 being dominant. Said et al. (2017) reported that most of the S. aureus in their study showed resistance to penicillin, while Goldstein et al. (2012) also demonstrated that 93% of MRSA isolates recovered from wastewaters in the USA were multidrug resistant. The study by Goldstein et al. (2012) examined the occurrence of MRSA and methicillin-susceptible S. aureus (MSSA) at US wastewater treatment plants. The study and findings were similar to this study since the presence of MRSA in a WWTP was investigated. Results from their study also showed 10 of 12 (83%) influent samples being MRSA-positive, while one of 12 (8%) effluent samples was MRSA-positive.
In the present study, it was shown that the mecA resistance gene was detected in 11 of the 12 S. aureus strains recovered from the WWTP sampled. This finding is corroborated by several previous studies (Wan & Chou 2014; Boopathy 2017). While Boopathy (2017) established the presence of methicillin-resistant Staphylococcus aureus (MRSA) in a rural sewage treatment plant, Wan and Chou (2014) examined the spreading of β-lactam resistance gene (mecA) and methicillin-resistant Staphylococcus aureus through municipal and swine slaughterhouse wastewaters.
The nuc gene was detected in all the S. aureus strains isolated from this study. However, the pvl virulence gene was detected in very few isolates. A similar study of clinical isolates (von Eiff et al. 2004) examined the prevalence of genes encoding for members of the staphylococcal leukotoxin family of Staphylococcus aureus. Their findings revealed 0.9 to 1.4% detection of pvl virulence gene.
The occurrence of CoNS in WWTPs has also been well documented. Faria et al. (2009) reported the survival of CoNS in treated effluents. On the other hand, Čuvalova et al. (2015) demonstrated that CoNS also occurred in drinking water. Of the 7 CoNS species identified in this study, Gómez et al. (2016) and Said et al. (2017) detected 5 species in their studies that focused on wastewater samples. These species included S. cohnii, S. haemolyticus, S. lentus, S. scuiri (Mammaliicoccus sciuri), and S. xylosus from wastewater samples. Borjesson et al. (2009) recovered S. cohnii, S. haemolyticus S. lentus, and S. sciuri from a municipal wastewater treatment plant. Antibiotics resistance by CoNS had also been documented. This was the case in the present study and several previous studies. Said et al. (2017) reported that CoNS isolated from wastewaters in Tunisia were resistant to several classes of antibiotics including beta-lactam antibiotics. Previously, Schwartz et al. (2003) had reported the occurrence of methicillin-resistant CoNS from wastewater environments. The detection of mecA resistance gene in CoNS has also been reported in recreational waters, community, and hospital wastewaters (Börjesson et al. 2009; Fogarty et al. 2015) and in other surface waters (Seyedmonir et al. 2015). Finding CoNS strains in the wastewater from the present North West Province of South Africa is thus not extraordinary.
The multiple antibiotic resistance (MAR) index of all 35 (100%) Staphylococcus spp. in our study exceeded the 0.2 value associated with highly antibiotic resistant strains. In a previous study (Oladipo et al. 2019), very high MAR index (> 0.2) were also recorded for S. aureus isolates from clinical and environmental sources. Multiple antibiotic resistance in bacteria is most commonly associated with the presence of plasmids which contain one or more resistance genes, each encoding a single antibiotic resistance phenotype. MAR index values greater than 0.2 indicate high risk source of contamination where antibiotics are often used. Our findings in the present study indicate that the presence of staphylococcal species exhibiting antibiotic resistance and harboring environmentally relevant virulence genes are of particular concern due to the possible link of community acquired MRSA and wastewater recycling for domestic, agricultural, and industrial use. The frequencies of resistance of S. aureus to beta-lactams antibiotics (AMP-CLO-AMC-CAZ-FOX) were high at all our sampling sites (influent, primary, and secondary effluents and final effluent). Several studies have shown S. aureus resistance to antibiotics such as penicillin, amoxicillin, and/or ampicillin have been isolated from both treated and untreated wastewater (Sahlstrom et al. 2004; Feng 2008). In a previous study carried out in the USA, increase percentages of Ery-, Amp-, and Pen- -resistant were also reported among staphylococcal species isolated from a WWTP (Goldstein et al. 2012).
