Abstract
Western redcedar (Thuja plicata; Cupressaceae; WRC) is an ecologically and economically important conifer species of the Pacific Northwest. Regeneration of WRC forests is affected by ungulate browsing, which removes current growth and hampers development of young trees. Monoterpenes make WRC foliage less palatable and can deter browsing. Genomic resources are required to advance knowledge of terpene accumulation and breeding of WRC for herbivore resistance. Unlike most conifers, WRC readily selfs to produce genotypes of reduced heterozygosity. We used seedlings of eight different fifth-generation selfed lines for monoterpene analysis and transcriptome sequencing. Trinity, Velvet/Oases, TransABySS, and SOAPdenovoTrans were used to generate independent transcriptome assemblies for each line. Sequence redundancy was reduced using the EvidentialGene pipeline. The best assembly, as determined by metrics of completeness, contiguity, and accuracy, was used to produce a WRC reference gene set of 28,279 sequences, of which 77% were annotated with significant BLASTp hits and 89% with significant InterProScan hits. An orthology-based approach was used to annotate gene families. Manually curated annotation identified 33 putative full-length terpene synthases (TPS). A maximum likelihood phylogeny revealed that WRC TPS cluster apart from those of Pinaceae within the gymnosperm TPS-d clade. Use of selfed lines enabled the development and annotation of a reduced-redundancy gene set for a gymnosperm of the Cupressaceae family. This gene set serves as a foundation for future functional characterization of WRC TPS and other defense genes and as a resource for the annotation of protein coding sequences in the WRC genome.



Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Baillie R, Drayton M, Pembleton L, Kaur S, Culvenor R, Smith K, Spangenberg G, Forster J, Cogan N (2017) Generation and characterisation of a reference transcriptome for Phalaris (Phalaris aquatica L.). Agronomy 7:14. https://doi.org/10.3390/agronomy7010014
Birol I, Raymond A, Jackman SD, Pleasance S, Coope R, Taylor GA, Yuen MMS, Keeling CI, Brand D, Vandervalk BP, Kirk H, Pandoh P, Moore RA, Zhao Y, Mungall AJ, Jaquish B, Yanchuk A, Ritland C, Boyle B, Bousquet J, Ritland K, MacKay J, Bohlmann J, Jones SJM (2013) Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shotgun sequencing data. Bioinformatics 29:1492–1497. https://doi.org/10.1093/bioinformatics/btt178
Blande D, Halimaa P, Tervahauta AI, Aarts MGM, Kärenlampi SO (2017) De novo transcriptome assemblies of four accessions of the metal hyperaccumulator plant Noccaea caerulescens. Sci Data 4:1–9. https://doi.org/10.1038/sdata.2016.131
Bohlmann J, Keeling CI (2008) Terpenoid biomaterials. Plant J 54:656–669
Bohlmann J, Meyer-Gauen G, Croteau R (1998) Plant terpenoid synthases: molecular biology and phylogenetic analysis. Proc Natl Acad Sci U S A 95:4126–4133. https://doi.org/10.1073/pnas.95.8.4126
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:421. https://doi.org/10.1186/1471-2105-10-421
Chen F, Tholl D, Bohlmann J, Pichersky E (2011) The family of terpene synthases in plants: a mid-size family of genes for specialized metabolism that is highly diversified throughout the kingdom. Plant J 66:212–229. https://doi.org/10.1111/j.1365-313X.2011.04520.x
De La Torre AR, Birol I, Bousquet J et al (2014) Insights into conifer giga-genomes. Plant Physiol 166:1724–1732. https://doi.org/10.1104/pp.114.248708
Debell JD, Morrell JJ, Gartner BL (1999) Within-stem variation in tropolone content and decay resistance of second-growth Western redcedar. For Sci 45:101–107
