Abstract
Background
Pulmonary sarcoidosis (SAR) and tuberculosis (TB) are two granulomatous lung-diseases and often pose a diagnostic challenge to a treating physicians.
Objective
The present study aims to explore the diagnostic potential of NMR based serum metabolomics approach to differentiate SAR from TB.
Materials and Method
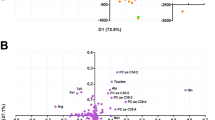
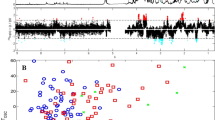
The blood samples were obtained from three study groups: SAR (N = 35), TB (N = 28) and healthy normal subjects (NC, N = 56) and their serum metabolic profiles were measured using 1D 1H CPMG (Carr-Purcell-Meiboom-Gill) NMR spectra recorded at 800 MHz NMR spectrometer. The quantitative metabolic profiles were compared employing a combination of univariate and multivariate statistical analysis methods and evaluated for their diagnostic potential using receiver operating characteristic (ROC) curve analysis.
Results
Compared to SAR, the sera of TB patients were characterized by (a) elevated levels of lactate, acetate, 3-hydroxybutyrate (3HB), glutamate and succinate (b) decreased levels of glucose, citrate, pyruvate, glutamine, and several lipid and membrane metabolites (such as very-low/low density lipoproteins (VLDL/LDL), polyunsaturated fatty acids, etc.).
Conclusion
The metabolic disturbances not only found to be well in concordance with various previous reports, these further demonstrated very high sensitivity and specificity to distinguish SAR from TB patients suggesting serum metabolomics analysis can serve as surrogate method in the diagnosis and clinical management of SAR.





Similar content being viewed by others
Data availability
The data that support the findings of this study has been uploaded on ZENODO (https://zenodo.org/record/ 5,593,941 | https://doi.org/10.5281/zenodo.5593941) and is available without undue reservation for further studies on request to the corresponding authors.
Abbreviations
- 1D/2D:
-
One/Two dimensional
- 3HB:
-
3-Hydroxy butyrate
- ANOVA:
-
One-way analysis of variance
- ATB:
-
Active Tuberculosis
- AUROC:
-
Area under ROC curve
- BCAAs:
-
Branched‐chain amino acids
- BMRB:
-
Biological Magnetic Resonance Data Bank
- CI:
-
Confidence interval
- CPMG:
-
Carr–Purcell–Meiboom–Gill
- CT:
-
Computerized tomography
- ELISA:
-
Enzyme-linked Immunosorbent Assay
- ESM:
-
Electronic Supplementary Material
- FID:
-
Free induction decay
- FT:
-
Fourier Transformation
- HMDB:
-
The Human Metabolome Database
- HSQC:
-
Heteronuclear Single Quantum Correlation
- IGRA:
-
Interferon-gamma Release Assay
- IL:
-
Interleukin
- IQR:
-
Interquartile range
- LDL:
-
Low-density lipoproteins
- LTBI:
-
Latent Tuberculosis infection
- MTB:
-
Mycobacterium tuberculosis
- MVA:
-
Multivariate analysis
- MW:
-
Molecular Weight
- NC:
-
Normal control
- NMR:
-
Nuclear Magnetic Resonance
- NSAID:
-
Non-steroidal anti-inflammatory drugs
- OPLS-DA:
-
PLS-DA with Orthogonal Signal Correction (OSC)
- PCA:
-
Principal component analysis
- PLS-DA:
-
Projection to least-squares discriminant analysis
- POC:
-
Point-of-care
- PTR:
-
Phenylalanine-to-tyrosine ratio
- RCF:
-
Relative centrifugal force
- ROC:
-
Receiver operating characteristic curve
- RPM:
-
Revolutions per minute
- SAR:
-
Sarcoidosis
- SD:
-
Standard deviation
- TB:
-
Tuberculosis
- TCA:
-
Tricarboxylic acid cycle
- TMAO:
-
Trimethylamine-N-oxide
- TOCSY:
-
Total Correlation Spectroscopy
- TST:
-
Tuberculin skin test
- VIP:
-
Variable importance in projection
- VLDL:
-
Very Low-density lipoproteins
- WHO:
-
World Health Organization
- μL/ml:
-
Microliter/Milliliter
References
Babu, K. (2013). Sarcoidosis in tuberculosis-endemic regions: India. Journal of Ophthalmic Inflammation and Infection, 3, 1–7.
