Abstract
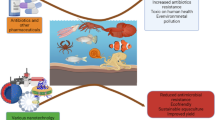
Biological treatment of swine liquid manure may be a favorable environment for the enrichment of bacteria carrying antibiotic resistance genes (ARGs), raising the alert about this public health problem. The present work sought to investigate the performance of a swine wastewater treatment plant (SWWTP), composed of a covered lagoon biodigester (CLB) followed by three facultative ponds, in the removal of usual pollutants, antibiotics, ARGs (blaTEM, ermB, qnrB, sul1, and tetA), and intI1. The SWWTP promoted a 70% of organic matter removal, mainly by the digester unit. The facultative ponds stood out in the solids’ retention carried from the anaerobic stage and contributed to ammonia volatilization. The detected antibiotic in the raw wastewater was norfloxacin (< 0.79 to 60.55 μg L−1), and the SWWTP seems to equalize peaks of norfloxacin variation probably due to sludge adsorption. CLB reduced the absolute abundance of ARGs by up to 2.5 log, while the facultative stage does not seem to improve the quality of the final effluent in terms of resistance elements. Considering the relative abundances, the reduction rates of total and ARG-carrying bacteria appear to be similar. Finally, correlation tests also revealed that organic matter and solids control in liquid manure treatment systems could help reduce the spread of ARGs after the waste final disposal.





Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the article, including the cited papers with respective DOI, and its supplementary information files.
References
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8(4):251–259. https://doi.org/10.1038/nrmicro2312
Apha A (2005) WWE, Standard Methods for the Examination of Water and Wastewater, twenty, first edn. American Public Health Association/American Water Works Association/Water Environment Federation, Washington DC, USA
Aquino SF et al. (2019) Fundamentals of anaerobic sewage treatment. In: Anaerobic reactors for sewage treatment: design, construction and operation. IWA Publishing.
Araújo IS, Oliveira JL, Alves RG, Belli Filho P, da Costa RH (2012) Avaliação de sistema de tratamento de dejetos suínos instalado no Estado de Santa Catarina. Rev Bras de Eng Agricola e Ambient 16(7):745–753. https://doi.org/10.1590/S1415-43662012000700007
ASAE. Agricultural Sanitation and Waste Management Commitec (2003) Manure Production and characteristics. ASAE, Standards D384:1
Bao J, Wang X, Gu J, Dai X, Zhang K, Wang Q, Ma J, Peng H (2020) Effects of macroporous adsorption resin on antibiotic resistance genes and the bacterial community during composting. Bioresour Technol 295:121997. https://doi.org/10.1016/j.biortech.2019.121997
Bengtsson-Palme J, Larsson DJ (2016) Concentrations of antibiotics predicted to select for resistant bacteria: proposed limits for environmental regulation. Environ Int 86:140–149. https://doi.org/10.1016/j.envint.2015.10.015
Chang J, Jiang T, Zhao M, Yang J, Wen Z, Yang F, Li G (2020) Variation pattern of antibiotic resistance genes and microbial community succession during swine manure composting under different aeration strategies. J Chem Technol Biotechnol 95(2):466–473. https://doi.org/10.1002/jctb.6097
Chee-Sanford JC, Mackie RI, Koike S, Krapac IG, Lin YF, Yannarell AC, Maxwell S, Aminov RI (2009) Fate and transport of antibiotic residues and antibiotic resistance genes following land application of maure waste. J Environ Qual 38(3):1086–1108. https://doi.org/10.2134/jeq2008.0128