Duan J, Xia C, Zhao G, Jia J, Kong X (2012) Optimizing de novo common wheat transcriptome assembly using short-read RNA-Seq data. BMC Genomics 13:392. https://doi.org/10.1186/1471-2164-13-392
Elbeltagy A, Nishioka K, Suzuki H, Sato T, Sato YI, Morisaki H, Mitsui H, Minamisawa K (2000) Isolation and characterization of endophytic bacteria from wild and traditionally cultivated rice varieties. Soil Sci Plant Nutr 463:617–629. https://doi.org/10.1080/00380768.2000.10409127
Emms DM, Kelly S (2015) OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol 16:157. https://doi.org/10.1186/s13059-015-0721-2
Foster AJ, Hall DE, Mortimer L, Abercromby S, Gries R, Gries G, Bohlmann J, Russell J, Mattsson J (2013) Identification of genes in Thuja plicata foliar terpenoid defenses. Plant Physiol 161:1993–2004. https://doi.org/10.1104/pp.112.206383
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Geddy R, Brown GG (2007) Genes encoding pentatricopeptide repeat (PPR) proteins are not conserved in location in plant genomes and may be subject to diversifying selection. BMC Genomics 8:130. https://doi.org/10.1186/1471-2164-8-130
Gesell A, Blaukopf M, Madilao L, Yuen MMS, Withers SG, Mattsson J, Russell JH, Bohlmann J (2015) The gymnosperm cytochrome P450 CYP750B1 catalyzes stereospecific monoterpene hydroxylation of (+)-sabinene in thujone biosynthesis in western redcedar. Plant Physiol 168:94–106. https://doi.org/10.1104/pp.15.00315
Gilbert D (2013) Gene-omes built from mRNA seq not genome DNA. In: 7th Annual Arthropod Genomics Symposium. Notre Dame
Gonzalez JS (2004) Growth, properties and uses of western red cedar. Forintek Canada Corp Spec Publ No SP-37R 37
Gordon SP, Tseng E, Salamov A, Zhang J, Meng X, Zhao Z, Kang D, Underwood J, Grigoriev IV, Figueroa M, Schilling JS, Chen F, Wang Z (2015) Widespread polycistronic transcripts in fungi revealed by single-molecule mRNA sequencing. PLoS One 10:e0132628. https://doi.org/10.1371/journal.pone.0132628
Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, Couger MB, Eccles D, Li B, Lieber M, MacManes MD, Ott M, Orvis J, Pochet N, Strozzi F, Weeks N, Westerman R, William T, Dewey CN, Henschel R, LeDuc RD, Friedman N, Regev A (2013) De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8:1494–1512. https://doi.org/10.1038/nprot.2013.084
Hall DE, Zerbe P, Jancsik S, Quesada AL, Dullat H, Madilao LL, Yuen M, Bohlmann J (2013) Evolution of conifer diterpene synthases: diterpene resin acid biosynthesis in lodgepole pine and jack pine involves monofunctional and bifunctional diterpene synthases. Plant Physiol 161:600–616. https://doi.org/10.1104/pp.112.208546
Hebda RJ, Mathewes RW (1984) Holocene history of cedar and native Indian cultures of the North American Pacific Coast. Science 225:711–713. https://doi.org/10.1126/science.225.4663.711
Hu X-G, Liu H, Jin Y, Sun YQ, Li Y, Zhao W, el-Kassaby YA, Wang XR, Mao JF (2016) De novo transcriptome assembly and characterization for the widespread and stress-tolerant conifer Platycladus orientalis. PLoS One 11:e0148985. https://doi.org/10.1371/journal.pone.0148985
Jones P, Binns D, Chang HY, Fraser M, Li W, McAnulla C, McWilliam H, Maslen J, Mitchell A, Nuka G, Pesseat S, Quinn AF, Sangrador-Vegas A, Scheremetjew M, Yong SY, Lopez R, Hunter S (2014) InterProScan 5: genome-scale protein function classification. Bioinformatics 30:1236–1240. https://doi.org/10.1093/bioinformatics/btu031
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Keeling CI, Bohlmann J (2006a) Diterpene resin acids in conifers. Phytochemistry 67:2415–2423
