Abstract
Microbiomes of freshwater basins intended for human use remain poorly studied, with very little known about the microbial response to in situ oil spills. Lake Pertusillo is an artificial freshwater reservoir in Basilicata, Italy, and serves as the primary source of drinking water for more than one and a half million people in the region. Notably, it is located in close proximity to one of the largest oil extraction plants in Europe. The lake suffered a major oil spill in 2017, where approximately 400 tons of crude oil spilled into the lake; importantly, the pollution event provided a rare opportunity to study how the lacustrine microbiome responds to petroleum hydrocarbon contamination. Water samples were collected from Lake Pertusillo 10 months prior to and 3 months after the accident. The presence of hydrocarbons was verified and the taxonomic and functional aspects of the lake microbiome were assessed. The analysis revealed specialized successional patterns of lake microbial communities that were potentially capable of degrading complex, recalcitrant hydrocarbons, including aromatic, chloroaromatic, nitroaromatic, and sulfur containing aromatic hydrocarbons. Our findings indicated that changes in the freshwater microbial community were associated with the oil pollution event, where microbial patterns identified in the lacustrine microbiome 3 months after the oil spill were representative of its hydrocarbonoclastic potential and may serve as effective proxies for lacustrine oil pollution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Petroleum hydrocarbons are one of the most common sources of anthropogenic pollution and cause frequent contamination of both aquatic and terrestrial environments. Due to their potential adverse effects on humans and the environment, primarily manifested through their mutagenic, carcinogenic, endocrine-disrupting, and teratogenic properties, petroleum hydrocarbons represent a public health concern in routine monitoring as their presence, if undetected, may cause irreparable damage (Zhang et al. 2016; Bashir et al. 2020). Releases of petroleum hydrocarbons in the environment due to production, operational use, and transportation are expected to only increase with an ever increasing global energy demand. The development of effective technologies and restoration strategies to overcome such anthropogenic pollution are therefore of great importance, but remain challenging. While various biological systems have been used for restoration of oil-polluted habitats, microbial restoration methods are the most commonly used and often the cheapest alternative. Such remedial technologies are based on the metabolic versatility of microbes which, unlike other complex organisms, have the capability to degrade a diversity of xenobiotic compounds. Microbes, including hydrocarbonoclastic microbes, therefore play an important role in environmental protection and human health through their active role in natural recovery, detoxification, and bioremediation (Xu et al. 2018). Recent advances in our understanding of microbial bioremediation have indicated that such processes do not occur in isolation in the environment and generally involve a consortia of microbes (Mukherjee et al. 2017). The availability of massively parallel sequencing technologies and sophisticated bioinformatic tools have facilitated holistic studies on microbial bioremediation and highlighted the need to study microbiomes in their totality.
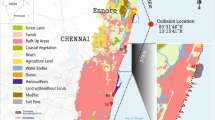
The most frequently documented contamination of water bodies in the 20th century are oil spills, with most occurring in the marine environment and being related to transport or production of crude oil (LJuhasz and Naidu 2000). Oil spills in freshwater ecosystems such as rivers and lakes, by comparison, are much rarer. The water basin and groundwater of Lake Pertusillo, an artificial freshwater reservoir in the Val d’Agri region of Italy, have been reported to be affected by oil extraction activities carried out in 27 oil wells around the lake by Ente Nazionale Idrocarburi (ENI 2019) (Fig. 1). Due to the close proximity of oil extraction activities with respect to Lake Pertusillo, the site was classified by the Italian Ministry of Environment as an environmentally sensitive site with potential for major accidents and adverse environmental impact (Ministry of the Environment 2018). Since 2010, several studies and field analyses have detected anomalous levels of heavy metal and hydrocarbons being present in the lake sediments and water (Colella 2012; Colella and D’Orsogna 2014); among others, seismicity induced by hydraulic fracturing and other injection-based oil extraction techniques have been implicated in such contaminations (Improta et al. 2015; Buttinelli et al. 2016). Furthermore, large-scale and repeated seasonal fish deaths have been observed in the lake during the last decade (De Pace et al. 2019). Recent studies carried out by the Italian Ministry of Health on the ichthyic fauna in Lake Pertusillo detected cyanotoxins, heavy metals, polychlorobiphenyles, and hydrocarbon contamination in several fish species sampled in the lake (De Pace 2015). In February 2017, Lake Pertusillo suffered a major oil spill where over 400 tons of crude oil was spilled from a nearby oil extraction plant into the lake (European Parliament 2017); although unfortunate, the oil spill offered a rare opportunity to study in situ microbial response to oil pollution in a lacustrine environment.