Seven distinct spa types were identified in this study with t091 being the most prevalent. Finding such a variety of spa types is a potential indication of diverse sources of isolation and that these could be from different geographical locations. This was also observed in a study in Nigeria (O'Malley et al. 2015; Ayeni et al. 2018) where spa type t091 was confirmed in nasal samples of clinical and poultry sources. In addition, Ilczyszyn et al. (2016) reported occurrence of spa type t091 amongst 5-year-old and younger patients in Poland. However, no data in searched databases could be found for a South African study on MRSA that had documented spa type t091 being associated with wastewaters.
The spa type t7835 had been associated with MRSA from clinical isolates in Nigeria (Kolawole et al. 2013), while spa type t447 had been reported in Netherlands and Spain. Also, spa type t6578 had been identified among swine (LA-MRSA) in Spain and the USA as ST398 (CC398), and subsequently detected in several companion and food-chain animals and humans (de Boer 2009). According to Smith et al. (2009), ST398 (CC398) has been well reported as a cause of livestock-associated (LA)-MRSA in Europe, while in Australia (Price et al. 2012) and the Americas (Grema et al. 2015), ST398 had been confirmed as a cause of LA-MRSA. Spa type t5126 had been identified in MRSA strains in Spain, USA, Germany, and France (https://spa.ridom.de/spa-t5126.shtml ), while spa type t061 had been associated with MRSA in the UK, Germany, and USA (von Eiff et al. 2008). Notably, spa type t657, sequence type (ST)772, was reported in this study among the pvl positive strains. This sequence type had been linked to community outbreak of CA-MRSA infections in some parts of the world, e.g., India (D'Souza et al. 2010) and Ireland (Edmundson et al. 2011).
In this study, S. aureus constituted 34% of the total recovered staphylococcal species which decreased as treatment progressed from influent to final effluent point. Similarly, studies conducted in Sweden, Spain, and the USA, respectively, reported 50–55% prevalence of S. aureus in WWTPs with decreased prevalence as treatment progressed (Börjesson et al. 2009; Goldstein et al. 2012; Gómez et al. 2016). In this study, the higher number of isolates from the final effluent which did not carry the mecA resistance gene could potentially be similar to the findings of Mao et al. (2015) that also reported a reduction of antibiotics resistance genes (ARGs) and mecA gene from raw influent point to the effluent.
In this study, three CoNS-S. nepalensis, S. arlettae, and S. cohnii from the WWTP which have not been widely reported were detected. Of these, S. cohnii carried the mecA resistance gene. Nováková et al. (2006) had earlier isolated S. nepalensis from human urine, S. arlettae from textile effluents (Elisangela et al. 2008), while S. cohnii had been recovered from wastewater samples (Börjesson et al. 2009; Gómez et al. 2016; Said et al. 2017). Staphylococcus cohnii is known for its associations with nosocomial infections (Chen et al. 2015) and had also been confirmed in infections in animals (Sousa et al. 2014).
Previously, CoNS had been regarded as non-pathogenic since most of these species are established by association between humans and animals (Otto 2010). However, their antibiotic resistance traits, possession of resistance, and virulence genes reveal evidence that these species could have detrimental human health outcomes and should therefore be studied more closely.
Conclusions
The present study demonstrates that environmental waters that receive WWTP effluent could be contaminated with MRSA and other potential pathogenic staphylococci. These findings indicate the possibility of treated wastewaters being a source for the dissemination of staphylococcal species, their resistance, and virulence genes to the environment which could have detrimental health impacts on the downstream users and consumers. The detection of a large proportion of MAR isolates in the present study is a cause for concern as this could pose health risks to humans and animals via resistant genetic elements that could be transferred from these isolates to other bacteria also of clinical importance. Therefore, a more effective treatment plan or treatment modification procedures of wastewaters such as ultraviolet (UV) radiation may therefore be crucial, especially if the water is to be reused. The findings of the present study are aspects that the managers of wastewater treatment plants, policy formulators, down-stream users, etc., should consider.