Chen J, Liu YS, Zhang JN, Yang YQ, Hu LX, Yang YY, Zhao JL, Chen FR, Ying GG (2017) Removal of antibiotics from piggery wastewater by biological aerated filter system: treatment efficiency and biodegradation kinetics. Bioresour Technol 238:70–77. https://doi.org/10.1016/j.biortech.2017.04.023
Chen Y, Zhang H, Luo Y, Song J (2012) Occurrence and dissipation of veterinary antibiotics in two typical swine wastewater treatment systems in east China. Environ Monit Assess 184(4):2205–2217. https://doi.org/10.1007/s10661-011-2110-y
Cheng DL, Ngo HH, Guo WS, Liu YW, Zhou JL, Chang SW, Nguyen DD, Bui XT, Zhang XB (2018) Bioprocessing for elimination antibiotics and hormones from swine wastewater. Sci Total Environ 621:1664–1682. https://doi.org/10.1016/j.scitotenv.2017.10.059
Couch M, Agga GE, Kasumba J, Parekh RR, Loughrin JH, Conte ED (2019) Abundances of tetracycline resistance genes and tetracycline antibiotics during anaerobic digestion of swine waste. J Environ Qual 48(1):171–178. https://doi.org/10.2134/jeq2018.09.0331
Da Cunha C, Freitas MG, da Silva Rodrigues DA, de Barros A, Ribeiro MC, Sanso AL, Afonso R (2021) Low-temperature partitioning extraction followed by liquid chromatography tandem mass spectrometry determination of multiclass antibiotics in solid and soluble wastewater fractions. J Chromatogr A 1650:462256. https://doi.org/10.1016/j.chroma.2021.462256
Dalto DB, da Silva CA (2020) A survey of current levels of trace minerals and vitamins used in commercial diets by the Brazilian pork industry—a comparative study. Trans. Anim Sci 4(4):1–18. https://doi.org/10.1093/tas/txaa195
Dartora V, Perdomo CC, Tumelero IL (1998) Manejo de dejetos de suínos. Boletim Informativo de Pesquisa e Extensão, Embrapa (in Portuguese)
Ding J, Zhu D, Wang Y, Wang H, Liang A, Sun H, Chen Q, Lassen MB, Lv M, Chen L (2021) Exposure to heavy metal and antibiotic enriches antibiotic resistant genes on the tire particles in soil. Sci Total Environ 792:148417. https://doi.org/10.1016/j.scitotenv.2021.148417
Dorival-García N, Zafra-Gómez A, Camino-Sánchez FJ, Navalón A, Vílchez JL (2013) Analysis of quinolone antibiotic derivatives in sewage sludge samples by liquid chromatography-tandem mass spectrometry: comparison of the efficiency of three extraction techniques. Talanta 106:104–118. https://doi.org/10.1016/j.talanta.2012.11.080
Embrapa. Empresa Brasileira de Pesquisa Agropecuária (2003) http://www.cnpsa.embrapa.br/SP/suinos/nutricao.html#:~:text=O%20consumo%20m%C3%A9dio%20%C3%A0%20vontade,por%20dia%2C%20dependendo%20da%20gen%C3%A9tica. Accessed 22 June 2023 (in Portuguese)
Fang H, Wang H, Cai L, Yu Y (2015) Prevalence of antibiotic resistance genes and bacterial pathogens in long-term manured greenhouse soils as revealed by metagenomic survey. Environ Sci Technol 49(2):1095–1104. https://doi.org/10.1021/es504157v
Feng L, Wu H, Zhang J, Brix H (2021) Simultaneous elimination of antibiotics resistance genes and dissolved organic matter in treatment wetlands: Characteristics and associated relationship. Chem Eng Technol 415:128966. https://doi.org/10.1016/j.cej.2021.128966
Ferris MJ, Muyzer G, Ward DM (1996) Denaturing gradient gel electrophoresis profiles of 16S rRNA-defined populations inhabiting a hot spring microbial mat community. Appl Environ Microbiol 62(2):340–346. https://doi.org/10.1128/AEM.62.2.340-346.1996
Gao L, Shi Y, Li W, Niu H, Liu J, Cai Y (2012) Occurrence of antibiotics in eight sewage treatment plants in Beijing, China. Chemosphere 86(6):665–671. https://doi.org/10.1016/j.chemosphere.2011.11.019