Lake Pertusillo and its surrounding area. Map of the Val d’Agri region of Italy with sites of interest including Lake Pertusillo, major oil extraction installations, groundwater flows, and streams draining into the lake, shown. The sampling sites for the present study, i.e., Madonna Grumentina and Monte Murro, are depicted with black location pins on the map
The coexistence of water resources in the proximity of highly polluting anthropogenic activities, such as oil extraction plants, petrochemical plants, and oil platforms represents a serious concern in conservation of freshwater ecosystems and as an extension, protection of human health (European Environment Agency 2019). However, since oil spills in freshwater environments are relatively rare, little is known of the microbial response to petroleum hydrocarbon contamination in these environments. In the present study, we have carried out culture-independent analyses of microbial communities in a lacustrine ecosystem impacted by oil spills where we have inferred changes in the lake microbiome in the aftermath of the pollution event along with metagenome prediction and consequent deductions on hydrocarbonoclastic functionalities. Our studies not only demonstrate changes in the lake microbial community due to oil pollution, but also identify principal hydrocarbonoclastic actors and major functional pathways involved in hydrocarbon degradation. Additionally, we also infer the microbial community succession in Lake Pertusillo after the oil pollution event. To our knowledge, our study represents one of the few studies carried out on oil pollution and subsequent microbial response in lacustrine ecosystems, and insights gained from the study should be useful in better monitoring and sustainable use of freshwater resources as well as designing of novel bioremediation strategies.
Materials and methods
Study site and sample collection
Samples were collected before and after the pollution event in February 2017, where 400 tons of crude oil was spilled from a nearby oil extraction plant run by Ente Nazionale Idrocarburi into Lake Pertusillo. Details for collected samples are given in Table 1. Certified analyses for detection of total petroleum hydrocarbons (TPH) in water samples were performed by the Regional Agency for Environment Protection of Basilicata (ARPAB) and civic associations (ARPAB 2017a; Cova Contro 2017a, b). Samples 1B, M16, and MM represent superficial samples and were collected from just under the water surface, whereas sample 2B was collected near the bottom of the lake; such a sampling design was pursued in order to understand changes in the microbial community structure along the water column. Water samples were collected using 1-L Pyrex glass bottles and stored in ice (4 °C) boxes; samples were later processed using a previously described ultrafiltration protocol (D’Ugo et al. 2016). The elutriate (50 mL) was used for downstream laboratory investigations. To study the metabolic potential of the microbiome, microcosms were prepared using sample 1B as an inoculum; the microcosm (sample 1B_LB) was supplemented with Luria-Bertani (LB) broth and incubated at 30 °C for a week. Inoculum from 1B_LB was further employed to enrich hydrocarbonoclastic bacteria in a microcosm supplemented with diesel (D) as the sole carbon source, i.e., sample 1B_LB_D, and incubated at 30 °C for a week.
For extraction of total genomic DNA, 5 mL of environmental and microcosm samples were centrifuged at 6000 rcf for 10 min at room temperature. The pellet was used for DNA extraction. Total environmental DNA was extracted from samples using the Genomic DNA tissue kit (Machery-Nagel, Thermo Fisher, Italy) and quantified using a spectrophotometer (NanoDrop, Thermo Fisher). High-throughput sequencing of V3-V4 and V3-V5 regions of the 16S rRNA gene for the microbiome was performed by Eurofins MWG Operon (Ebersberg, Germany). All sequencing data generated in the study has been deposited in the NCBI Sequence Read Archive (SRA) under the BioProject number PRJNA412797.
Bioinformatic analysis
16S rRNA sequences generated in the study were evaluated for quality using FastQC (Andrews 2010), and subsequently processed in mothur (Schloss et al. 2009) with trimming of adapters, primers, and barcodes. Additional quality control was performed using the following criteria: maxhomop = 6, maxambig = 0, minlength = 200, qwindowaverage = 30, bdiffs = 1, pdiffs = 2, and tdiffs = 2. Mothur was subsequently used to align quality-filtered sequences to the SILVA (Quast et al. 2013) database. Chimeric sequences were removed using the mothur implementation of vsearch (Rognes et al. 2016). QIIME (Quantitative Insights Into Microbial Ecology) was used for open reference OTU (Operational Taxonomic Units) calling and taxonomic assignment of high-quality 16S rRNA sequences against the SILVA nr v132 release. Taxonomic classification was summarized in QIIME and subsequently visualized in R with ggplot2 (Wickham 2009). Metabolic reconstruction for 16S rRNA datasets was conducted in PICRUSt v2.0.0 (Douglas et al. 2019). Metagenomes for the 16S rRNA datasets were predicted using BIOM abundance files and representative sequence files as input; these were generated in QIIME during OTU calling and taxonomic assignment, respectively. Predicted metagenomes were further collapsed into MetaCyc pathways by the MinPath (Langille et al. 2013) implemented within PICRUSt 2.0.0 and abundances for the same were also obtained. Inferred metagenomes were visualized as cladograms generated with the standalone graphical tool GraPhlan v0.95 (Ye and Doak 2009).
Results
Changes in microbial community composition observed at the Phylum level
To better understand the changes in hydrocarbonoclastic properties of the Lake Pertusillo microbiome after the oil spill, bioinformatic analysis was directed towards the study of bacteria putatively capable of degrading hydrocarbons. To this end, 16S rRNA gene sequence analysis of lake samples was performed for samples collected both before (MM and M16) and after oil spill (1B and 2B) (Fig. 1). Comprehensive taxonomic analysis was carried out for each sample, along with predictive metabolic reconstruction to elucidate the metabolic potential of the microbial community.