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Adekanmbi AO, Soyoye OF, Adelowo OO (2019) Characterization of methicillin-resistance gene mecA in coagulase negative staphylococci (CoNS) recovered from wastewater of two healthcare facilities in Nigeria. Gene Reports 17:100541. https://doi.org/10.1016/j.genrep.2019.100541
Adesoji TO, Egyir B, Shittu AO (2020) Antibiotic-resistant staphylococci from the wastewater treatment plant and grey-water samples in Obafemi Awolowo University, Ile-Ife, Nigeria. J Water Health 18(6):890–898. https://doi.org/10.2166/wh.2020.019
Agoro MA, Okoh OO, Adefisoye MA, Okoh AI (2018) Physicochemical properties of wastewater in three typical South African sewage works. Pol J Environ Stud 27(2):491–499. https://doi.org/10.15244/pjoes/74156
Amirsoleimani A, Brion M, Diene SM, François P, Richard EM (2019) Prevalence and characterization of Staphylococcus aureus in wastewater treatment plants by whole genomic sequencing. Water Res 158:193–202. https://doi.org/10.1016/j.watres.2019.04.035
Angelakis AN, Asano T, Bahri A, Jimenez BE, Tchobanoglous G (2018) Water reuse: from ancient to modern times and the future. Front Environ Sci 6:26. https://doi.org/10.3389/fenvs.2018.00026
Ayeni FA, Ruppitsch W, Allerberger F (2018) Molecular characterization of clonal lineage and staphylococcal toxin genes from S. aureus in Southern Nigeria. Peer J 9(6):e5204. https://doi.org/10.7717/peerj.5204
Azuma T, Usui M, Hayashi T (2022) Inactivation of antibiotic-resistant bacteria in wastewater by ozone-based advanced water treatment processes. Antibiotics 11:210. https://doi.org/10.3390/antibiotics11020210
Boopathy R (2017) Presence of methicillin resistant Staphylococcus aureus (MRSA) in sewage treatment plant. Bioresour Tech 240:144–148. https://doi.org/10.1016/j.biortech.2017.02.093
Börjesson S, Matussek A, Melin S, Löfgren S, Lindgren PE (2010) Methicillin-resistant Staphylococcus aureus (MRSA) in municipal wastewater: an uncharted threat? J Appl Microbiol 108(4):1244–1251. https://doi.org/10.1111/j.1365-2672.2009.04515.x
Börjesson S, Melin S, Matussek A, Lindgren PE (2009) A seasonal study of the mecA gene and Staphylococcus aureus including methicillin-resistant S. aureus in a municipal wastewater treatment plant. Water Res 43(4):925–932. https://doi.org/10.1016/j.watres.2008.11.036
Botai JO, Botai CM, de Wit JP, Muthoni M, Adeola AM (2019) Analysis of drought progression physiognomies in South Africa. Water 11:299. https://doi.org/10.3390/w11020299
Cheesbrough M (2006) District laboratory practice in tropical countries. Part 2, 2nd edn. Cambridge University Press, pp 64–66
Chen HJ, Hung WC, Lin YT, Tsai JC, Chiu HC, Hsueh PR, Teng LJ (2015) A novel fusidic acid resistance determinant, fusF, in Staphylococcus cohnii. J Antimicrob Chemother 70:416–419. https://doi.org/10.1093/jac/dku408
Chidamba LC, Cilliers E, Bezuidenhout CC (2016) Spatial and temporal variations in pollution indicator bacteria in the Lower Vaal River South Africa. Water Environ Res 88(11):2142–2149. https://doi.org/10.2175/106143016X14733681695528
Clinical and Laboratory Standards Institute (CLSI) (2014) Performance standards for antimicrobial susceptibility testing; Twenty-fourth informational supplement. CLSI Document M100-S24, Wayne, 34(1)
Coetzee I, Bezuidenhout CC, Bezuidenhout JJ (2017) Triclosan resistant bacteria in sewage effluent and cross-resistance to antibiotics. Water Sci Tech 76(6):1500–1509. https://doi.org/10.2166/wst.2017.335
Collivignarelli MC, Abbà A, Miino MC, Caccamo FM, Torretta V, Rada EC, Sorlini S (2021) Disinfection of wastewater by UV-based treatment for reuse in a circular economy perspective. Where are we at? Int J Environ Res Public Health 18(1):77. https://doi.org/10.3390/ijerph18010077
