Abstract
Unexpectedly high levels of field susceptibility to the fusiform rust fungus observed for slashXloblolly hybrid families in the Cooperative Forest Genetics Research Program Pine Hybrid Trials led to several hypotheses concerning causation of the observed susceptibility. One of these hypotheses involved the failure of major resistance genes to appropriately function in this particular hybrid combination. This current work, involving the loblolly pine resistance gene Fr1 and a fusiform rust inoculum avirulent against Fr1 resistance in a greenhouse trial, delineates the investigation of major gene resistance for a particular slashXloblolly hybrid family. In this trial, the Fr1 resistance allele, derived from the heterozygous (Fr1/fr1) loblolly male parent and transferred to hybrid offspring that should have been segregating 1:1 for resistance, was fully penetrant. Likewise, in the pure-species loblolly control, the Fr1 resistance allele was again fully penetrant, and the performances of our hybrid family and the loblolly control family (both of which segregated for Fr1 resistance) were comparable. These results by inductive reasoning refute the hypothesis that major resistance genes are not appropriately functional in a slashXloblolly hybrid background.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 1994, the Cooperative Forest Genetics Research Program (CFGRP) installed a series of 11 pine hybrid trials in the lower coastal plain of the southeastern USA that included the slash pine (Pinus elliottii var. elliottii) as the maternal parent in crosses (numbers of families consistent across all sites given) with Caribbean pines (Pinus caribaea var. bahamensis, 23 families and P. caribaea var. hondurensis, 23 families), loblolly pine (Pinus taeda, 19 families), and an F1 family from an Australian tree improvement program (P. elliottii var. elliottii X P. caribaea var. hondurensis, 20 families). All pollen sources, except for the F1 family from Australia, were mixes of about 20 pollen parents. Also included in the trials were open-pollinated families from the same slash maternal (18 families) and loblolly pine pollen parents (18 families) used in the crossing (Lopez-Upton et al. 1999, 2000; Gezan et al. 2004). These tests were assessed for growth, tip moth (Rhyacionia spp.) incidence, and fusiform rust disease incidence.
Genetic variation for resistance to the fusiform rust fungus Cronartium quercuum Berk. Miyabe ex Shirai f. sp. fusiforme (subsequently called Cqf) is well known in southern pines (Barber et al. 1957; Jewell and Mallett 1967; Wells et al. 1982; Kuhlman and Powers 1988; Snow et al. 1990; Hodge et al. 1993; McKeand et al. 1999; Isik et al. 2008), as is pathogenic variation in Cqf (Snow and Kais 1970; Powers 1980; Kuhlman and Matthews 1993). Further experimentation has shown specific host–pathogen interactions (Griggs and Walkinshaw 1982; Kinloch and Walkinshaw 1991; Nelson et al. 1993), leading to the hypothesis that gene-for-gene interactions are important factors in this pathosystem. Subsequent genetic mapping experiments revealed specific resistance genes (Fr genes) in the host (Wilcox et al. 1996; Amerson et al. 1997, 2005; Jordan 1997; Li 2003), while recent experiments with the pathogen led to the demonstration (Kubisiak et al. 2005) and mapping of a functional avirulence gene (Avr1) in the pathogen, with this mapping effort conclusively demonstrating the gene-for-gene interaction of the host Fr1 resistance gene and the pathogen Avr1 gene (Kubisiak et al. 2010). These findings collectively have led to a general acceptance that gene-for-gene interactions indeed play a large role in the determination of fusiform rust disease development (i.e., gall formation). Recently, Nelson et al. (2010) presented the experimental approach and data analysis that underlie this gene-for-gene conclusion and delineated the practical implications for optimal deployment of southern pine genotypes for fusiform rust resistance given the virulence allele frequencies in the local Cqf populations.