Keeling CI, Bohlmann J (2006b) Genes, enzymes and chemicals of terpenoid diversity in the constitutive and induced defence of conifers against insects and pathogens. New Phytol 170:657–675
Keeling CI, Weisshaar S, Ralph SG, Jancsik S, Hamberger B, Dullat HK, Bohlmann J (2011) Transcriptome mining, functional characterization, and phylogeny of a large terpene synthase gene family in spruce (Picea spp.). BMC Plant Biol 11:43. https://doi.org/10.1186/1471-2229-11-43
Kotera E, Tasaka M, Shikanai T (2005) A pentatricopeptide repeat protein is essential for RNA editing in chloroplasts. Nature 433:326–330. https://doi.org/10.1038/nature03229
Lamesch P, Berardini TZ, Li D, Swarbreck D, Wilks C, Sasidharan R, Muller R, Dreher K, Alexander DL, Garcia-Hernandez M, Karthikeyan AS, Lee CH, Nelson WD, Ploetz L, Singh S, Wensel A, Huala E (2012) The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40:D1202–D1210. https://doi.org/10.1093/nar/gkr1090
Letunic I, Bork P (2016) Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw290
Lurin C, Andrés C, Aubourg S, Bellaoui M, Bitton F, Bruyère C, Caboche M, Debast C, Gualberto J, Hoffmann B, Lecharny A, le Ret M, Martin-Magniette ML, Mireau H, Peeters N, Renou JP, Szurek B, Taconnat L, Small I (2004) Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 16:2089–2103. https://doi.org/10.1105/tpc.104.022236
Martin DM, Fäldt J, Bohlmann J (2004) Functional characterization of nine Norway spruce TPS genes and evolution of gymnosperm terpene synthases of the TPS-d subfamily. Plant Physiol 135:1908–1927. https://doi.org/10.1104/pp.104.042028
Martin JA, Wang Z (2011) Next-generation transcriptome assembly. Nat Rev Genet 12:671–682. https://doi.org/10.1038/nrg3068
Miller JR, Koren S, Sutton G (2010) Assembly algorithms for next-generation sequencing data. Genomics 95:315–327
Morris PI, Stirling R (2012) Western red cedar extractives associated with durability in ground contact. Wood Sci Technol 46:991–1002. https://doi.org/10.1007/s00226-011-0459-2
Nakasugi K, Crowhurst R, Bally J, Waterhouse P (2014) Combining transcriptome assemblies from multiple de novo assemblers in the allo-tetraploid plant Nicotiana benthamiana. PLoS One 9:e91776. https://doi.org/10.1371/journal.pone.0091776
Neale DB, Wegrzyn JL, Stevens KA, Zimin AV, Puiu D, Crepeau MW, Cardeno C, Koriabine M, Holtz-Morris AE, Liechty JD, Martínez-García PJ, Vasquez-Gross HA, Lin BY, Zieve JJ, Dougherty WM, Fuentes-Soriano S, Wu LS, Gilbert D, Marçais G, Roberts M, Holt C, Yandell M, Davis JM, Smith KE, Dean JFD, Lorenz W, Whetten RW, Sederoff R, Wheeler N, McGuire PE, Main D, Loopstra CA, Mockaitis K, deJong PJ, Yorke JA, Salzberg SL, Langley CH (2014) Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies. Genome Biol 15:R59. https://doi.org/10.1186/gb-2014-15-3-r59
Nelson DR (2009) The cytochrome p450 homepage. Hum Genomics 4:59–65. https://doi.org/10.1186/1479-7364-4-1-59
O’Connell LM, Ritland K (2005) Post-pollination mechanisms promoting outcrossing in a self-fertile conifer, Thuja plicata (Cupressaceae). Can J Bot Can Bot 83:335–342. https://doi.org/10.1139/b05-007
Okamoto S, Yu F, Harada H, Okajima T, Hattan JI, Misawa N, Utsumi R (2011) A short-chain dehydrogenase involved in terpene metabolism from Zingiber zerumbet. FEBS J 278:2892–2900. https://doi.org/10.1111/j.1742-4658.2011.08211.x
Orsini L, Gilbert D, Podicheti R, Jansen M, Brown JB, Solari OS, Spanier KI, Colbourne JK, Rush D, Decaestecker E, Asselman J, de Schamphelaere KAC, Ebert D, Haag CR, Kvist J, Laforsch C, Petrusek A, Beckerman AP, Little TJ, Chaturvedi A, Pfrender ME, de Meester L, Frilander MJ (2016) Daphnia magna transcriptome by RNA-Seq across 12 environmental stressors. Sci Data 3:160030. https://doi.org/10.1038/sdata.2016.30