Taxonomic analysis of Lake Pertusillo samples revealed that phylum Proteobacteria was the predominant phylum in oil-contaminated samples (1B and 2B). The proportion of phylum Proteobacteria in the microbiome increased notably from 52% before oil spill (sample M16), to 81% and 77% (samples 1B and 2B, respectively) after the pollution event (Supplementary Figure S1). Additionally, the M16 sample, collected at same coordinates as samples 1B and 2B before the oil spill, showed much higher levels (24%) of bacteria assigned to phylum Actinobacteria (Supplementary Figure S1). The phylum was appreciably reduced in proportion in samples 1B and 2B (5.4% and 4.3%, respectively) compared to the M16 sample indicating an adverse response of actinobacterial microbes to oil contamination in the lake. Surprisingly, in the MM sample, Actinobacteria represented the dominant phylum and not Proteobacteria (Supplementary Figure S1).
To understand if hydrocarbonoclastic properties of Lake Pertusillo microbes were primarily sequestered in the Proteobacterial phylum, 1B samples were enriched for hydrocarbon-degrading bacteria. To do this, 1B samples were added to a rich medium (Luria-Bertani (LB) broth) without any hydrocarbons and incubated for a week (sample 1B_LB). Taxonomic analysis of the 1B_LB microbiome revealed that in the absence of hydrocarbons, the microbial community composition changes considerably; this was represented by a decline of phylum Proteobacteria (from 81% to 23%) and increase of phylum Firmicutes (from 0.1% to 75%; Supplementary Figure S1 and S2). When hydrocarbons were re-introduced into the 1B_LB sample (1B_LB_D), where diesel (D) was provided as the sole energy source, the microbiota displayed a rapid shift towards a Proteobacteria-dominated population (99%; Supplementary Figure S2). Thus, it can be observed that, while environmental samples 1B and 2B (total petroleum hydrocarbon, TPH: 900 μg/L) and the microcosm with diesel (1B_LB_D) maintain a microbiome structure dominated by Proteobacteria, an absence of hydrocarbons in the 1B_LB sample caused a drastic reduction of Proteobacteria and increase in Firmicutes; taken together, these indicate that most hydrocarbonoclastic bacteria in Lake Pertusillo belong to phylum Proteobacteria.
Changes in microbial community composition observed at the Genus level
To understand the Lake Pertusillo microbiome further and how it changed when exposed to petroleum hydrocarbon contamination, we analysed the Pertusillo microbiome before and after the oil spill at the genus level and compared it with microbiomes from two pristine Bulgarian reservoirs (Supplementary Table S1) (Iliev et al. 2017). Our analyses revealed that contaminated samples from Lake Pertusillo were enriched for alphaproteobacterial and betaproteobacterial genera with specialized hydrocarbonoclastic competence, as has been observed previously in other oil-contaminated ecosystems (Table S1) (Yang et al. 2014; Kostka et al. 2011). Hydrogenophaga sp., a member of the Betaproteobacteria family Comamonadaceae and previously reported in oil sludge samples in Saudi Arabia (Albokari et al. 2015), was found to be notably enriched in the two contaminated samples 1B and 2B (1B: 22.7%; 2B: 8.9%) in comparison to lake samples before oil spills (MM: 0.0%, M16: 0.0%) (Supplementary Table S1). Similar differences in abundances were observed for Hydrogenophaga when compared to pristine Bulgarian lacustrine ecosystems where all five sites analyzed showed a Hydrogenophaga abundance under 1% (Supplementary Table S1).
Another member of class Betaproteobacteria, Acidovorax sp., is also well represented in the oil-contaminated lake microbiome, showing a notable increase compared to unpolluted Lake Pertusillo samples (1B: 3.5%; 2B: 12.2% ; MM 0.0% and M16 0.0%) and pristine Bulgarian lake samples (Supplementary Table S1).
Afipia sp., belonging to Class Alphaproteobacteria, was enriched in the two contaminated Lake Pertusillo samples (1B: 3.5%; 2B: 12.2%) compared to all other lake samples (Supplementary Table S1). Interestingly, Reyranella sp. (Class Alphaproteobacteria) was also highly enriched in the oil-contaminated Lake Pertusillo samples (1B: 7.3%; 2B: 2.%) in comparison to unpolluted Lake Pertusillo samples and Bulgarian water basin samples (Supplementary Table S1).
Dyadobacter sp., a member of Phylum Bacteroidetes, was found to be notably enriched only in the oil-contaminated sample 1B (1B: 2.79%, Supplementary Table S1). The versatile genus Variovorax (Class Betaproteobacteria) was also enriched only in sample 1B (1B: 17.8 %; Supplementary Table S1).
In samples analyzed along the contaminated water column, i.e., samples 1B and 2B, we detected two Alphaproteobacterial genera, Sphingopyxis (1B: 1.1%; 2B: 2.2% and Hirschia (1B: 0.8%; 2B: 1.2% Supplementary Table S1).