Badar, F., Azfar, S. F., Ahmad, I., Yasmeen, S., & Kirmani, S. (2011a). Diagnostic difficulties in differentiating sarcoidosis from tuberculosis. Oman Medical Journal, 26, 210–211.
Beckonert, O., Keun, H. C., Ebbels, T. M., Bundy, J., Holmes, E., Lindon, J. C., & Nicholson, J. K. (2007). Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nature Protocols, 2, 2692–2703.
Bunsow, E., Ruiz-Serrano, M. A. J. S., Roa, P. L. P., Kestler, M., Viedma, D. G., & Bouza, E. (2014). Evaluation of GeneXpert MTB/RIF for the detection of Mycobacterium tuberculosis and resistance to rifampin in clinical specimens. Journal of Infection, 68, 338–343.
Chaiyachat, P., Kaewseekhao, B., Chaiprasert, A., Kamolwat, P., Nonghanphithak, D., Phetcharaburanin, J., Sirichoat, A., Ong, R.T.-H., & Faksri, K. (2023). Metabolomic analysis of Mycobacterium tuberculosis reveals metabolic profiles for identification of drug-resistant tuberculosis. Scientific Reports, 13, 8655.
Chakaya, J., Khan, M., Ntoumi, F., Aklillu, E., Fatima, R., Mwaba, P., Kapata, N., Mfinanga, S., Hasnain, S. E., Katoto, P. D. (2021). Global Tuberculosis Report 2020-Reflections on the Global TB burden, treatment and prevention efforts. International Journal of Infectious Diseases 113(Supplement 1), S7–S12.
Chen, Y., Wu, J., Tu, L., Xiong, X., Hu, X., Huang, J., Xu, Z., Zhang, X., Hu, C., & Hu, X. (2013a). 1 H-NMR spectroscopy revealed Mycobacterium tuberculosis caused abnormal serum metabolic profile of cattle. PLoS ONE, 8, e74507.
Chen, Y., Wu, J., Tu, L., Xiong, X., Hu, X., Huang, J., Xu, Z., Zhang, X., Hu, C., & Hu, X. (2013b). 1H-NMR spectroscopy revealed mycobacterium tuberculosis caused abnormal serum metabolic profile of cattle. PLoS ONE, 8, e74507.
Chen, X., Ye, J., Lei, H., & Wang, C. (2022). Novel potential diagnostic serum biomarkers of metabolomics in osteoarticular tuberculosis patients: A preliminary study. Frontiers in Cellular and Infection Microbiology, 12, 90.
Cho, Y., Park, Y., Sim, B., Kim, J., Lee, H., Cho, S. N., Kang, Y., & Lee, S. G. (2020). Identification of serum biomarkers for active pulmonary tuberculosis using a targeted metabolomics approach. Scientific Reports, 10, 1–11.
Chong, J., Soufan, O., Li, C., Caraus, I., Li, S., Bourque, G., Wishart, D. S., & Xia, J. (2018). MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Research, 46, 486–494.
Conde, R., Laires, R., Goncalves, L. G., Rizvi, A., Barroso, C., Villar, M., Macedo, R., Simoes, M. J., Gaddam, S., & Lamosa, P. (2022). Discovery of serum biomarkers for diagnosis of tuberculosis by NMR metabolomics including cross-validation with a second cohort. Biomedical Journal, 45, 654–664.
Das, M. K., Bishwal, S. C., Das, A., Dabral, D., Badireddy, V. K., Pandit, B., Varghese, G. M., & Nanda, R. K. (2015). Deregulated tyrosine-phenylalanine metabolism in pulmonary tuberculosis patients. Journal of Proteome Research, 14, 1947–1956.
De Buck, J., Shaykhutdinov, R., Barkema, H. W., & Vogel, H. J. (2014b). Metabolomic profiling in cattle experimentally infected with Mycobacterium avium subsp. paratuberculosis. PLoS ONE, 9, 111872.
Debik, J., Sangermani, M., Wang, F., Madssen, T. S., & Giskedegnrd, G. F. (2022). Multivariate analysis of NMR-based metabolomic data. NMR in Biomedicine, 35, 4638.
Du Preez, I., & Luies, L. (2017). Metabolomics biomarkers for tuberculosis diagnostics: Current status and future objectives. Biomarkers, 11, 179–194.