Proost S, Van BM, Vaneechoutte D et al (2015) PLAZA 3.0: an access point for plant comparative genomics. Nucleic Acids Res 43:D974–D981. https://doi.org/10.1093/nar/gku986
Rigault P, Boyle B, Lepage P, Cooke JEK, Bousquet J, MacKay JJ (2011) A white spruce gene catalog for conifer genome analyses. Plant Physiol 157:14–28. https://doi.org/10.1104/pp.111.179663
Ringer KL, Davis EM, Croteau R (2005) Monoterpene metabolism. Cloning, expression, and characterization of (-)-isopiperitenol/(-)-carveol dehydrogenase of peppermint and spearmint. Plant Physiol 137:863–872. https://doi.org/10.1104/pp.104.053298
Robertson G, Schein J, Chiu R, Corbett R, Field M, Jackman SD, Mungall K, Lee S, Okada HM, Qian JQ, Griffith M, Raymond A, Thiessen N, Cezard T, Butterfield YS, Newsome R, Chan SK, She R, Varhol R, Kamoh B, Prabhu AL, Tam A, Zhao YJ, Moore RA, Hirst M, Marra MA, Jones SJM, Hoodless PA, Birol I (2010) De novo assembly and analysis of RNA-seq data. Nat Methods 7:909–912. https://doi.org/10.1038/nmeth.1517
Russell JH, Burdon RD, Yanchuk AD (2003) Inbreeding depression and variance structures for height and adaptation in self- and outcross Thuja plicata families in varying environments. For Genet 10:171–184
Russell JH, Ferguson DC (2008) Preliminary results from five generations of a western redcedar (Thuja plicata) selection study with self-mating. Tree Genet Genomes 4:509–518. https://doi.org/10.1007/s11295-007-0127-8
Russell JH, Yanchuk AD (2012) Breeding for growth improvement and resistance to multiple pests in Thuja plicata. Gen Tech Rep 240:40–44
Schuler MA, Werck-Reichhart D (2003) Functional genomics of P450s. Annu Rev Plant Biol 54:629–667. https://doi.org/10.1146/annurev.arplant.54.031902.134840
Schulz MH, Zerbino DR, Vingron M, Birney E (2012) Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28:1086–1092. https://doi.org/10.1093/bioinformatics/bts094
Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31:3210–3212. https://doi.org/10.1093/bioinformatics/btv351
Smith-Unna R, Boursnell C, Patro R, Hibberd JM, Kelly S (2016) TransRate: reference-free quality assessment of de novo transcriptome assemblies. Genome Res 26:1134–1144. https://doi.org/10.1101/gr.196469.115
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stevens KA, Wegrzyn JL, Zimin A, Puiu D, Crepeau M, Cardeno C, Paul R, Gonzalez-Ibeas D, Koriabine M, Holtz-Morris AE, Martínez-García PJ, Sezen UU, Marçais G, Jermstad K, McGuire PE, Loopstra CA, Davis JM, Eckert A, de Jong P, Yorke JA, Salzberg SL, Neale DB, Langley CH (2016) Sequence of the sugar pine megagenome. Genetics 204:1613–1626. https://doi.org/10.1534/genetics.116.193227
Tatusov RL, Koonin EV, Lipman DJ (1997) A genomic perspective on protein families. Science 278:631–637. https://doi.org/10.1126/science.278.5338.631
Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar SK, Troggio M, Pruss D, Salvi S, Pindo M, Baldi P, Castelletti S, Cavaiuolo M, Coppola G, Costa F, Cova V, Dal Ri A, Goremykin V, Komjanc M, Longhi S, Magnago P, Malacarne G, Malnoy M, Micheletti D, Moretto M, Perazzolli M, Si-Ammour A, Vezzulli S, Zini E, Eldredge G, Fitzgerald LM, Gutin N, Lanchbury J, Macalma T, Mitchell JT, Reid J, Wardell B, Kodira C, Chen Z, Desany B, Niazi F, Palmer M, Koepke T, Jiwan D, Schaeffer S, Krishnan V, Wu C, Chu VT, King ST, Vick J, Tao Q, Mraz A, Stormo A, Stormo K, Bogden R, Ederle D, Stella A, Vecchietti A, Kater MM, Masiero S, Lasserre P, Lespinasse Y, Allan AC, Bus V, Chagné D, Crowhurst RN, Gleave AP, Lavezzo E, Fawcett JA, Proost S, Rouzé P, Sterck L, Toppo S, Lazzari B, Hellens RP, Durel CE, Gutin A, Bumgarner RE, Gardiner SE, Skolnick M, Egholm M, van de Peer Y, Salamini F, Viola R (2010) The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42:833–839. https://doi.org/10.1038/ng.654