Metabolic reconstruction of Lake Pertusillo microbiomes
To functionally characterize the Lake Pertusillo microbiome, metabolic reconstruction was conducted in PICRUSt v2.0.0 with metabolic pathways being inferred using MinPath. Bioinformatic analysis confirmed a hydrocarbonoclastic metabolic landscape for samples 1B and 2B, wherein multiple metabolic pathways involved in degradation of aromatic hydrocarbons were detected (Fig. 2). These include metabolic pathways involved in the degradation of aromatic hydrocarbons such as catechol, phenol, phthalate, ethylbenzene, protocatechuate, and toluene; metabolic pathways involved in mineralization of chlorinated hydrocarbons such as PCBs and other chloroaromatic and sulfonated aromatic hydrocarbons were also detected (Fig. 2b and Supplementary Table S2).
a Microbiome contributions of OTUs at the genus level in sample 1B. b Metabolic reconstruction of sample 1B and 2B. A subset of pathways related to hydrocarbon degradation and nitrogen and sulfur metabolism for samples 1B and 2B is depicted in the circular cladogram. Each ring of the cladogram from the center denotes a level of the pathway ontology. Corresponding labels are found from the outermost ring to the final clade in the sector. Each clade is depicted by a circular marker
Discussion
Microbial response to oil contamination in freshwater ecosystems, especially lacustrine habitats, is poorly studied. Studies on the microbial community composition and metabolic landscape of polluted lacustrine environments contribute to our understanding of the dynamics of microbial response and in turn, facilitate better conservation of water resources. Recently, high-throughput sequencing has facilitated the advancement of microbial ecological studies in polluted habitats with bioinformatic approaches being used to reveal metagenomic characteristics of bacterial response to petroleum hydrocarbon contamination in diverse ecosystems. Using 16S rRNA gene sequences as an input in PICRUSt (Douglas et al. 2019) and other similar bioinformatic tools (Aßhauer et al. 2015), it is possible to predict metagenomics features and functional compositions of microbiomes; subsequently, critically important and enriched taxa and functional pathways can be inferred as biomarkers that can be effectively used to distinguish diverse oil-polluted environments (Mukherjee et al. 2017).
Proteobacteria dominate the oil-polluted lacustrine microbiome
In the present study, Proteobacteria were found to be the dominant phylum in the oil polluted Lake Pertusillo microbiome. The Proteobacterial dominance of an oil-contaminated Lake Pertusillo microbiome is in agreement with previous observations from other ecosystems which suffered oil pollution events (Mukherjee et al. 2017). For example, Yang et al. have previously reported a similar shift in microbial community structure towards phylum Proteobacteria upon oil contamination in permafrost habitats (Yang et al. 2014). Members of this phylum, such as those belonging to the classes Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria, are known to be capable of using various hydrocarbons as their sole carbon source (Kostka et al. 2011). Unsurprisingly, when evaluated at the genus level, contaminated samples from Lake Pertusillo were found to be enriched for alphaproteobacterial and betaproteobacterial genera.
Hydrogenophaga sp., a member of the Betaproteobacteria family Comamonadaceae, was found to be notably enriched in the two contaminated samples 1B and 2B. The genus has previously been isolated from microbial consortia in benzene-contaminated sites (Fahy et al. 2008) along with Acidovorax sp. and Pseudomonas sp., both of which were also detected in our study. The presence of Hydrogenophaga sp. was also described in microcosms set up using groundwater from a BTEX (benzene, toluene, ethylbenzene, and xylenes) contaminated site as inoculum with a mixture of toluene and benzene used as sole carbon sources (Aburto and Peimbert 2011). Hydrogenophaga sp. have additionally been reported to play an important role in PAH-degrading (polycyclic aromatic hydrocarbons) microbial communities (Martin et al. 2012). Furthermore, a role for Hydrogenophaga in polychlorinated biphenyls (PCBs) degradation has been proposed previously (Lambo and Patel 2006). These data are largely in agreement with a previous study, which detected PCBs in ichthyic fauna of Lake Pertusillo (De Pace 2015) and with the ARPAB (ARPAB 2017b) that stresses that PCBs such as PAH are present in the sediment of the entire reservoir with higher concentrations in 2017. Acidovorax, another betaproteobacterial genera, was well represented in the oil-contaminated samples. Acidovorax has previously been found to be the most abundant dominant bacterial species in the sludge of an Alberta oil sand tailing pond (Singleton et al. 2009). Like other members of Comamonadaceae, Acidovorax is frequently encountered in association with PAH degradation (Singleton et al. 2018). Denitrifying Acidovorax sp. has been isolated from terrestrial subsurface sediments exposed to mixed-waste contamination; in particular, this genus has been detected in nitrate-reducing microbial consortia cultivated with alkylated aromatic compounds, revealing their important role in hydrocarbons and nitrate-polluted waters (Sperfeld et al. 2018). The versatile betaproteobacterial genus Variovorax sp. was enriched only in sample 1B; the genus has been previously reported for hydrocarbonoclastic properties such as biosynthesis of biosurfactants (Franzetti et al. 2012) and PAH degradation in the presence of nitrates (Eriksson et al. 2003).