Results from the CFGRP hybrid trials for growth and fusiform rust resistance have been reported at age 3 years by Lopez-Upton et al. (1999) and at age 8 years by Gezan et al. (2004). A most interesting result was seen for fusiform rust incidence (scored as presence or absence of galls) in the maternal slashXloblolly pine hybrid families (F1SL) compared to the rust incidence for the pure slash pine and loblolly pine open-pollinated families. At age 3 years across 11 trials, the mean of the F1SL families was significantly higher than the mean between the two pure species, and by age 8 years, the mean rust incidence across all 11 trials and across all F1SL families was 42.6%, while the mean rust incidences across the trials for the open-pollinated slash and loblolly pine families were 36.9% and 27.9%, respectively. The rust incidence mean of the F1SL families was not only significantly greater than the mean of the two sets of pure-species open-pollinated families (approximate test for heterosis), but was also significantly greater than the mean for the poorer performing species. Furthermore, no open-pollinated pure-species family had a mean incidence across the 11 trials equal to or greater than the mean for F1SL families, and the mean incidence for each pure-species slash maternal family was always less than the comparable F1SL family. These data appear contrary to the expectation that the F1SL families, on average, should have displayed resistance levels at least comparable to those of the OP parental families, on average.
Given these observations regarding hybrid susceptibility, we hypothesized that major resistance genes (i.e., Fr genes), as previously recognized in the loblolly pine, are not appropriately functional in F1SL hybrid backgrounds (i.e., that major resistance genes in the pure species and in hybrids may not function in the same manner). To test a specific case of this hypothesis, we sought to determine if disease phenotype expression, conferred by the heterozygous Fr1 resistance gene discovered in loblolly pine selection 10-005 and tractable (based on allele inheritance) in its pure-species progeny with genetic markers (Wilcox et al. 1996), would differ in pure-species loblolly 10-005 progeny and progeny of a slash (highly disease susceptible parent) x loblolly 10-005 cross when both progeny sets were challenged with a single genotype Cqf isolate that was avirulent to Fr1. In such a challenge with pure-species loblolly 10-005 progeny where the other parent was highly susceptible, the expectation is that most or all of the progeny deemed (based on genetic markers) to have the Fr1 resistance allele should be non-diseased (gall free), while most or all of the progeny deemed to have the fr1 non-resistance allele should be diseased (galled). In our comparison of pure-species loblolly 10-005 progeny and slash x 10-005 progeny, we used random amplification of polymorphic DNA (RAPD) markers (genetic markers more fully described in the “Materials and Methods” section) linked to the Fr1 gene in the loblolly 10-005 parent for determination of the Fr1 vs. fr1 allele status of a given progeny.
Materials and methods
Plant materials
Hybrid seeds used for this study were made by crossing rust-susceptible slash pine 003-01-053, as the female, with pollen derived from loblolly pine 10-005 which is heterozygous for fusiform rust resistance gene Fr1. Loblolly pine seeds used as a control/check were obtained from a full-sib cross of loblolly tree 10-005, as the female, with pollen from a highly rust-susceptible loblolly individual, USDA Forest Service tree 4666-4 (Kuhlman 1992; Wilcox et al. 1996; Kuhlman et al. 1997) that lacks Fr1 resistance (Wilcox et al. 1996; Kuhlman et al. 1997).
Genetic marker background and preliminary studies
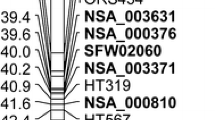
In loblolly pine selection, 10-005 RAPD markers J7_470 (previously called J7_485; Wilcox et al. 1996) and AJ4_420 are tightly linked to each other at a distance of 1.3 cm (H. Amerson, unpublished data) and also tightly linked to the Fr1 gene (Wilcox et al. 1996; H. Amerson, unpublished data), with Fr1 most likely residing in the interval between these markers (H. Amerson, unpublished data), but precise positioning of Fr1 awaits further study.
In tree 10-005, the + (band present) allele of marker J7_470 and the − (band absent) allele of marker AJ4_420 are coupled with each other and the Fr1 resistance allele, while −J7_470 and +AJ4_420 are coupled with each other and the fr1 non-resistance allele (H. Amerson, unpublished data, cited in Myburg et al. 2006). Since RAPD markers are dominant markers, it was necessary that one and, preferably, both markers (J7_470, AJ4_420) be homozygous null (uniformly band absent in the slash female) if one or both of the markers were to be successfully used to make allele calls (Fr1 vs. fr1 inherited from the loblolly parent) in the hybrid progeny. Prior to the start of our inoculation study (described below), preliminary marker investigations of haploid megagametophyte DNA from the hybrid seeds (the megagametophytic nuclear DNA in a pine seed is equivalent to the maternal contribution to the nuclear DNA of the embryo) and diploid needle DNA from the hybrid progeny revealed that the slash parent was indeed homozygous null for both markers J7_470 and AJ4_420. Hence, both markers were suitable for assessing the loblolly10-005 allele contribution (Fr1 vs. fr1) in each hybrid progeny as well as to the pure-species loblolly progeny.