Visser EA, Wegrzyn JL, Steenkmap ET, Myburg AA, Naidoo S (2015) Combined de novo and genome guided assembly and annotation of the Pinus patula juvenile shoot transcriptome. BMC Genomics 16:1057. https://doi.org/10.1186/s12864-015-2277-7
Vourc’h G, De Garine-Wichatitsky M, Labbé A et al (2002) Monoterpene effect on feeding choice by deer. J Chem Ecol 28:2411–2427. https://doi.org/10.1023/A:1021423816695
Warren RL, Keeling CI, Saint YMM et al (2015) Improved white spruce (Picea glauca) genome assemblies and annotation of large gene families of conifer terpenoid and phenolic defense metabolism. Plant J 83:189–212. https://doi.org/10.1111/tpj.12886
Wegrzyn JL, Liechty JD, Stevens KA, Wu LS, Loopstra CA, Vasquez-Gross HA, Dougherty WM, Lin BY, Zieve JJ, Martinez-Garcia PJ, Holt C, Yandell M, Zimin AV, Yorke JA, Crepeau MW, Puiu D, Salzberg SL, de Jong PJ, Mockaitis K, Main D, Langley CH, Neale DB (2014) Unique features of the loblolly pine (Pinus taeda L.) megagenome revealed through sequence annotation. Genetics 196:891–909. https://doi.org/10.1534/genetics.113.159996
Xie Y, Wu G, Tang J, Luo R, Patterson J, Liu S, Huang W, He G, Gu S, Li S, Zhou X, Lam TW, Li Y, Xu X, Wong GKS, Wang J (2014) SOAPdenovo-Trans: de novo transcriptome assembly with short RNA-Seq reads. Bioinformatics 30:1660–1666. https://doi.org/10.1093/bioinformatics/btu077
Yagi Y, Tachikawa M, Noguchi H, Satoh S, Obokata J, Nakamura T (2013) Pentatricopeptide repeat proteins involved in plant organellar RNA editing. RNA Biol 10:1419–1425. https://doi.org/10.4161/rna.24908
Zerbe P, Hamberger B, Yuen MMS, Chiang A, Sandhu HK, Madilao LL, Nguyen A, Hamberger B, Bach SS, Bohlmann J (2013) Gene discovery of modular diterpene metabolism in nonmodel systems. Plant Physiol 162:1073–1091. https://doi.org/10.1104/pp.113.218347
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18:821–829. https://doi.org/10.1101/gr.074492.107
Zharkikh A, Troggio M, Pruss D, Cestaro A, Eldrdge G, Pindo M, Mitchell JT, Vezzulli S, Bhatnagar S, Fontana P, Viola R, Gutin A, Salamini F, Skolnick M, Velasco R (2008) Sequencing and assembly of highly heterozygous genome of Vitis vinifera L. cv Pinot Noir: problems and solutions. J Biotechnol 136:38–43. https://doi.org/10.1016/j.jbiotec.2008.04.013
Zimin AV, Stevens KA, Crepeau MW, Puiu D, Wegrzyn JL, Yorke JA, Langley CH, Neale DB, Salzberg SL (2017) An improved assembly of the loblolly pine mega-genome using long-read single-molecule sequencing. Gigascience 6:1–4. https://doi.org/10.1093/GIGASCIENCE/GIW016
Zulak KG, Bohlmann J (2010) Terpenoid biosynthesis and specialized vascular cells of conifer defense. J Integr Plant Biol 52:86–97
Acknowledgements
We thank Dr. Carol Ritland and Ms. Karen Reid for excellent project management support, Dr. Timothy J. Sexton for technical assistance, and the McGill University and Génome Québec Innovation Centre for sequencing services. The research was supported with funds from the Natural Sciences and Engineering Research Council of Canada (NSERC Discovery Grant) and funds to JB and JHR from Genome British Columbia, Genome Canada, and the British Columbia Ministry of Forests, Lands, Natural Resource Operations and Rural Development (MFLNRORD) for the CEDaR User Partnership Project (UPP-002, Genome BC) and the CEDaR Applied Genomics Partnership Project (184CED-GAPP, Genome Canada and Genome BC). TJS is supported by a NSERC Postgraduate Doctoral fellowship.