Du Preez, I., & Luies, L. (2019). The application of metabolomics toward pulmonary tuberculosis research. Tuberculosis, 115, 126–139.
Du Bois, R. M., Goh, N., McGrath, D., & Cullinan, P. (2003). Is there a role for microorganisms in the pathogenesis of sarcoidosis? Journal of Internal Medicine, 253, 4–17.
Dubaniewicz, A. (2010). Mycobacterium tuberculosis heat shock proteins and autoimmunity in sarcoidosis. Autoimmunity Reviews, 9, 419–424.
Dubaniewicz, A. (2013). Microbial and human heat shock proteins as “danger signals” in sarcoidosis. Human Immunology, 74, 1550–1558.
Dubaniewicz, A., Holownia, A., Kalinowski, L., Wybieralska, M., Dobrucki, I. T., & Singh, M. (2013). Is mycobacterial heat shock protein 16 kDa, a marker of the dormant stage of Mycobacterium tuberculosis, a sarcoid antigen? Human Immunology, 74, 45–51.
Dubey, D., Chaurasia, S., Guleria, A., Kumar, S., Modi, D. R., Misra, R., & Kumar, D. (2019). Metabolite assignment of ultrafiltered synovial fluid extracted from knee joints of reactive arthritis patients using high resolution NMR spectroscopy. Magnetic Resonance in Chemistry, 57, 30–43.
Dubey, D., Kumar, S., Rawat, A., Guleria, A., Kumari, R., Ahmed, S., Singh, R., Misra, R., & Kumar, D. (2021). NMR-based metabolomics revealed the underlying inflammatory pathology in reactive arthritis synovial joints. Journal of Proteome Research, 20, 5088–5102.
Duchemann, B., Triba, M. N., Guez, D., Rzeznik, M., Savarin, P., Nunes, H., Valeyre, D., Bernaudin, J. F., & Le Moyec, L. (2016). Nuclear magnetic resonance spectroscopic analysis of salivary metabolome in sarcoidosis. Sarcoidosis Vasculitis and Diffuse Lung Disease, 33, 10–16.
Eriksson, L., Johansson, E., Kettaneh-Wold, N., & Wold, S. (1999). Introduction to multi-and megavariate data analysis using projection methods (PCA & PLS). Umetrics AB 1999, 1–490
Frediani, J. K., Jones, D. P., Tukvadze, N., Uppal, K., Sanikidze, E., Kipiani, M., Tran, V. T., Hebbar, G., Walker, D. I., & Kempker, R. R. (2014). Plasma metabolomics in human pulmonary tuberculosis disease: A pilot study. PLoS ONE, 9, e108854.
Fuchs, J. E., Huber, R. G., Von Grafenstein, S., Wallnoefer, H. G., Spitzer, G. M., Fuchs, D., & Liedl, K. R. (2012). Dynamic regulation of phenylalanine hydroxylase by simulated redox manipulation. PLoS ONE, 7, e53005.
Geamanu, A., Gupta, S. V., Bauerfeld, C., & Samavati, L. (2016). Metabolomics connects aberrant bioenergetic, transmethylation, and gut microbiota in sarcoidosis. Metabolomics, 12, 35.
Gebremicael, G.; Amare, Y.; Challa, F.; Gebreegziabxier, A.; Medhin, G.; Wolde, M.; Kassa, D (2017). Lipid profile in tuberculosis patients with and without human immunodeficiency virus infection. International Journal of Chronic Diseases 2017, Article ID 3843291
Geisler, S., Gostner, J. M., Becker, K., Ueberall, F., & Fuchs, D. (2013). Immune activation and inflammation increase the plasma phenylalanine-to-tyrosine ratio. Pteridines, 24, 27–31.
Gleeson, L. E., Sheedy, F. J., Palsson-McDermott, E. M., Triglia, D., O’Leary, S. M., O’Sullivan, M. P., O’Neill, L. A., & Keane, J. (2016). Cutting edge: Mycobacterium tuberculosis induces aerobic glycolysis in human alveolar macrophages that is required for control of intracellular bacillary replication. The Journal of Immunology, 196, 2444–2449.
Guleria, A., Misra, D. P., Rawat, A., Dubey, D., Khetrapal, C. L., Bacon, P., Misra, R., & Kumar, D. (2015). NMR-Based Serum Metabolomics Discriminates Takayasu Arteritis from Healthy Individuals: A Proof-of-Principle Study. Journal of Proteome Research, 14, 3372–3381.