Čuvalova Z, Pipova M, Kantikova M, Brtkova A, Fajber J (2015) Virulence factors and antimicrobial resistance of coagulase-negative staphylococci isolated from drinking water. Open Life Sci 10:328–338. https://doi.org/10.1515/biol-2015-0034
de Boer E, Zwartkruis-Nahuis JT, Wit B, Huijsdens XW, de Neeling AJ, Bosch T et al (2009) Prevalence of methicillin-resistant Staphylococcus aureus in meat. Int J Food Microbiol 134:52–56. https://doi.org/10.1016/j.ijfoodmicro.2008.12.007
D’Souza N, Rodrigues C, Mehta A (2010) Molecular characterization of methicillin-resistant Staphylococcus aureus with emergence of epidemic clones of sequence type (ST) 22 and ST 772 in Mumbai, India. J Clin Microbiol 48:1806–1811. https://doi.org/10.1128/jcm.01867-09
du Plessis A (2019a) Current and future water scarcity and stress. In: Water as an inescapable risk. Springer Water. Springer, Cham. https://doi.org/10.1007/978-3-030-03186-2_2
du Plessis A (2019b) Evaluation of Southern and South Africa’s freshwater resources. In: water as an inescapable risk. Springer Water. Springer, Cham. https://doi.org/10.1007/978-3-030-031862_7
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
Edmundson SP, Hirpara KM, Bennett D (2011) The effectiveness of methicillin-resistant Staphylococcus aureus colonisation screening in asymptomatic healthcare workers in an Irish orthopaedic unit. Eur J Clin Microbiol Infect Dis 30:1063–1066. https://doi.org/10.1007/s10096-011-1192-3
Elisangela F, Andrea Z, Fabio D, de Guimaro MC, Ragagnin R, Durrant L, Artur C-P (2008) Biodegradation of textile azo dyes by a facultative Staphylococcus arlettae strain VN-11 using a sequential microaerophilic/aerobic process. Int Biodeter Biodegrad 63(3):280–288. https://doi.org/10.1016/j.ibiod.2008.10.003
Faria C, Vaz-Moreira I, Serapicos E, Nunes OC, Manaia CM (2009) Antibiotic resistance in coagulase negative staphylococci isolated from wastewater and drinking water. Sci Total Environ 407:3876–3882. https://doi.org/10.1016/j.scitotenv.2009.02.034
Feng Y, Chen CJ, Su LH, Hu S, Yu J, Chiu CH (2008) Evolution and pathogenesis of Staphylococcus aureus: lessons learned from genotyping and comparative genomics. FEMS Microbiol Rev. 32(1):23–37. https://doi.org/10.1111/j.1574-6976.2007.00086.x
Fogarty LR, Haack SK, Johnson HE, Brennan AK, Isaacs NM, Spencer C (2015) Staphylococcus aureus and methicillin-resistant S. aureus (MRSA) at ambient freshwater beaches. J Water Health 13(3):680–692. https://doi.org/10.2166/wh.2014.278
Ghernaout D, Elboughdiri N (2020) Is not it time to stop using chlorine for treating water? Oalib 7:1–11. https://doi.org/10.4236/oalib.1106007
Gillen AL, Conrad J, Cargill M (2015) The Genesis and emergence of community-associated methicillin-resistant Staphylococcus aureus (CA-MRSA): An Example of evolution in action? Ans Res J 8:391–401. https://digitalcommons.liberty.edu/bio_chem_fac_pubs
Goldstein RE, Micallef SA, Gibbs SG, Davis JA, He X, George A, Kleinfelter LM, Schreiber NA, Mukherjee S, Sapkota A, Joseph SW, Sapkota AR (2012) Methicillin-resistant Staphylococcus aureus (MRSA) detected at four U.S. wastewater treatment plants. Environ Health Perspect 120:1551–1558. https://doi.org/10.1289/ehp.1205436
Gómez P, Lozano C, Benito D, Estepa V, Tenorio C, Zarazaga M, Torres C (2016) Characterization of Staphylococci in urban wastewater treatment plants in Spain, with detection of methicillin-resistant Staphylococcus aureus ST398. Environ Pollut 212:71–76. https://doi.org/10.1016/j.envpol.2016.01.038
Grema HA, Geidam YA, Gadzama GB, Ameh JA, Suleiman A (2015) Methicillin resistant Staphyloccus aureus (MRSA): a review. Adv Anim Vet Sci 3(2):79–98. https://doi.org/10.14737/journal.aavs/2015/3.2.79.98