Afipia sp., belonging to class Alphaproteobacteria, was enriched in the two contaminated Lake Pertusillo samples. Similar to Acidovorax sp., Afipia sp. is a well-known degrader of polycyclic aromatic substances (Willumsen et al. 2005) and can degrade PAHs using nitrates as oxidizing agents; such combination of PAH degradation with denitrification by Afipia sp. has been observed in nitrate-contaminated groundwater (Green et al. 2010). Reyranella sp., an alphaproteobacterial genus which was found to be enriched in oil-contaminated samples, has been previously reported in degradation of asphaltenes, a particularly recalcitrant fraction of crude oil (Song et al. 2018). In samples analyzed along the water column, i.e., samples 1B and 2B, the alphaproteobacterial genera Sphingopyxis and Hirschia were detected. Both these genera have been previously identified to play important roles in microbiome successions in relation to oil spills (Rodriguez et al. 2015). In the first phase (during the oil spill), sand microbiomes in beaches affected by the Deep Water Horizon oil spill showed a prevalence of microbes that degrade aliphatic hydrocarbons and were replaced in the second phase (2–3 months after the oil spill) with a PAH-degrading microbial community enriched in Sphingopyxis and Hirschia (Rodriguez et al. 2015).
Dyadobacter sp., a member of Phylum Bacteroidetes, was found to be enriched only in the oil-contaminated sample 1B; members of the genus are particularly specialized in degradation of complex aromatic hydrocarbons such as azaarenes, which are relatively water soluble, nitrogen-containing heterocyclic aromatic hydrocarbons that have been reported to leach from contaminated soils and sediments into aqueous habitats (Pereria et al. 1983). The isolation and study of these enriched species identified herein, among others, could help us better understand lacustrine microbial response to oil pollution and may serve as excellent tools for future bioremediation interventions and as microbial proxies for oil pollution in lakes; taken together, these will contribute to protection of freshwater ecosystems and reduction of public health risks. Indeed, recent further studies concentrated on the hydrocarbonoclastic biofilms formed on the water surface after the Lake Pertusillo oil pollution event have revealed unique electrogenic structural and biochemical properties of the biofilm and sheds further light on possible mechanisms of microbial detoxification of oil-contaminated lacustrine habitats (D’Ugo et al. 2021). Overall, the presence of a hydrocarbonoclastic microbial community characterized by an abundance of proteobacterial genera such as Hydrogenophaga, Acidovorax, Reyranella, and Variovorax among others, highlighted the lake's pollution status, indicating a community shaped by residual recalcitrant hydrocarbons derived from the oil spill 3 months earlier and with significant complex hydrocarbon degradation potential.
Metabolic landscape of polluted lacustrine microbiome reveals microbial hydrocarbonoclastic potential
Detection of hydrocarbonoclastic pathways involved in the degradation of complex aromatic hydrocarbons in metagenomes predicted through PICRUSt is consistent with the hydrocarbon degradation potential of several genera found to be enriched in petroleum hydrocarbon-contaminated Pertusillo samples. Indeed, microbial genera such Hydrogenophaga, Acidovorax, Reyranella, Variovorax, and others, found to be enriched in polluted samples as described above, exhibit multiple metabolic pathways involved in degradation of aromatic hydrocarbons. For example, mineralization pathways for pyrene and benzo-[a]-pyrene along with other high-molecular weight PAHs have been reported previously for Hydrogenophaga sp. (Yan et al. 2017). Additionally, metabolic pathways for the degradation of chloroaromatics have also been identified in Hydrogenophaga sp., including transformation of 2,4′-dichlorobiphenyl (2,4′-DCB) into 2- and 4-chlorobenzoic acid (2- and 4-CBA; Lambo and Patel 2006). Furthermore, degradation of sulfo-nitroaromatic compounds have also been reported for Hydrogenophaga sp.; indeed, Gan et al. reported the mineralization of 4-aminobenzenesulfonate (4-ABS) by Hydrogenophaga sp. strain PBC involving enzymatic conversions catalyzed by 4-sulfocatechol 1,2-dioxygenase, 3-sulfomuconate cycloisomerase, and 3,4-dioxygenase enzymes (Gan et al. 2011). The versatility of Hydrogenophaga sp. as a degrader of PAHs could explain the increased abundance of the genus in oil-contaminated samples, as mentioned above (Supplementary Table S1). PAH degrading capabilities have also been reported for other genera found to be abundant in oil-contaminated samples, i.e., Acidovorax, Reyranella, and Variovorax, with metabolic pathways employing diverse hydrocarbonoclastic enzymes such as ring-hydroxylating dioxygenase (Singleton et al. 2009), catechol 1,2-dioxygenase (El Azhari et al. 2010), and hydrolytic esterases (Wang and Gu 2006), among others, being identified in them.