Guleria, A., Pratap, A., Dubey, D., Rawat, A., Chaurasia, S., Sukesh, E., Phatak, S., Ajmani, S., Kumar, U., Khetrapal, C. L., Bacon, P., Misra, R., & Kumar, D. (2016). NMR based serum metabolomics reveals a distinctive signature in patients with Lupus Nephritis. Science and Reports, 6, 35309.
Guleria, A., Kumar, A., Kumar, U., Raj, R., & Kumar, D. (2018). NMR Based Metabolomics: An Exquisite and Facile Method for Evaluating Therapeutic Efficacy and Screening Drug Toxicity. Current Topics in Medicinal Chemistry, 18, 1827–1849.
Guleria, A., Kumar, U., Kumar, D., Anuja, A. K., Singh, M. K., Sharma, P., Agarwal, V., Misra, R., & Gupta, L. (2021). NMR-based serum and muscle metabolomics for diagnosis and activity assessment in idiopathic inflammatory myopathies. Analytical Science Advances, 2, 515–526.
Gupta, D., Agarwal, R., Aggarwal, A. N., & Jindal, S. K. (2012). Sarcoidosis and tuberculosis: The same disease with different manifestations or similar manifestations of different disorders. Current Opinion in Pulmonary Medicine, 18, 506–516.
Gupta, L., Ahmed, S., Jain, A., & Misra, R. (2018). Emerging role of metabolomics in rheumatology. International Journal of Rheumatic Diseases, 21, 1468–1477.
Gupta, L., Guleria, A., Rawat, A., Kumar, D., & Aggarwal, A. (2021). NMR-based clinical metabolomics revealed distinctive serum metabolic profiles in patients with spondyloarthritis. Magnetic Resonance in Chemistry, 59, 85–98.
Haas, C. T., Roe, J. K., Pollara, G., Mehta, M., & Noursadeghi, M. (2016). Diagnostic “omics” for active tuberculosis. BMC Medicine, 14, 1–19.
Huang, H., Han, Y. S., Chen, J., Shi, L. Y., Wei, L. L., Jiang, T. T., Yi, W. J., Yu, Y., Li, Z. B., & Li, J. C. (2020). The novel potential biomarkers for multidrug-resistance tuberculosis using UPLC-Q-TOF-MS. Experimental Biology and Medicine, 245, 501–511.
Jain, A., Kumar, A., Singh, H., Rai, M. K., Chaturvedi, S., Guleria, A., Nath, A., Kumar, D., Misra, D. P., & Agarwal, V. (2018). 114 Nuclear magnetic resonance (NMR) based serum metabolomics in sarcoidosis and tuberculosis: search for a biomarker. Rheumatology, 57, 075–338.
Kim, M.-J., Wainwright, H. C., Locketz, M., Bekker, L.-G., Walther, G. B., Dittrich, C., Visser, A., Wang, W., Hsu, F.-F., & Wiehart, U. (2010). Caseation of human tuberculosis granulomas correlates with elevated host lipid metabolism. EMBO Molecular Medicine, 2, 258–274.
Koeken, V. A., Lachmandas, E., Riza, A., Matzaraki, V., Li, Y., Kumar, V., Oosting, M., Joosten, L. A., Netea, M. G., & van Crevel, R (2019). Role of Glutamine Metabolism in Host Defense Against Mycobacterium tuberculosis Infection. The Journal of infectious diseases, 219 (10), 1662–1670.
Kumar, R., Goel, N., & Gaur, S. N. (2012). Sarcoidosis in north Indian population: A retrospective study. Indian Journal of Chest Diseases and Allied Sciences, 54, 99.
Kumar, U., Jain, A., Guleria, A., Misra, D. P., Goel, R., Danda, D., Misra, R., & Kumar, D. (2020). Circulatory glutamine/glucose ratio for evaluating disease activity in Takayasu arteritis: A NMR based serum metabolomics study. Journal of Pharmaceutical and Biomedical Analysis, 180, 113080.
Kumar, U., Mehta, P., Kumar, S., Jain, A., Guleria, A., Kumar, R., Misra, R., & Kumar, D. (2021). Circulatory histidine levels as predictive indicators of disease activity in takayasu arteritis. Analytical Science Advances, 2, 527–535.