Gillings MR (2014) Integrons: past, present, and future. Microbiol Mol Biol Rev 78(2):257–277. https://doi.org/10.1128/MMBR.00056-13
Goldstein C, Lee MD, Sanchez S, Hudson C, Phillips B, Register B, Grady M, Liebert C, Summers AO, White DG, Maurer JJ (2001) Incidence of class 1 and 2 integrases in clinical and commensal bacteria from livestock, companion animals, and exotics. Antimicrob Agents Chemother 45(3):723–726. https://doi.org/10.1128/AAC.45.3.723-726.2001
Gomes RP, Pena CB, Rezende J, Coutrim MX, Afonso RJ (2017) Validation of a new high-throughput method to determine urinary S-phenylmercapturic acid using low-temperature partitioning extraction and ultra high performance liquid chromatography-mass spectrometry. J Sep Sci 40(2):550–557. https://doi.org/10.1002/jssc.201600540
He Y, Yuan Q, Mathieu J, Stadler L, Senehi N, Sun R, Alvarez PJJ (2020) Antibiotic resistance genes from livestock waste: occurrence, dissemination, and treatment. npj. Clean Water 3(4):1–11. https://doi.org/10.1038/s41545-020-0051-0
Huang X, Zheng J, Tian S, Liu C, Liu L, Wei L, Fan H, Zhang T, Wang L, Zhu G, Xu K (2019) Higher temperatures do not always achieve better antibiotic resistance gene removal in anaerobic digestion of swine manure. Appl Environ Microbiol 85(7):e02878–e02818. https://doi.org/10.1128/AEM.02878-18
Jara D, Bello-Toledo H, Domínguez M, Cigarroa C, Fernández P, Vergara L, Quezada-Aguiluz M, Opazo-Capurro A, Lima CA, González-Rocha G (2020) Antibiotic resistance in bacterial isolates from freshwater samples in Fildes Peninsula, King George Island, Antarctica. Sci Rep 10(1):3145. https://doi.org/10.1038/s41598-020-60035-0
Jindal A, Kocherginskaya S, Mehboob A, Robert M, Mackie RI, Raskin L, Zilles JL (2006) Antimicrobial use and resistance in swine waste treatment systems. Appl Environ Microbiol 72(12):7813–7820. https://doi.org/10.1128/AEM.01087-06
Kunz A, Higarashi MM, de Oliveira PA (2005) Tecnologias de manejo e tratamento de dejetos de suínos estudadas no Brasil. Cadernos de Ciência Tecnologia 22(3):651–665. https://doi.org/10.35977/0104-1096.cct2005.v22.8663 (in Portuguese)
Kunz A, Steinmetz RLR, Amaral AC (2022) Fundamentos da digestão anaeróbia, purificação do biogás, uso e tratamento do digestato, 2nd edn. Sbera: Embrapa Suínos e Aves, Concórdia, p 211 (in Portuguese)
Li B, Zhang T (2010) Biodegradation and adsorption of antibiotics in the activated sludge process. Environ Sci Technol 44(9):3468–3473. https://doi.org/10.1021/es903490h
Li J, Wang T, Shao B, Shen J, Wang S, Wu Y (2012) Plasmid-mediated quinolone resistance genes and antibiotic residues in wastewater and soil adjacent to swine feedlots: potential transfer to agricultural lands. Environ Health Perspect 120(8):1144–1149. https://doi.org/10.1289/ehp.1104776
Li X, Liu C, Chen Y, Huang H, Ren T (2018) Antibiotic residues in liquid manure from swine feedlot and their effects on nearby groundwater in regions of North China. Environ Sci Pollut Res Int 25(12):11565–11575. https://doi.org/10.1007/s11356-018-1339-1
Lin H, Sun W, Yu Q, Ma J (2020) Acidic conditions enhance the removal of sulfonamide antibiotics and antibiotic resistance determinants in swine manure. Environ Pollut 263:114439. https://doi.org/10.1016/j.envpol.2020.114439
Liu Y, Cheng D, Xue J, Weaver L, Wakelin SA, Feng Y, Li Z (2020) Changes in microbial community structure during pig manure composting and its relationship to the fate of antibiotics and antibiotic resistance genes. J Hazard Mater 389:122082. https://doi.org/10.1016/j.jhazmat.2020.122082