Huang JJ, Hu HY, Tang F, Li Y, Lu SQ, Lu Y (2011) Inactivation and reactivation of antibiotic-resistant bacteria by chlorination in secondary effluents of a municipal wastewater treatment plant. Water Res 45(9):2775–2781. https://doi.org/10.1016/j.watres.2011.02.026
Igbinosa IH, Okoh AI (2012) Antibiotic susceptibility profile of Aeromonas species isolated from wastewater treatment plant. Sci World J 764563:6. https://doi.org/10.1100/2012/764563
Igbinosa EO, Beshiru A, Akporehe LU, Ogofure AG (2016) Detection of methicillin-resistant staphylococci isolated from food producing animals: A public health implication. Vet Sci 3:1–11. https://doi.org/10.3390/vetsci3030014
Ilczyszyn WM, Sabat AJ, Akkerboom V, Szkarlat A, Klepacka J, Sowa-Sierant I, Wasik B, Kosecka-Strojek M, Buda A, Miedzobrodzki J, Friedrich AW (2016) Clonal Structure and characterization of Staphylococcus aureus strains from invasive infections in paediatric patients from South Poland: association between age, spa types, clonal complexes, and genetic markers. PLoS One. 11(3):e0151937. https://doi.org/10.1371/journal.pone.0151937. Erratum in: PLoS One. 2016;11(4):e0154485A
Jordaan K, Bezuidenhout CC (2013) The impact of physico-chemical water quality parameters on bacterial diversity in the Vaal River, South Africa. Water SA 39(3):385–396. https://doi.org/10.4314/wsa.v39i3.7
Kiliça S, Firatb M, Yilmaz S, Ateş A (2023) A novel assessment framework for evaluation of the current implementation level of water and wastewater management practices. Water Supply 23(5):1787. https://doi.org/10.2166/ws.2023.117
Kolawole DO, Adeyanju A, Schaumburg F, Akinyoola AL, Lawal OO, Amusa YB, Köck R, Becker K (2013) Characterization of colonizing Staphylococcus aureus isolated from surgical wards’ patients in a Nigerian university hospital PLoS ONE 8:e68721. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0068721
Krumperman PH (1983) Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl Environ Microbiol. 46(1):165–70. https://doi.org/10.1128/aem.46.1.165-170.1983
Kumar RGL, Kasim KA, Lekshmi M, Nayak BB, Kumar S (2016) Incidence of methicillin-resistant staphylococci in fresh seafood. Adv Microbiol 6:399–406. https://doi.org/10.4236/aim.2016.66039
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E., Goodfellow M. (eds) Nucleic acids Techniques in Bacterial Systematics. pp. 115–147 https://doi.org/10.12691/jaem-2-4-11
Liu SS, Qu HM, Yang D, Hu H, Liu WL, Qiu ZG, Hou AM, Guo J, Li JW, Shen ZQ, Jin N (2018) Chlorine disinfection increases both intracellular and extracellular antibiotic resistance genes in a full-scale wastewater treatment plant. Water Res 136:131–136. https://doi.org/10.1016/j.watres.2018.02.036
Mann BC, Bezuidenhout JJ, Bezuidenhout CC (2019) Biocide resistant and antibiotic cross resistant potential pathogens from sewage and river water. Water Sci Tech 80(3):551–562. https://doi.org/10.2166/wst.2019.300
McClure J, Conly JM, Lau G (2006) Novel multiplex PCR assay for detection of the staphylococcal virulence marker Panton-Valentine leukocidin genes and simultaneous discrimination of methicillin-susceptible from resistant staphylococci. J Clin Microbiol 44:1141–1144. https://doi.org/10.1128/jcm.44.3.1141-1144.2006
Malassa H, Al-Qutob M, Al-Khatib M, Al-Rimawi F (2013) Determination of different trace heavy metals in ground water of South West Bank/Palestine by ICP/MS. J Environ Protect 4(08):818. https://doi.org/10.4236/jep.2013.48096
Madhaiyan M, Wirth JS, Saravanan VS (2020) Phylogenomic analyses of the Staphylococcaceae family suggest the reclassification of five species within the genus Staphylococcus as heterotypic synonyms, the promotion of five subspecies to novel species, the taxonomic reassignment of five Staphylococcus species to Mammaliicoccus gen. nov., and the formal assignment of Nosocomiicoccus to the family Staphylococcaceae. Int J Syst Evol Microbiol 70(11):5926–5936. https://doi.org/10.1099/ijsem.0.004498