Lee, W., VanderVen, B. C., Fahey, R. J., & Russell, D. G. (2013). Intracellular Mycobacterium tuberculosis exploits host-derived fatty acids to limit metabolic stress. Journal of Biological Chemistry, 288, 6788–6800.
Liebenberg, C., Luies, L., & Williams, A. A. (2021). Metabolomics as a tool to investigate HIV/TB co-infection. Frontiers in Molecular Biosciences, 8, 692823.
Loke, W. S. J., Herbert, C., Thomas, P. S. (2013) Sarcoidosis: Immunopathogenesis and immunological markers. International Journal of Chronic Diseases, 2013 (928601), 1–13.
McGaha, T. L., Huang, L., Lemos, H., Metz, R., Mautino, M., Prendergast, G. C., & Mellor, A. L. (2012). Amino acid catabolism: A pivotal regulator of innate and adaptive immunity. Immunological Reviews, 249, 135–157.
Muhammed, H., Kumar, D., Dubey, D., Kumar, S., Chaurasia, S., Guleria, A., Majumder, S., Singh, R., Agarwal, V., & Misra, R. (2020). Metabolomics analysis revealed significantly higher synovial Phe/Tyr ratio in reactive arthritis and undifferentiated spondyloarthropathy. Rheumatology, 59, 1587–1590.
Murr, C., Grammer, T. B., Meinitzer, A., Kleber, M. E., März, W., Fuchs, D, (2014). Immune activation and inflammation in patients with cardiovascular disease are associated with higher phenylalanine to tyrosine ratios: the ludwigshafen risk and cardiovascular health study. Journal of amino acids 2014(783730), 1–6.
Neurauter, G., Schrocksnadel, K., Scholl-Burgi, S., Sperner-Unterweger, B., Schubert, C., Ledochowski, M., & Fuchs, D. (2008). Chronic immune stimulation correlates with reduced phenylalanine turnover. Current Drug Metabolism, 9, 622–627.
Nicholson, J. K., Foxall, P. J., Spraul, M., Farrant, R. D., & Lindon, J. C. (1995). 750 MHz 1H and 1H–13C NMR spectroscopy of human blood plasma. Analytical Chemistry, 67, 793–811.
Nikam, C., Kazi, M., Nair, C., Jaggannath, M., Manoj, M., Vinaya, R., Shetty, A., & Rodrigues, C. (2014). Evaluation of the Indian TrueNAT micro RT-PCR device with GeneXpert for case detection of pulmonary tuberculosis. International Journal of Mycobacteriology, 3, 205–210.
Ouyang, X., Dai, Y., Wen, J. L., & Wang, L. X. (2011). 1H NMR-based metabolomic study of metabolic profiling for systemic lupus erythematosus. Lupus, 20, 1411–1420.
Ploder, M., Neurauter, G., Spittler, A., Schroecksnadel, K., Roth, E., & Fuchs, D. (2008). Serum phenylalanine in patients post trauma and with sepsis correlate to neopterin concentrations. Amino Acids, 35, 303–307.
Régo, A. M., da Silva, D. A., Ferreira, N. V., de Pina, L. C., Evaristo, J. A., Evaristo, G. P. C., Nogueira, F. C., Ochs, S. M., Amaral, J. J., & Ferreira, R. B. (2021). Metabolic profiles of multidrug resistant and extensively drug resistant Mycobacterium tuberculosis unveiled by metabolomics. Tuberculosis, 126, 102043.
Schnappinger, D., Schoolnik, G. K., & Ehrt, S. (2006). Expression profiling of host pathogen interactions: How Mycobacterium tuberculosis and the macrophage adapt to one another. Microbes and Infection, 8, 1132–1140.
Sharma, S. K., & Mohan, A. (2004). Sarcoidosis in India: Not so rare. Journal, Indian Academy of Clinical Medicine, 5, 12–21.
Shi, L., Salamon, H., Eugenin, E. A., Pine, R., Cooper, A., & Gennaro, M. L. (2015). Infection with Mycobacterium tuberculosis induces the Warburg effect in mouse lungs. Scientific Reports, 5, 18176.
Shi, L., Eugenin, E. A., & Subbian, S. (2016). Immunometabolism in Tuberculosis. Frontiers in Immunology, 7, 150.
Shimazu, T., Hirschey, M. D., Newman, J., He, W., Shirakawa, K., Le Moan, N., Grueter, C. A., Lim, H., Saunders, L. R., & Stevens, R. D. (2013). Suppression of oxidative stress by b-hydroxybutyrate, an endogenous histone deacetylase inhibitor. Science, 339, 211–214.