Host materials, fungal materials, and inoculations
For this study, the 120 F1SL and 120 loblolly progenies (controls) were inoculated with basidiospores derived from the Cqf single genotype isolate 0-5-32 (avirulent against Fr1 resistance). The progeny were assessed for rust disease status and evaluated for allele status (Fr1 vs. fr1), determined by genetic markers. Loblolly control and F1SL hybrid and seeds were sown in vermiculite and greenhouse-germinated. Subsequent to germination, germlings/seedlings were individually transplanted into Ray Leach super cells containing a 3:4:3 peat/perlite/vermiculite potting mixture and numbered, following the standard protocol of the USDA Forest Service Resistance Screening Center (RSC) in Asheville, NC (Knighten et al. 1988). The F1SL progeny were produced and maintained at the RSC. The loblolly controls were initially produced and maintained at the North Carolina State University in Raleigh, NC, to facilitate megagametophyte harvest shortly after the time of transplant, and transferred to the Asheville facility at ~5 weeks postsowing. Inoculum production and progeny inoculation at 8 weeks postsowing followed standard RSC protocols (Knighten et al. 1988), except that inoculum concentration was raised to 100,000 basidiospores/ml. Final seedling disease assessments were conducted at 7 months postinoculation. For each living seedling, the disease phenotype was scored as either diseased (gall present), non-diseased (no gall), or in a few cases, ambiguous (gall status not clear). Four F1SL hybrid seedlings and one loblolly seedling died before phenotype assessment. These five individuals and two loblolly seedlings with ambiguous phenotypes were deleted from the marker–phenotype analyses.
DNA isolation and RAPD marker protocols
The haploid megagametophyte was harvested from each individual loblolly seedling shortly after transplantation (while the megagametophyte was still attached to the cotyledons) and maintained individually frozen at −20 to −80°C. Diploid needle tissue was collected from each F1SL seedling and maintained individually frozen at −20 to −80°C. Numbered identities for each individual sample (megagametophyte or needles) and its corresponding seedling were maintained throughout the entire experiment. To begin DNA isolation, each previously frozen megagametophyte or needle sample (~50 mg or less) was positioned between a ¼-inch ceramic ball (bottom) and a ¼-inch ceramic cylinder (Qbiogene; http//www.qubiogene.com) in individual 2-ml screw cap microcentrifuge tubes to facilitate specimen grinding. A lysis mixture warmed to ~65°C, either Puregene lysis solution plus Proteinase K for megagametophytes or Qiagen lysis buffer AP1 plus RNase A for needles, was added to each tube. Grinding (megagametophytes, 25 s at 4.5 speed; needles, 30 s at 5.0 speed) was performed in a Bio-101 Thermo Savant grinder (Carlsbad, CA), and genomic DNA was isolated from each megagametophyte sample following a modified Puregene protocol (PUREGENE DNA Isolation Kit, Gentra Systems; Minneapolis, MN; Kubisiak et al. 2010), with the final isolated DNA rehydrated in 100-ul of low TE (10 mM Tris–HCl, pH 8.0 and 0.1 mM EDTA, pH 8.0). Genomic DNA was isolated from each ground needle sample following a modified Qiagen DNeasy 96 protocol (Qiagen DNeasy 96 Plant Kit Handbook, Qiagen Inc.; Valencia, CA) similar to that reported by Li (2003), but amended to employ a porous/breathable tape seal (included in the Qiagen kit) atop the DNeasy 96 plate for all centrifugations of the DNeasy 96 plate, and by elution of final isolated DNA in 200 ul of Qiagen AE solution. For our isolation protocol, high speed centrifugation of samples in a 96-well plate format was accomplished in either a Qiagen–Sigma 4–15 or 4K15 centrifuge at 6,000 rpm which equates to 5,635 and 5,796×g, respectively. DNA concentration for each sample was estimated by visual comparison with known lambda standards following electrophoresis in 0.8% agarose gels. Haploid DNA (from megagametophytes) was diluted to 1.0 ng/μl, and diploid DNA (from needles) was diluted to 1.5 ng/μl for use in RAPD reactions. Haploid and diploid samples were treated the same except that RAPD reactions for haploid samples utilized 5 ng of template DNA, while those for diploid samples utilized 7.5 ng of template DNA. RAPD reactions (with noted haploid vs. diploid template differences), PCR conditions/specifications, gel electrophoresis, DNA band visualization, and recording (for our study markers) all followed the protocol of Myburg et al. (2006). Markers J7_470 and AJ4_420 were scored as band present vs. absent by visual assessment. Samples having the +J7_470 (band present) and the −AJ4_420 (band absent) profiles were deemed to have the Fr1 resistance allele inherited from parent 10-005, while samples having the −J7_470/+AJ4_420 profile were deemed to have the fr1 non-resistance allele inherited from parent 10-005. Two of the hybrid samples showing recombination between markers J7_470 and AJ4_420 were deleted from the marker–phenotype analysis, as the allele call was considered confounded.