Maimon A, Friedler E, Gross A (2014) Parameters affecting greywater quality and its safety for reuse. Sci Total Environ 487:20–25. https://doi.org/10.1016/j.scitotenv.2014.03.133
Mao D, Shuai Y, Rysz M, Luo Y, Yang F, Li F, Hou J, Mu Q, Alvarez PJJ (2015) Prevalence and proliferation of antibiotic resistance genes in two municipal wastewater treatment plants. Water Res 85:458–466. https://doi.org/10.1016/j.watres.2015.09.010
Montanaro L, Ravaioli S, Ruppitsch W, Campoccia D, Pietrocola G, Visai L, Speziale P, Allerberger F, Arciola CR (2016) Molecular Characterization of a prevalent ribocluster of methicillin-sensitive Staphylococcus aureus from orthopedic implant infections. Correspondence with MLST CC30. Front Cell Infect Microbiol 16(6):8. https://doi.org/10.3389/fcimb.2016.00008
Nanoukona C, Argemi X, Sogbo F, Orekan J, Keller D, Affolabi D, Schramm F, Riegel P, Baba-Moussa L, Prévost G (2017) Pathogenic features of clinically significant coagulase-negative staphylococci in hospital and community infections in Benin. Int J Med Microbiol 307:75–82. https://doi.org/10.1016/j.ijmm.2016.11.001
Nováková D, Pantůcek R, Petrás P, Koukalová D, Sedlácek I (2006) Occurrence of Staphylococcus nepalensis strains in different sources including human clinical material. FEMS Microbiol Lett 263(2):163–168. https://doi.org/10.1111/j.1574-6968.2006.00408.x
O’Malley SM, Emele FE, Nwaokorie FO, Idika N, Umeizudike AK, Emeka-Nwabunnia I, Hanson BM, Nair R, Wardyn SE, Smith TC (2015) Molecular typing of antibiotic-resistant Staphylococcus aureus in Nigeria. J Infect Public Health 8(2):187–193. https://doi.org/10.1016/j.jiph.2014.08.001
Okwara FN, Obimbo EM, Wafula EM, Murila FV (2004) Bacteraemia, urinary tract infection and malaria in hospitalised febrile children in Nairobi: is there an association? East Afr Med J 81(1):47–51. https://doi.org/10.4314/eamj.v81i1.8795
Oladipo AO, Oladipo OG, Bezuidenhout CC (2019) Multi-drug resistance traits of methicillin-resistant Staphylococcus aureus and other Staphylococcal species from clinical and environmental sources. J Water Health 17(6):930–943. https://doi.org/10.2166/wh.2019.177
Oladipo OG, Ezeokoli OT, Maboeta MS, Bezuidenhout JJ, Tiedt LR, Jordaan A, Bezuidenhout CC (2018a) Tolerance and growth kinetics of bacteria isolated from gold and gemstone mining sites in response to heavy metal concentrations. J Environ Manage 212C:357–366. https://doi.org/10.1016/j.jenvman.2018.01.038
Oladipo OG, Awotoye OO, Olayinka A, Bezuidenhout CC, Maboeta MS (2018b) Heavy metal tolerance traits of filamentous fungi isolated from gold and gemstone mining sites. Brazil J Microbiol 49:29–37. https://doi.org/10.1016/j.bjm.2017.06.003
Okoh AI, Igbinosa EO (2010) Antibiotic susceptibility profiles of some Vibrio strains isolated from wastewater final effluents in a rural community of the Eastern Cape Province of South Africa. BMC Microbiol 10:143. https://doi.org/10.1186/1471-2180-10-143
Otto M (2010) Staphylococcus colonization of the skin and antimicrobial peptides. Expert Rev Dermatol 5:183–195. https://doi.org/10.1586/edm.10.6
Othman R, Bakar MZA, Kasim Zamri-Saad M (2014) Improving the reproductive performance of buffaloes in Sabah. Malaysia. J Anim Health Prod 2(1):1–4
Plano LRW, Garza AC, Shibata T, Elmir SM, Kish J, Sinigalliano CD, Gidley ML, Gary M, Withum K, Fleming LE, Solo-Gabriele HM (2011) Shedding of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus from adult and pediatric bathers in marine waters. BMC Microbiol 11(1)5. https://doi.org/10.1186/1471-2180-11-5
Porrero MC, Valverde A, Mateos A, Cantón R, Gortázar C, Fernández Garayzábal JF, Domínguez L (2016) Staphylococcus aureus genetic lineages found in urban effluents and river water. Int J Water Wastewater Treat 2(2). https://doi.org/10.16966/2381-5299.117