Although it was not possible to analyze the microbial community structure of Lake Pertusillo during the oil spill, our results indicate that hydrocarbonoclastic capacities of the lacustrine microbiome in the aftermath of the oil spill were highly skewed towards mineralization of complex hydrocarbons including PAHs, choroaromatics, nitroaromatics, and sulfonated aromatic compounds with few metabolic pathways for degradation of aliphatic petroleum hydrocarbons detected. This suggests that there may have been a shift in the microbiome towards a specific hydrocarbonoclastic competence at this stage, i.e., 3 months after the oil spill, where the structure of the Pertusillo microbiome could be shaped by the residual presence of more recalcitrant, complex hydrocarbons such as benzene derivatives, toluene, PAH, PCB, and BTEX, among others, after most aliphatic hydrocarbons had been degraded. Such an observation is substantiated further by detection of several genera of microbes known to specialize in PAH and complex hydrocarbon degradation, as described above, as well as previously reported similar observations from the Deep Water Horizon oil spill where the second stage of microbial successions was dominated by PAH degrading microbes (Rodriguez et al. 2015; Martirani-Von Abercron et al. 2017). Importantly, predictive metabolic reconstruction using PICRUSt demonstrated that advanced bioinformatic pipelines can be successfully employed in dissecting complex freshwater ecosystem metabolomes, where hydrocarbonoclastic contributions of members of microbial communities in polluted habitats can be inferred and in turn, provide important insights.
Conclusions
While multiple studies have been conducted on microbial response to petroleum hydrocarbon contamination in the marine environment, little is known about microbial response to oil spills in lacustrine and freshwater environments owing to scarcity of studies on the same. In the present study, we have described the microbial response to an oil spill in Lake Pertusillo in February 2017, including taxonomic and functional characterizations for polluted and non-polluted samples. Our study has identified several taxa such as Hydrogenophaga, Variovorax, Reyranella, Acidovorax, and Afipia, among others, that may be used as microbial proxies for oil pollution in freshwater environments. Additionally, functional characterization of microbial response using bioinformatic approaches identified several metabolic pathways that may be involved in mineralization of Lake Pertusillo oil pollutants. Microbial successions in an oil-contaminated Lake Pertusillo revealed similarities with observations reported from marine oil spills, where mature microbial communities specialized in degradation of recalcitrant complex petroleum hydrocarbons. The detection of a specifically modified microbial community after a suspected pollution event, in this case concerning petroleum hydrocarbons, allows us to devise hypotheses about the causes and beginning of such events, acting as a “cold case” detection even after 3 months had elapsed following the occurrence. Since most freshwater ecosystems are used as sources of potable drinking water, microbial proxies identified herein and methods for detecting alterations in the lacustrine microbiome and its metabolic potential can be valuable tools in detecting pollution events and preventing major public health complications. To our knowledge, the present study is one of the few conducted on oil spills in lacustrine environments and will contribute further to our understanding of the origins and temporal dynamics of microbiome alterations in freshwater ecosystems impacted by petroleum hydrocarbon contamination.
Change history
05 May 2021
A Correction to this paper has been published: https://doi.org/10.1007/s11356-021-14244-3
References
Aburto A, Peimbert M (2011) Degradation of a benzene-toluene mixture by hydrocarbon-adapted bacterial communities. Ann Microbiol 61(3):553–562
Albokari M, Mashhour I, Alshehri M, Boothman C, Al-Enezi M (2015) Characterization of microbial communities in heavy crude oil from Saudi Arabia. Ann Microbiol 65(1):95–104
Andrews S (2010). FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc.
ARPAB (2017a). http://www.arpab.it/risorse_idriche/public/RELAZIONE%20PERTUSILLO_19102017_pubblicato_rev_031117.pdf (last access 24/09/2020)
ARPAB 2017b. http://www.arpab.it/risorse_idriche/rapporti2017.asp, last access 20/06/2019
Aßhauer KP, Wemheuer B, Daniel R, Meinicke P (2015) Tax4Fun: predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics 31(17):2882–2884. https://doi.org/10.1093/bioinformatics/btv287
Bashir I, Lone FA, Bhat RA, Mir SA, Dar ZA, Dar SA (2020) Concerns and threats of contamination on aquatic ecosystems bioremediation and biotechnology. Jan 27:1–26. https://doi.org/10.1007/978-3-030-35691-0_1
Buttinelli M, Improta L, Bagh S, Chiarabba C (2016) Inversion of inherited thrusts by wastewater injection induced seismicity at the Val d’Agri oilfield (Italy). Sci Rep 6:37165. https://doi.org/10.1038/srep37165
Colella A (2012) Hydrocarbons and metals in waters and sediments of the Pertusillo Lake, Italy. Fresenius Environ Bull 21(10a):3003–3011
Colella A, D’Orsogna MR (2014) Hydrocarbon contamination in waters and sediments of the Pertusillo freshwater reservoir, Val D’agri, Southern Italy. Fresenius Environ Bull 23(12b):3286–3295
Cova Contro (2017a).http://analizebasilicata.altervista.org/blog/ancora-idrocarburi-ed-azoto-nel-pertusillo-anche-il-quarto-campione-e-contaminato, last access 20/06/2019
Cova Contro (2017b). http://analizebasilicata.altervista.org/blog/idrocarburi-nel-pertusillo-presenti-in-tre-campioni-su-tre-oltre-a-fosfati-e manganese/?doing_wp_cron=1517936436.7830340862274169921875, last access 20/06/2019
D’Ugo E, Marcheggiani S, Fioramonti I, Giuseppetti R, Spurio R, Helmi K, Guillebault D, Medlin LK, Simeonovski I, Boots B, Breitenbach U, Koker L, Albay M, Mancini L (2016) Detection of human enteric viruses in freshwater from European countries. Food Environ Virol 8:206–214. https://doi.org/10.1007/s12560-016-9238-4
D’Ugo E, Bertuccini L, Spadaro F, Giuseppetti R, Iosi F, Santavenere F, Giuliani F, Gricia M, Rodomonte A, Lovecchio N, Mukherjee A, Bucci P, Bruno M, Stellacci E, Bernardo A, Magurano F (2021) Water Res 197:117092. https://doi.org/10.1016/j.watres.2021.117092
De Pace R (2015). Convegno “Gli animali quali indicatori biologici dell’inquinamento ambientale: strategie di monitoraggio.” Calvello, PT (Italy) 6 June 2015. http://docplayer.it/1712392-Gli-animali-quali-indicatori-biologici-dell-inquinamento-ambientale-strategie-di-monitoraggio.html.