Shin, J. H., Yang, J. Y., Jeon, B. Y., Yoon, Y. J., Cho, S. N., Kang, Y. H., Ryu, D. H., & Hwang, G. S. (2011). 1H NMR-based metabolomic profiling in mice infected with Mycobacterium tuberculosis. Journal of Proteome Research, 10, 2238–2247.
Singh, V., Kaur, C., Chaudhary, V. K., Rao, K. V., & Chatterjee, S. M. (2015). tuberculosis secretory protein ESAT-6 induces metabolic flux perturbations to drive foamy macrophage differentiation. Scientific Reports, 5, 12906.
Somashekar, B. S., Amin, A. G., Tripathi, P., MacKinnon, N., Rithner, C. D., Shanley, C. A., Basaraba, R., Henao-Tamayo, M., Kato-Maeda, M., & Ramamoorthy, A. (2012). Metabolomic signatures in guinea pigs infected with epidemic-associated W-Beijing strains of Mycobacterium tuberculosis. Journal of Proteome Research, 11, 4873–4884.
Stanley, S. A., & Cox, J. S. (2013). Host–Pathogen interactions during mycobacterium tuberculosis infections. In: J. Pieters, & J. McKinney (Eds.), Pathogenesis of mycobacterium tuberculosis and its Interaction with the host organism. Current Topics in Microbiology and Immunology (vol. 374). Springer, Berlin, Heidelberg. https://doi.org/10.1007/82_2013_332
Statement on Sarcoidosis Joint Statement of the American Thoracic Society (ATS), the European Respiratory Society (ERS) and the World Association of Sarcoidosis and Other Granulomatous Disorders (WASOG) adopted by the ATS Board of Directors and by the ERS Executive Committee, February 1999. Am J Respir Crit Care Med 1999, 160, 736–755.
Szymanska, E., Saccenti, E., Smilde, A. K., & Westerhuis, J. A. (2012). Double-check: Validation of diagnostic statistics for PLS-DA models in metabolomics studies. Metabolomics, 8, 3–16.
Tang, H., Wang, Y., Nicholson, J. K., & Lindon, J. C. (2004). Use of relaxation-edited one-dimensional and two dimensional nuclear magnetic resonance spectroscopy to improve detection of small metabolites in blood plasma. Analytical Biochemistry, 325, 260–272.
Thillai, M., Eberhardt, C., Lewin, A. M., Potiphar, L., Hingley-Wilson, S., Sridhar, S., Macintyre, J., Kon, O. M., Wickremasinghe, M., & Wells, A. (2012). Sarcoidosis and tuberculosis cytokine profiles: Indistinguishable in bronchoalveolar lavage but different in blood. PLoS ONE, 7, e38083.
Thulasidoss, K., Asokan, L., Chandra, P., & Rejliwal, P. (2017). The clinical conundrum of diagnosing and treating systemic sarcoidosis in a high TB burden area. BMJ case reports, 2017. https://doi.org/10.1136/bcr-2016-218741.
Tuck, M. K., Chan, D. W., Chia, D., Godwin, A. K., Grizzle, W. E., Krueger, K. E., Rom, W., Sanda, M., Sorbara, L., & Stass, S. (2009). Standard operating procedures for serum and plasma collection: Early detection research network consensus statement standard operating procedure integration working group. Journal of Proteome Research, 8, 113–117.
Vrieling, F., Alisjahbana, B., Sahiratmadja, E., van Crevel, R., Harms, A. C., Hankemeier, T., Ottenhoff, T. H., & Joosten, S. A. (2019). Plasma metabolomics in tuberculosis patients with and without concurrent type 2 diabetes at diagnosis and during antibiotic treatment. Scientific Reports, 9, 18669.
Wannemacher, R. W., Jr., Klainer, A. S., Dinterman, R. E., & Beisel, W. R. (1976). The significance and mechanism of an increased serum phenylalanine-tyrosine ratio during infection. The American Journal of Clinical Nutrition, 29, 997–1006.
Weiner, J., 3rd., Parida, S. K., Maertzdorf, J., Black, G. F., Repsilber, D., Telaar, A., Mohney, R. P., Arndt-Sullivan, C., Ganoza, C. A., & Faé, K. C. (2012). Biomarkers of Inflammation, Immunosuppression and Stress Are Revealed by Metabolomic Profiling of Tuberculosis Patients. PLoS ONE, 7, e40221.