Liu C, Li G, Qin X, Xu Y, Wang J, Wu G, Feng H, Ye J, Zhu C, Li X, Zheng X (2022) Profiles of antibiotic- and heavy metal-related resistance genes in animal manure revealed using a metagenomic analysis. Ecotoxicol Environ Saf 239:113655. https://doi.org/10.1016/j.ecoenv.2022.113655
Lu XM, Lu PZ (2019) Synergistic effects of key parameters on the fate of antibiotic resistance genes during swine manure composting. Environ Pollut 252(Pt B):1277–1287. https://doi.org/10.1016/j.envpol.2019.06.073
Luo G, Li B, Li LG, Zhang T, Angelidaki I (2017) Antibiotic resistance genes and correlations with microbial community and metal resistance genes in full-scale biogas reactors as revealed by metagenomic analysis. Environ Sci Technol 51(7):4069–4080. https://doi.org/10.1021/acs.est.6b05100
Mao D, Yu S, Luo Y, Yang F, Li F, Hou J, Mu Q, Alvarez P (2015) Prevalence and proliferation of antibiotic resistance genes in two municipal wastewater treatment plants. Water Res 85:458–466. https://doi.org/10.1016/j.watres.2015.09.010
MAPA. Ministério da Agricultura, Pecuária e Abastecimento (2021) Aditivos. https://www.gov.br/agricultura/pt-br/assuntos/insumos-agropecuarios/insumos-pecuarios/alimentacao-animal/aditivos. Accessed 03 May 2021
Mukaka MM (2012) Statistics corner: a guide to appropriate use of correlation coefficient in medical research. Malawi Med J 24(3):69–71
Ndegwa PM, Zhu J, Luo A (2001) Effects of solid levels and chemical additives on removal of solids and phosphorus in swine manure. J Environ Eng 127(12):1111–1115. https://doi.org/10.1061/(ASCE)0733-9372(2001)127:12(1111)
Nguyen THT, Nguyen HD, Le MH, Nguyen TTH, Nguyen TD, Nguyen DL, Nguyen QH, Nguyen TKO, Michalet S, Dijoux-Franca MG, Pham HN (2023) Efflux pump inhibitors in controlling antibiotic resistance: outlook under a heavy metal contamination context. Molecules 28(7):2912. https://doi.org/10.3390/molecules28072912
Okuda T, Yamashita N, Tanaka H, Matsukawa H, Tanabe K (2009) Development of extraction method of pharmaceuticals and their occurrences found in Japanese wastewater treatment plants. Environ Int 35(5):815–820. https://doi.org/10.1016/j.envint.2009.01.006
Oliveira PAV (1993) Manual de manejo e utilização dos dejetos de suínos. In: Embrapa Suínos e Aves-Documentos (INFOTECA-E). (in Portuguese)
Palhares JCP, Calijuri MDC (2007) Caracterização dos afluentes e efluentes suinícolas em sistemas de crescimento/terminação e qualificação de seu impacto ambiental. Ciência Rural 37(2):502–509. https://doi.org/10.1590/S0103-84782007000200032
Pallares-Vega R, Blaak H, van der Plaats R, de Roda Husman AM, Hernandez Leal L, van Loosdrecht M, Weissbrodt DG, Schmitt H (2019) Determinants of presence and removal of antibiotic resistance genes during WWTP treatment: a cross-sectional study. Water Res 161:319–328. https://doi.org/10.1016/j.watres.2019.05.100
Pereira A, Paranhos AGO, de Aquino S, Silva SQ (2021) Distribution of genetic elements associated with antibiotic resistance in treated and untreated animal husbandry waste and wastewater. Environ Sci Pollut Res Int 28:26380–26403. https://doi.org/10.1007/s11356-021-13784-y
Pollock J, Muwonge A, Hutchings MR, Mainda G, Bronsvoort BM, Gally DL, Corbishley A (2020) Resistance to change: AMR gene dynamics on a commercial pig farm with high antimicrobial usage. Sci Rep 10:1708. https://doi.org/10.1038/s41598-020-58659-3
PubChem. National Center for Biotechnology Information (2021) https://pubchem.ncbi.nlm.nih.gov/compound/Norfloxacin. Accessed 05 May 2021.