Price LB, Stegger M, Hasman H (2012) Staphylococcus aureus CC398: host adaptation and emergence of methicillin resistance in livestock. Evidence of CC398 human to animal host jump. MBio 3:305–311. https://doi.org/10.1128/mBio.00305-11
Rosenberg Goldstein RE, Micallef SA, Gibbs SG, Davis JA, He X, George A, Kleinfelter LM, Schreiber NA, Mukherjee S, Sapkota A, Joseph SW, Sapkota AR (2012) Methicillin-resistant Staphylococcus aureus (MRSA) detected at four U.S. wastewater treatment plants. Environ Health Perspect 120(11):1551–1558. https://doi.org/10.1289/ehp.1205436
Sahlstrom L, Aspan A, Bagge E, Danielsson-Tham L, Albihn A (2004) Bacterial pathogen incidences in sludge from Swedish sewage treatment plants. Water Res 38:1989–1994. https://doi.org/10.1016/j.watres.2004.01.031
Said MB, Abbassi MS, Gómez P, Ruiz-Ripa L, Sghaier S, Ibrahim C, Torres C, Hassen A (2017) Staphylococcus aureus isolated from wastewater treatment plants in Tunisia: occurrence of human and animal associated lineages. J Water Health 15(4):638–643. https://doi.org/10.2166/wh.2017.258
Salgot M, Folch M (2018) Wastewater treatment and water reuse. Curr Opin in Environ Sci Health 2:64–74. https://doi.org/10.1016/j.coesh.2018.03.005
Salvador-Oke KT, King P, Olowoyo JO (2018) Bacteriological examination and physico chemical properties of streams receiving industrial effluents in Rosslyn, Pretoria, South Africa. J Environ Sci Manage 21(2):7–15. https://doi.org/10.47125/jesam/2018_2/02
Schwartz T, Kohnen W, Jansen B, Obst U (2003) Detection of antibiotic-resistant bacteria and their resistance genes in wastewater, surface water, and drinking water biofilms. FEMS Microbiol Ecol. 43(3):325–335. https://doi.org/10.1111/j.1574-6941.2003.tb01073.x
Seyedmonir E, Yilmaz F, Icgen B (2015) mecA gene dissemination among staphylococcal and non-staphylococcal isolates shed in surface waters. Bull Environ Contam Toxicol 95:131–138. https://doi.org/10.1007/s00128-015-1510z
Shi P, Jia SY, Zhang XX, Zhang T, Cheng SP, Li AM (2013) Metagenomic insights into chlorination effects on microbial antibiotic resistance in drinking water. Water Res 47(1):111–120. https://doi.org/10.1016/j.watres.2012.09.046
Shopsin B, Gomez M, Montgomery SO, Smith DH, Waddington M, Dodge DE, Bost DA, Riehman M, Naidich S, Kreiswirth BN (1999) Evaluation of protein A gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J Clin Microbiol 37(11):3556–3563. https://doi.org/10.1128/JCM.37.11.3556-3563.1999
Smith TC, Male MJ, Harper AL, Kroeger JS, Tinkler GP, Moritz ED, Capuano AW, Herwaldt LA, Diekema DJ (2009) Methicillin-resistant Staphylococcus aureus (MRSA) strain ST398 is present in midwestern U.S. swine and swine workers. PLoS One 4(1):e4258. https://doi.org/10.1371/journal.pone.0004258
Sousa M, Silva N, Igrejas G, Silvac F, Sargo R, Alegria N, Benito D, Gomez P, Lozano C, Gomez-Sanz E, Torres C, Caniça M, Poeta P (2014) Antimicrobial resistance determinants in Staphylococcus spp. recovered from birds of prey in Portugal. Vet Microbiol 171:436–440. https://doi.org/10.1016/j.vetmic.2014.02.034
Thapaliya D, Hellwig EJ, Kadariya J, Grenier D, Jefferson AJ, Dalman M, Kennedy K, DiPerna M, Orihill A, Taha M, Smith TC (2017) Prevalence and characterization of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus on public recreational beaches in Northeast Ohio. Geo Health 1:320–332. https://doi.org/10.1002/2017GH000106
Thompson JM, Gundogdu A, Stratton HM, Katouli M (2012) Antibiotic resistant Staphylococcus aureus in hospital wastewaters and sewage treatment plants with special reference to methicillin-resistant Staphylococcus aureus (MRSA). J Appl Microbiol 114:44–54. https://doi.org/10.1111/jam.12037