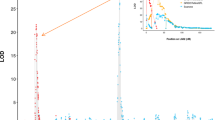
Taking into account the previously noted seven seedlings for which phenotype data were missing or ambiguous and the two DNA samples showing marker recombination, the 120-member hybrid sample set was reduced to 114 members for marker–phenotype analysis, while the 120-member loblolly sample set was reduced to 117 members for marker–phenotype analysis. Marker–phenotype logarithm of odds (LOD) associations, indicative of allele (Fr1 or fr1)–phenotype associations, were evaluated using Map Maker Macintosh 2.0 software (Lander et al. 1987). P values presented from Map Maker are likelihood of the odds tests (a form of likelihood ratio test) of the complete versus reduced model for inclusion or exclusion of the marker.
Results and discussion
Chi square (χ 2) assessments at the 0.05 level for marker segregation in both the loblolly (including markers from the 3 deleted samples) and the hybrid progeny (including markers which were available for 4 of the 6 deleted samples) revealed that segregation did not significantly deviate from 1:1. Because samples (2 F1SL hybrids) showing recombination between markers were excluded from the marker–phenotype analysis, markers J7_470 and AJ4_420 furnished identical information, but the small number of recombinants observed for the F1SL hybrid samples, as well as the 1:1 segregation, indicated that the F1SL marker data were not atypical of those expected for loblolly family 10-005.
The pure-species loblolly pine control family that was composed of 117 10-005 maternal full-sibs performed as expected from previous experimentation (Table 1). All individuals with the marker profile indicative of the Fr1 allele were non-galled (hence, the Fr1 resistance allele was fully penetrant), and 92% of the individuals with the alternative marker profile, indicative of the fr1 allele, had galls. Marker–phenotype LOD associations were 26.25 (p value < 10−5).
The results from the association of disease phenotypes (gall vs. no gall) with marker states for the 114 F1SL individuals were extremely compelling (Table 2), with LOD scores of 21.74 (p value < 10−5). No individuals carrying the marker profile indicative of the Fr1 allele were galled, while those with the marker profile indicative of the fr1 allele were galled at 84%. Again, the only exceptions to marker profile expectations were the fr1 non-galled individuals (Table 2). The failure of some supposedly susceptible individuals to develop galls (these being termed “escapes”) is a common phenomenon observed in pine-Cqf inoculation trials (Nelson et al. 2010; Kayihan et al. 2010), and a heritable component for escape rate was observed in clonal materials of 63 full-sib loblolly pine families challenged with Cqf (Kayihan et al. 2010).
This greenhouse experiment, using seedlings challenged with basidiospores derived from a Cqf single genotype isolate avirulent against Fr1 resistance, has clearly demonstrated that the Fr1 resistance allele (inherited from the loblolly 10-005 pollen parent) was fully penetrant/functional in the F1SL study family where the maternal slash parent was chosen only for susceptibility to Cqf. Since there is a common theoretical mode of action for other Fr genes, there would be no reason to speculate that other Fr genes would not also function as well in F1SL families. Furthermore, Fr1 resistance has been shown to extend to 7-year-old (Wilcox et al. 1996) and 3-year-old (Kayihan et al. 2010) loblolly pine field plantings. From these facts, the major conclusion to be drawn is that the poor Cqf resistance, demonstrated by F1SL families in the CFGRP Pine Hybrid Trials, most probably was not due to the inability of the Fr genes to function.