Data archiving statement
The sequence data supporting this work can be found at the NCBI BioProject Database under BioProject ID PRJNA399722. In addition, sequences of the gene lists described in this paper and their annotations are also available in Files S1–S5 and File S9.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Dardick
Electronic supplementary material
Figure S1
Representative four-month old WRC seedling used for RNA isolation and sequencing. (JPEG 52 kb)
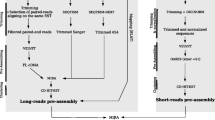
Figure S2
Pipeline for the de novo assembly and redundancy reduction of the WRC gene set, carried out for each inbred S5 line separately. (JPEG 208 kb)
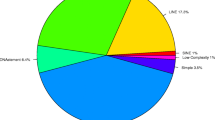
Figure S3
Monoterpene profiles of foliar samples for 12 different monoterpenes across the eight S5 lines. (JPEG 937 kb)
Figure S4
Results of the BUSCO gene set completeness assessment. The reduced-redundancy gene set for WRC S5 Line 4 was found to be the most complete, with the lowest number of missing orthologs. (JPEG 438 kb)
Table S1
Summary for transcriptome assemblies for WRC S5 lines. (DOCX 14 kb)
Table S2
Results of the Conditional Reciprocal Best BLAST (CRBB) analysis. (DOCX 15 kb)
Table S3
Results of the BLASTp analysis of transcriptome assemblies against the longest predicted proteins (n = 1000) in the P. glauca and A. thaliana reference gene sets. (DOCX 13 kb)
File S1
Sequences of 241 plant terpene synthase (TPS) used in construction of a maximum-likelihood phylogeny of plant TPS. (TXT 184 kb)
File S2
Sequences of 126 gymnosperm and a single P. patens TPS used in construction of a maximum-likelihood phylogeny of gymnosperm TPS. (TXT 101 kb)
File S3
Sequence data for the core WRC gene set. Gene set containing the 28,279 core, reduced-redundancy protein sequences for predicted ORFs as produced by the EvidentialGene pipeline. (TXT 12858 kb)
File S4
Sequence data for the alternate WRC gene set. Gene set containing 40,691 additional putative protein-coding sequences, which may be potential gene isoforms or paralogs. (TXT 18875 kb)
File S5
Summary of significant BLASTp and InterProScan hits for the main reduced-redundancy gene set of Line 4. BLAST columns are as described in the BLAST Command Line Applications User Manual (https://www.ncbi.nlm.nih.gov/books/NBK279690/). The pipeline for BLASTing and filtering hits is described in the Methods section. GO names are separated by Biological Process (P), Molecular Function (F) and Cellular Component (C). The InterPro ID column lists all InterPro domains found for the queried sequence. Top PFAM hit describes the hit with the highest score against the PFAM database for each sequence, using an e-value cut-off of 1e-5. (XLSX 5321 kb)
File S6
Statistical summary of orthogroup analysis for all sequences assigned to orthogroups. Of the 498,235 protein coding sequences from 16 different plant species submitted for orthogroup analysis, 391,179 were successfully assigned to 19,660 orthogroups. The majority of orthogroups (11,616) had an average of less than one gene per species; the largest orthogroup (3201 genes) had an average of 151–200 genes per species. A large number of orthogroups (5614) had members from only two species; however, a similarly large number (3835) had members from all 16 species. (XLSX 15 kb)
File S7
Statistical summary of orthogroup analysis results for each species. The species with the lowest amount of genes assigned to orthogroups was P. patens, with only 57.5% of sequences assigned; the highest was P. glauca with 92.4%. 90.4% of our WRC gene set was successfully assigned to orthogroups; 0.1% of WRC sequences were in species-specific orthogroups. (XLSX 21 kb)
File S8
Summary of orthogroup composition and function. The number of orthogroup members from each species, together with the total number of genes in each orthogroup and the top five PFAM hits for each orthogroup. The largest orthogroup, with 3201 genes consisted mainly of pentatricopetide-repeat containing protein-coding genes, a large protein family in plants with little functional redundancy (Lurin et al. 2004). (XLSX 1665 kb)
File S9
Sequence data for 33 putative full-length TPS genes from the WRC gene set. Putative TPS were identified using BLASTp, InterProScan and orthogroup analysis, and after removal of partial ORFs and proteins less than 400 aa long were reduced to a set of 33 putative full-length TPS. (TXT 26 kb)
Rights and permissions
About this article
Cite this article
Shalev, T.J., Yuen, M.M.S., Gesell, A. et al. An annotated transcriptome of highly inbred Thuja plicata (Cupressaceae) and its utility for gene discovery of terpenoid biosynthesis and conifer defense. Tree Genetics & Genomes 14, 35 (2018). https://doi.org/10.1007/s11295-018-1248-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-018-1248-y