von Eiff C, Friedrich AW, Peters G, Becker K (2004) Prevalence of genes encoding for members of the staphylococcal leukotoxin family among clinical isolates of Staphylococcus aureus. Diagn Microbiol Infect Dis 49:157–162. https://doi.org/10.1016/j.diagmicrobio.2004.03.009
von Eiff C, Maas D, Sander G, Friedrich AW, Peters G, Becker K (2008) Microbiological evaluation of a new growth-based approach for rapid detection of methicillin-resistant Staphylococcus aureus. J Antimicro Chemo 61(6):1277–1280. https://doi.org/10.1093/jac/dkn122
Wan MT, Chou CC (2014) Spreading of b-lactam resistance gene (mecA) and methicillin-resistant Staphylococcus aureus through municipal and swine slaughterhouse wastewaters. Water Res 64:288–295. https://doi.org/10.1016/j.watres.2014.07.014
Wang A, Lin C, Shen Z, Liu Z, Xu H, Cheng J, Wen X (2020) Effects of pre-oxidation on haloacetonitrile and trichloronitromethane formation during subsequent chlorination of nitrogenous organic compounds. Int J Environ Res Public Health 17:1046. https://doi.org/10.3390/ijerph17031046
WHO (2017a) Antimicrobial resistance. http://www.who.int/mediacentre/news/releases/2017/bacteria-antibioticsneeded/en/. Accessed 3 Sept. 2019
WHO (2017b) World health organization. guidelines for drinking‑water quality. Fourth edition incorporating the first addendum. WHO library cataloguing-in-publication data, pp 631. https://www.who.int/publications/i/item/9789241549950. Accessed 06 Jun 2023
Yang CM, Lin MF, Liao PC, Yeh HW, Chang BV, Tang TK, Cheng C, Sung CH, Liou ML (2009) Comparison of antimicrobial resistance patterns between clinical and wastewater strains in a regional hospital in Taiwan. Lett Appl Microbiol 48:560–565. https://doi.org/10.1111/j.1472-765X.2009.02572.x
Acknowledgements
The authors sincerely appreciate the following: Messrs. Abram Mahlatsi, Michael Adebowale, Israel Ojo, and Mrs. Lee Chenhaka for their assistance during the study. The support of operators at the local WWTP in sampling is also acknowledged.
Funding
Open access funding provided by North-West University. The authors received financial support from the North-West University, South Africa, the research support in part by the National Research Foundation (NRF) of South Africa for the Grant No. 113824 and the Water Research Commission (WRC), South Africa (project K5/2347/3).
Author information
Authors and Affiliations
Contributions
Conceptualization: Cornelius Bezuidenhout. Data curation: Adegboyega Oladipo. Formal analysis: Adegboyega Oladipo, Oluwatosin Oladipo, Cornelius Bezuidenhout. Supervision: Cornelius Bezuidenhout. Validation: Cornelius Bezuidenhout, Oluwatosin Oladipo. Writing, review and editing: Oluwatosin Oladipo, Adegboyega Oladipo, Cornelius Bezuidenhout.
Corresponding author
Ethics declarations
Ethical statement
This study was approved by the Health Research Ethics Committee (HREC) of the Faculty of Health Sciences, North-West University, South Africa, under ethics number NWU-00122–17-S1.
Consent to participate
All the authors are in mutual agreement with the content of the manuscript and submission to the Environmental Science and Pollution Research Journal.
Consent for publication
All the authors support that the manuscript be published in Environmental Science and Pollution Research Journal.
Competing interests
The authors declare no competing interests.
Disclaimer
The views expressed are those of the authors and not of the funders.
Additional information
Responsible Editor: Diane Purchase
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oladipo, A.O., Oladipo, O.G. & Bezuidenhout, C.C. Detection of mecA positive staphylococcal species in a wastewater treatment plant in South Africa. Environ Sci Pollut Res 30, 117165–117178 (2023). https://doi.org/10.1007/s11356-023-30319-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-30319-9