World Health Organization 2020 Global Tuberculosis Burden. who. int/iris/bitstream/handle/10665/336069/9789240013131-eng. pdf 2021,
Wu, T., Xie, C., Han, J., Ye, Y., Weiel, J., Li, Q., Blanco, I., Ahn, C., Olsen, N., Putterman, C., Saxena, R., & Mohan, C. (2012). Metabolic disturbances associated with systemic lupus erythematosus. PLoS ONE, 7, e37210.
Xia, J., & Wishart, D. S. (2011). Web-based inference of biological patterns, functions and pathways from metabolomic data using MetaboAnalyst. Nature Protocols, 6, 743–760.
Xia, J., Psychogios, N., Young, N., & Wishart, D. S. (2009). MetaboAnalyst: A web server for metabolomic data analysis and interpretation. Nucleic Acids Research, 37, W652–W660.
Zangerle, R., Kurz, K., Neurauter, G., Kitchen, M., Sarcletti, M., & Fuchs, D. (2010). Increased blood phenylalanine to tyrosine ratio in HIV-1 infection and correction following effective antiretroviral therapy. Brain, Behavior, and Immunity, 24, 403–408.
Zhou, Y., Li, H. P., Li, Q. H., Zheng, H., Zhang, R. X., Chen, G., & Baughman, R. P. (2008). Differentiation of sarcoidosis from tuberculosis using real-time PCR assay for the detection and quantification of Mycobacterium tuberculosis. Sarcoidosis, Vasculitis, and Diffuse Lung Diseases: Official Journal of WASOG, 25, 93–99.
Zhou, A., Ni, J., Xu, Z., Wang, Y., Lu, S., Sha, W., Karakousis, P. C., & Yao, Y. F. (2013). Application of 1H NMR spectroscopy-based metabolomics to sera of tuberculosis patients. Journal of Proteome Research, 12, 4642–4649.
Zhou, A., Ni, J., Xu, Z., Wang, Y., Zhang, H., Wu, W., Lu, S., Karakousis, P. C., & Yao, Y. F. (2015). Metabolomics specificity of tuberculosis plasma revealed by 1 H NMR spectroscopy. Tuberculosis, 95, 294–302.
Acknowledgements
DK acknowledges the Department of Medical Education, Govt. of Uttar Pradesh for supporting the High Field NMR Facility at Centre of Biomedical Research, Lucknow, India. RR acknowledges the receipt of a SRF fellowship from CSIR, INDIA.The manuscript communication number is IU/R&D/2023-MCN0002213.
Funding
This research received intramural funding from the Centre of Biomedical Research (CBMR), Lucknow (Project No. CBMR/IMR/0010/2021 | PI: Dr. Dinesh Kumar).
Author information
Authors and Affiliations
Contributions
AJ and VA conceptualized the idea; AJ, MKR, HS, KS, SC, AN, DPM, VA identified the patients and collected the samples. AK, RR, DD, SY, and ARK prepared the NMR samples and performed the NMR experiments; MKR, SY, ARK and DK: NMR data analysis and multivariate/univariate statistics. DK and AG wrote the first draft. AJ, VA and DK critically reviewed and revised the draft. All authors read and verified the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
On behalf of all authors, the corresponding authors state that there is no conflict of interest. The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11306_2023_2052_MOESM1_ESM.doc
Supplementary file1 Figure S1: Multivariate exploratory ROC analysis based on eighteen discriminatory metabolites of diagnostic potential. Figure S2: The box-cum-whisker plots showing quantitative differences for eighteen discriminatory serum metabolic entities between SAR and TB. Table S1: The serum metabolic features indexed top to bottom based on variable importance in projection (VIP) score values derived from the PLS-DA analysis of complete and pruned CPMG data matrix. Table S2: The spectral features strategically selected from the 0.02 ppm binned CPMG data matrix for univariate statistical and ROC curve analysis. Table S3: Serum metabolic features identified for statistically significant difference between the study-groups using ANOVA statistics. (DOC 8617 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rai, M.K., Yadav, S., Jain, A. et al. Clinical metabolomics by NMR revealed serum metabolic signatures for differentiating sarcoidosis from tuberculosis. Metabolomics 19, 92 (2023). https://doi.org/10.1007/s11306-023-02052-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-023-02052-4