The cause or causes of higher-than-expected fusiform rust incidence (high Cqf susceptibility) in F1SL families previously tested (CFGRP tests) in the field remain elusive. Certainly, there is a possibility of differences in growth phenology between slashXloblolly hybrids and pure-species families (Munoz et al. 2010). A difference in growth phenology, where flushing at different times may occur, could cause exposure to different pathogen genotypes and possibly different (perhaps higher) inoculum levels. Also, if hybrid shoots failed to develop woodiness as quickly as the pure species, that likely would result in an extended period of succulence and susceptibility. The tests of these and other hypotheses related to the susceptibility phenomenon await further experimentation.
References
Amerson HV, Jordan AP, Kuhlman EG, O'Malley DM, Sederoff RR (1997) Genetic basis of fusiform rust disease resistance in loblolly pine. In: Proceedings of the 24th biennial southern forest tree improvement conference, Orlando, FL, June 9–12, 1997 403 pp
Amerson HV, Kubisiak TL, Garcia SA, Kuhlman EG, Nelson CD, McKeand SE, Mullin TJ, Li B (2005) Interacting genes in the pine-fusiform rust forest pathosystem. In: Proceedings of the 28th biennial southern forest tree improvement conference, June 21–23, 2005, Raleigh, NC, US Department of Agriculture, Forest Service, p. 60. At http://wwwtreesearchfsfedus/pubs/25641
Barber JC, Dorman KW, Bauer E (1957) Slash pine progeny tests indicate genetic variation in resistance to rust. US Dep Agric For Serv Southeast For Exp Stn Res Note 104
Gezan SA, Huber DA, Medina AM, Parisi L, Powell GL (2004) 46th Annual progress report, cooperative forest genetics research program, University of Florida pp 12–14
Griggs MM, Walkinshaw CH (1982) Diallel analysis of genetic resistance to Cronartium quercuum fsp fusiforme in slash pine. Phytopathology 72:816–818
Hodge GR, White TL, Schmidt RA, Allen JE (1993) Stability of rust infection ratios for resistant and susceptible slash and loblolly pine across rust hazard levels. S Jour Appl For 17(4):188–192
Isik F, Amerson HV, Whetten RW, Garcia SA, Li B, McKeand S (2008) Resistance assessments of elite loblolly pine families to fusiform rust inocula in greenhouse testing. Can J For Res 38:2687–2696
Jewell FF, Mallett SL (1967) Testing slash pine for rust resistance. For Sci 13:413–418
Jordan AP (1997) Fusiform rust disease resistance and genomic mapping in loblolly pine. MS thesis, NC State Univ Raleigh, NC p 105
Kayihan GC, Nelson CD, Huber DA, Amerson HV, White TL, Davis JM (2010) Clonal evaluation for fusiform rust disease resistance: effects of pathogen virulence and disease escape. Can J For Res 40:1042–1050
Kinloch BB, Jr, Walkinshaw CH (1991) Resistance to fusiform rust in southern pines: how is it inherited? In: Hiratsuka Y, Samoil JK, Blenis PV, Crane PE, Laishley BL (eds) Rusts of Pine Proc IUFRO Rusts of Pine Working Party Conf Inf rep NOR-X-317, Edmonton, Canada pp. 219–228
Knighten JL, Young CH, McCartney TC, Anderson RL (1988) Resistance screening center procedures manual: a step-by-step guide used in the operational screening of southern pines for resistance to fusiform rust. USDA Forest Service Forest Pest Management Report 83-1-18 (revised 1988) p. 62
Kubisiak TL, Amerson HV, Nelson CD (2005) Genetic interaction of the fusiform rust fungus with resistance gene Fr1 in loblolly pine. Phytopathology 95:376–380
Kubisiak TL, Anderson CL, Amerson HV, Smith JA, Davis JM, Nelson CD (2010) A genomic map enriched for markers linked to Avr1 in Cronartium quercuum f. sp. fusiforme. Fungal Genet Biol (in press)