De Pace R, Storelli MM, Barone G, Messineo V, Bruno M (2019) Co-occurrence of polychlorinated biphenyls, cyanotoxins and trace elements in commercial fish species from a freshwater protected area (Pertusillo Lake, Southern Italy). Journal of Geography, Environment and Earth Science International 22:1–14. https://doi.org/10.9734/jgeesi/2019/v22i430152
Douglas GM et al (2019) PICRUSt2: an improved and extensible approach for metagenome inference. bioRxiv preprint. https://doi.org/10.1101/672295
El Azhari N, Devers-Lamrani M, Chatagnier G, Rouard N, Martin-Laurent F (2010 May 15) (2010). Molecular analysis of the catechol-degrading bacterial community in a coal wasteland heavily contaminated with PAHs. J Hazard Mater 177(1-3):593–601. https://doi.org/10.1016/j.jhazmat.2009.12.074
ENI (2019). https://www.eniday.com/en/education_en/largest-oil-field-europe/last access: 12/6/2019
Eriksson M, Sodersten E, Yu Z, Dalhammar G, Mohn WW (2003) Degradation of polycyclic aromatic hydrocarbons at low temperature under aerobic and nitrate-reducing conditions in enrichment cultures from northern soils. Appl Environ Microbiol. Jan 69(1):275–284
European Environment Agency (2019). https://www.eea.europa.eu/themes/water/european-waters/water-use-and-environmental-pressures
European Parliament (2017). www.europarl.europa.eu/doceo/document/P-8-2017-001300_EN.html?redirect (last access september 24, 2020).
Fahy A, Ball AS, Lethbridge G, Timmis KN, McGenity TJ (2008) Isolation of alkali-tolerant benzene-degrading bacteria from a contaminated aquifer. Lett Appl Microbiol 47(1):60–66
Franzetti A, Gandolfi I, Raimondi C, Bestetti G, Banat IM, Smyth TJ, Papacchini M, Cavallo M, Fracchia L (2012) Environmental fate, toxicity, characteristics and potential applications of novel bioemulsifiers produced by Variovorax paradoxus 7bCT5. Bioresour Technol 108:245–251
Gan HM, Ibrahim Z, Shahir S, Yahya A (2011) Identification of genes involved in the 4-aminobenzenesulfonate degradation pathway of Hydrogenophaga sp. PBC via transposon mutagenesis. FEMS Microbiol Lett 318(2):108–114. https://doi.org/10.1111/j.1574-6968.2011.02245.x
Green SJ et al (2010) Denitrifying bacteria isolated from terrestrial subsurface sediments exposed to mixed-waste contamination. Appl Environ Microbiol. May 76(10):3244–3254
Iliev I et al (2017) Metagenomic profiling of the microbial freshwater communities in two Bulgarian reservoirs. Basic Microbiol 57(8):669–679
Improta L, Valoroso L, Piccinini D, Chiarabba C (2015) A detailed analysis of wastewater-induced seismicity in the Val d’Agri oil field (Italy). Geophys Res Lett 42:2682–2690. https://doi.org/10.1002/2015GL063369
Kostka JE, Prakash O, Overholt WA, Green SJ, Freyer G, Canion A et al (2011) Hydrocarbon-degrading bacteria and the bacterial community response in Gulf of Mexico beach sands impacted by the deepwater horizon oil spill. Appl Environ Microbiol 77:7962–7974. https://doi.org/10.1128/AEM.05402-11
Lambo AJ, Patel TR (2006) Cometabolic degradation of polychlorinated biphenyls at low temperature by psychrotolerant bacterium Hydrogenophaga sp. Curr Microbiol 53(1):48–52
Langille MG et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(9):814–821
LJuhasz A, Naidu R (2000) Bioremediation of high molecular weight polycyclic aromatic hydrocarbons: a review of the microbial degradation of benzo[a]pyrenel. Int Biodeterior Biodegradation 45(1–2):57–88
Martin F et al (2012) Betaproteobacteria dominance and diversity shifts in the bacterial community of a PAH-contaminated soil exposed to phenanthrene. Environ Pollut Mar 162:345–353
Martirani-Von Abercron SM, Marín P, Solsona-Ferraz M, Castañeda-Cataña MA, Marqués S (2017) Naphthalene biodegradation under oxygen-limiting conditions: community dynamics and the relevance of biofilm-forming capacity. Microb Biotechnol 10(6):1781–1796
Ministry of the Environment (2018).https://www.minambiente.it/sites/default/files/archivio/allegati/stabilimenti_rischio_industriale/2018/basilicata_31_12_2018. pdf ; last access: 20/6/2019
Mukherjee A, Chettri B, Langpoklakpam JS, Basak P, Prasad A, Mukherjee MA, Bhattacharyya KM, Singh AK, Chattopadhyay D (2017) Bioinformatic approaches including predictive metagenomic profiling reveal characteristics of bacterial response to petroleum hydrocarbon contamination in diverse environments. Sci Rep 7(1):1108. https://doi.org/10.1038/s41598-017-01126-3