Qian X, Gu J, Sun W, Wang X, Li H (2019) Effects of passivators on antibiotic resistance genes and related mechanisms during composting of copper-enriched pig manure. Sci Total Environ 674:383–391. https://doi.org/10.1016/j.scitotenv.2019.04.197
Roberts MC (2008) Update on macrolide-lincosamide-streptogramin, ketolide, and oxazolidinone resistance genes. FEMS Microbiol Lett 282(2):147–159. https://doi.org/10.1111/j.1574-6968.2008.01145.x
Rogers HR (1996) Sources, behaviour and fate of organic contaminants during sewage treatment and in sewage sludges. Sci Total Environ 185(1-3):3–26. https://doi.org/10.1016/0048-9697(96)05039-5
Seifrtová M, Nováková L, Lino C, Pena A, Solich P (2009) An overview of analytical methodologies for the determination of antibiotics in environmental waters. Anal Chim Acta 649(2):158–179. https://doi.org/10.1016/j.aca.2009.07.031
Seiler C, Berendonk TU (2012) Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Microbiol 3:399. https://doi.org/10.3389/fmicb.2012.00399
Shi Y, Liu J, Zhuo L, Yan X, Cai F, Luo W, Ren M, Liu Q, Yu Y (2020) Antibiotics in wastewater from multiple sources and surface water of the Yangtze River in Chongqing in China. Environ Monit Assess 192(3):159. https://doi.org/10.1007/s10661-020-8108-6
Sui Q, Zhang J, Tong J, Chen M, Wei Y (2017) Seasonal variation and removal efficiency of antibiotic resistance genes during wastewater treatment of swine farms. Environ Sci Pollut Res Int 24(10):9048–9057. https://doi.org/10.1007/s11356-015-5891-7
Tao CW, Hsu BM, Ji WT, Hsu TK, Kao PM, Hsu CP, Shen SM, Shen TY, Wan TJ, Huang YL (2014) Evaluation of five antibiotic resistance genes in wastewater treatment systems of swine farms by real-time PCR. Sci Total Environ 496:116–121. https://doi.org/10.1016/j.scitotenv.2014.07.024
USDA. United States Department of Agriculture (2020) Market and Trade Data. https://apps.fas.usda.gov/psdonline/app/index.html#/app/advQuery. Accessed 03 May 2021.
Van Boeckel TP, Pires J, Silvester R, Zhao C, Song J, Criscuolo NG, Gilbert M, Bonhoeffer S, Laxminarayan R (2019) Global trends in antimicrobial resistance in animals in low- and middle-income countries. Science 365(6459):eaaw1944. https://doi.org/10.1126/science.aaw1944
Van den Meersche T, Rasschaert G, Haesebrouck F, Van Coillie E, Herman L, Van Weyenberg S, Daeseleire E, Heyndrickx M (2019) Presence and fate of antibiotic residues, antibiotic resistance genes and zoonotic bacteria during biological swine manure treatment. Ecotoxicol Environ Saf 175:29–38. https://doi.org/10.1016/j.ecoenv.2019.01.127
Van Goethem MW, Pierneef R, Bezuidt OK, Van De Peer Y, Cowan DA, Makhalanyane TP (2018) A reservoir of ‘historical’ antibiotic resistance genes in remote pristine Antarctic soils. Microbiome 6(40):1–12. https://doi.org/10.1186/s40168-018-0424-5
Vaz-Moreira I, Harnisz M, Abreu-Silva J, Rolbiecki D, Korzeniewska E, Luczkiewicz A, Manaia CM, Plaza G (2022) Antibiotic resistance in wastewater, does the context matter? Poland and Portugal as a case study. Crit Rev Environ Sci Technol 52(23):4194–4216. https://doi.org/10.1080/10643389.2021.2000828
Vivan M, Kunz A, Stolberg J, Perdomo C, Techio VH (2010) Eficiência da interação biodigestor e lagoas de estabilização na remoção de poluentes em dejetos de suínos. Rev Bras de Eng Agricola e Ambient 14(3):320–325. https://doi.org/10.1590/S1415-43662010000300013
Wang L, Oda Y, Grewal S, Morrison M, Michel FC Jr, Yu Z (2012) Persistence of resistance to erythromycin and tetracycline in swine manure during simulated composting and lagoon treatments. Microb Ecol 63(1):32–40. https://doi.org/10.1007/s00248-011-9921-9
Wang L, Wang J, Wang J, Zhu L, Yang L, Yang R (2019) Distribution characteristics of antibiotic resistant bacteria and genes in fresh and composted manures of livestock farms. Sci Total Environ 695:133781. https://doi.org/10.1016/j.scitotenv.2019.133781
Xia X, Wang Z, Fu Y, Du XD, Gao B, Zhou Y, He J, Wang Y, Shen J, Jiang H, Wu Y (2019) Association of colistin residues and manure treatment with the abundance of mcr-1 gene in swine feedlots. Environ Int 127:361–370. https://doi.org/10.1016/j.envint.2019.03.061
Xu W, Zhang G, Li X, Zou S, Li P, Hu Z, Li J (2007) Occurrence and elimination of antibiotics at four sewage treatment plants in the Pearl River Delta (PRD). South China. Water Res 41(19):4526–4534. https://doi.org/10.1016/j.watres.2007.06.023
Xu Y, Tan L, Li Q, Zheng X, Liu W (2022) Sublethal concentrations of heavy metals Cu2+ and Zn2+ can induce the emergence of bacterial multidrug resistance. Environ Technol Innov 27:102379. https://doi.org/10.1016/j.eti.2022.102379
Yuan K, Yu K, Yang R, Zhang Q, Yang Y, Chen E, Lin L, Luan T, Chen W, Chen B (2019) Metagenomic characterization of antibiotic resistance genes in Antarctic soils. Ecotoxicol Environ Saf 176:300–308. https://doi.org/10.1016/j.ecoenv.2019.03.099
Zhang J, Lu T, Chai Y, Sui Q, Shen P, Wei Y (2019) Which animal type contributes the most to the emission of antibiotic resistance genes in large-scale swine farms in China? Sci Total Environ 658:152–159. https://doi.org/10.1016/j.scitotenv.2018.12.175
Zhang M, Liu YS, Zhao JL, Liu WR, He LY, Zhang JN, Ying GG (2018) Occurrence, fate and mass loadings of antibiotics in two swine wastewater treatment systems. Sci Total Environ 639:1421–1431. https://doi.org/10.1016/j.scitotenv.2018.05.230
Zhang Y, Snow DD, Parker D, Zhou Z, Li X (2013) Intracellular and extracellular antimicrobial resistance genes in the sludge of livestock waste management structures. Environ Sci Technol 47(18):10206–10213. https://doi.org/10.1021/es401964s
Zheng W, Zhang Z, Liu R, Lei Z (2018) Removal of veterinary antibiotics from anaerobically digested swine wastewater using an intermittently aerated sequencing batch reactor. Res J Environ Sci 65:8–17. https://doi.org/10.1016/j.jes.2017.04.011
Acknowledgements
The authors would also like to thank Prof. Juliana Calabria de Araújo, from the Microbiology Laboratory of the Department of Sanitary and Environmental Engineering at the Federal University of Minas Gerais, for providing the positive controls for qPCR.
Funding
This work was supported by the Coordination for the Improvement of Higher Education Personnel (CAPES – financing code 001), Brazilian National Health Foundation (FUNASA- grand number 2510001557612017-21), and National Council for Research and Scientific Development (CNPq – grand number 423101/2018-8).
Author information
Authors and Affiliations
Contributions
Andressa R Pereira: conceptualization, methodology, data curation, software, formal analysis, validation, investigation, and writing—original draft; Lucimeire A B Fonseca: methodology, investigation; Aline G O Paranhos: investigation, methodology, writing—review and editing; Camila C R F Cunha: investigation, methodology, writing—review and editing; Sérgio F Aquino: visualization, writing—review and editing, supervision, resources, funding acquisition; Silvana Q Silva: conceptualization, visualization, writing—review and editing, project administration, supervision, resources, funding acquisition.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Gerald Thouand
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 224 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pereira, .R., de Ávila Barbosa Fonseca, L., Paranhos, A.G. et al. Role of a typical swine liquid manure treatment plant in reducing elements of antibiotic resistance. Environ Sci Pollut Res 30, 91803–91817 (2023). https://doi.org/10.1007/s11356-023-28823-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-28823-z