Pereria WE, Rostad CE, Garbarino JR, Hult MF (1983) Groundwater contamination by organic bases derived from coal-tar waste. Environ Toxicol Chem 2:283–294
Quast C et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools Quast C. Nucleic Acids Res 41:590–596
Rodriguez RLM, Overholt WA, Hagan C, Huettel M, Kostka JE, Konstantinidis KT (2015) Microbial community successional patterns in beach sands impacted by the Deepwater Horizon oil spill. ISME J 9(9):1928–1940
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584 eCollection
Schloss PD et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Singleton DR, Ramirez LG, Aitken MD (2009) Characterization of a polycyclic aromatic hydrocarbon degradation gene cluster in a phenanthrene-degrading Acidovorax strain. Appl Environ Microbiol 75(9):2613–2620
Singleton DR, Lee J, Dickey AN, Stroud A, Scholl EH, Wright FA, Aitken MD (2018) Polyphasic characterization of four soil-derived phenanthrene-degrading Acidovorax strains and proposal of Acidovorax carolinensis sp. nov. Syst Appl Microbiol 41(5):460–472
Song WF et al (2018) Shifts of the indigenous microbial communities from reservoir production water in crude oil- and asphaltene-degrading microcosms. Int Biodeterior Biodegradation 132:18–29
Sperfeld M, Diekert G, Studenik S, Sperfeld M, Diekert G, Studenik S (2018). Community dynamics in a nitrate-reducing microbial consortium cultivated with p-alkylated vs. non-p-alkylated aromatic compounds. FEMS Microbiol Ecol
Wang YP, Gu JD (2006) Degradability of dimethyl terephthalate by Variovorax paradoxus T4 and Sphingomonas yanoikuyae DOS01 isolated from deep-ocean sediments. Ecotoxicology 15:549–557
Wickham H (2009) ggplot2: Elegant Graphics for Data Analysis. Springer, London, p 210
Willumsen PA, Johansen JE, Karlson U, Hansen BM (2005) Isolation and taxonomic affiliation of N-heterocyclic aromatic hydrocarbon-transforming bacteria. Appl Microbiol Biotechnol 67(3):420–428
Xu X et al (2018) Petroleum hydrocarbon-degrading bacteria for the remediation of oil pollution under aerobic conditions: a perspective analysis. Front Microbiol 9:2885. https://doi.org/10.3389/fmicb.2018.02885
Yan Z, Zhanga Y, Wu H, Yangab M, Zhangab H, Haoac Z, Jianga H (2017) Isolation and characterization of a bacterial strain Hydrogenophaga sp. PYR1 for anaerobic pyrene and benzo[a]pyrene biodegradation. RSC Adv 7:46690–46698. https://doi.org/10.1039/C7RA09274A
Yang S, Wen X, Zhao L, Shi Y, Jin H (2014) Crude oil treatment leads to shift of bacterial communities in soils from the deep active layer and upper permafrost along the China-Russia Crude Oil Pipeline route. PLoS One 9(5):e96552. https://doi.org/10.1371/journal.pone.009655
Ye Y, Doak TG (2009) A parsimony approach to biological pathway reconstruction/inference for genomes and metagenomes. PLoS Comput Biol 5(8):e1000465
Zhang Y, Dong S, Wang H, Tao S, Kiyama R (2016) Biological impact of environmental polycyclic aromatic hydrocarbons (ePAHs) as endocrine disruptors. Environ Pollut 213:809–824. https://doi.org/10.1016/j.envpol.2016.03.050
Acknowledgements
We thank Cosimo Curianò for his help in preparation of figures.
Availability of data and materials
The datasets generated and/or analysed during the current study are available at https://figshare.com/s/93d3b639c6f1f7487ef1.
Funding
Intramural funding.
Author information
Authors and Affiliations
Contributions
E.D.: conceptualization, methodology, writing-original draft preparation, visualization, validation, writing-reviewing. M.B.: validation, investigation, writing-reviewing. A.M.: methodology, validation, software, investigation, writing-reviewing. R.G.: validation, investigation. R.D.: conceptualization. F.M.: software, writing-reviewing, funding acquisition.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(PDF 1993 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
D’Ugo, E., Bruno, M., Mukherjee, A. et al. Characterization of microbial response to petroleum hydrocarbon contamination in a lacustrine ecosystem. Environ Sci Pollut Res 28, 26187–26196 (2021). https://doi.org/10.1007/s11356-021-13885-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-13885-8