Kuhlman EG (1992) Interaction of virulent single-gall isolates of Cronartium quercuum fsp fusifiorme and resistant families of loblolly pine. For Sci 38:641–651
Kuhlman EG, Matthews FR (1993) Variation in virulence among single-aeciospore isolates from single-gall isolates of Cronartium quercuum fsp fusiforme. Can J For Res 23:67–71
Kuhlman EG, Powers HR Jr (1988) Resistance responses in half-sib loblolly pine progenies after inoculation with Cronartium quercuum fsp fusiforme. Phytopathology 78:484–487
Kuhlman EG, Amerson HV, Jordan AP, Pepper WD (1997) Inoculum density and expression of major gene resistance to fusiform rust disease in loblolly pine. Plant Dis 81:597–600
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Li H (2003) Major gene detection for fusiform rust resistance using segregation analysis and linkage analysis in loblolly pine. PhD thesis, NC State Univ, Raleigh, NC
Lopez-Upton J, White TL, Huber DA (1999) Taxon and family differences in survival, cold hardiness, early growth, and rust incidence of loblolly pine, slash pine and some pine hybrids. Silvae Genet 48:303–313
Lopez-Upton J, White TL, Huber DA (2000) Species differences in early growth and rust incidence of loblolly and slash pine. Forest Ecol Manag 132:211–222
McKeand SE, Li B, Amerson HV (1999) Genetic variation in fusiform rust resistance in loblolly pine across a wide geographic range. Silvae Genet 48:255–260
Munoz PR, Huber DA, Butnor JR (2010) Phenotypic analysis of first-year traits in a pseudo-backcross family (slash x loblolly) x slash and the open-pollinated families of the pure species progenitors. Tree Genet Genomes (in press)
Myburg H, Morse AM, Amerson HV, Kubisiak TL, Huber D, Osborne JA, Garcia SA, Nelson CD, Davis JM, Covert SF, van Zyl L (2006) Differential gene expression in loblolly pine (Pinus taeda L) challenged with the fusiform rust fungus, Cronartium quercuum fsp fusiforme. Physiol Mol Plant Pathol 68:79–91
Nelson CD, Doudrick RL, Nance WL, Hamaker JM, Capo B (1993) Specificity of host:pathogen genetic interaction for fusiform rust disease on slash pine. In: Proceedings of the 22nd southern forest tree improvement conference, June 14–17, 1993, Atlanta, Georgia, USA, pp 403–411
Nelson CD, Kubisiak TL, Amerson HV (2010) Unraveling and managing fusiform rust disease: a model approach for coevolved forest tree pathosystems. For Path 40:67–72
Powers HR Jr (1980) Pathogenic variation among single-aeciospore isolates of Cronatium quercuum f sp fusiforme. For Sci 26:280–282
Snow GA, Kais AG (1970) Pathogenic variability in isolates of Cronartium fusiforme from five southern states. Phytopathology 60:1730–1731
Snow GA, Matthews FR, Nance WL, Foster GS (1990) Effects of pollen source on loblolly pine resistance to Cronartium quercuum fsp fusiforme. For Sci 36:304–312
Wells OO, Switzer GL, Nance WL (1982) Genotype-environment interaction in rust resistance in Mississippi loblolly pine. For Sci 28:797–809
Wilcox PL, Amerson HV, Kuhlman EG, Liu BH, O'Malley DM, Sederoff RR (1996) Detection of a major gene for resistance to fusiform rust disease in loblolly pine by genomic mapping. Proc Natl Acad Sci USA 93:3859–3864
Acknowledgments
We appreciate and acknowledge the support of the NCSU College of Natural Resources, the NCSU Forest Biotechnology Laboratory, and the Cooperative Forest Genetics Research Program at the University of Florida. We would further like to acknowledge Plum Creek (Paul Belonger and Lorin Clark) for making the hybrid cross.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Dean
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Huber, D., Amerson, H. Performance of the loblolly pine fusiform rust disease resistance gene (Fr1) in a slashXloblolly pine hybrid family. Tree Genetics & Genomes 7, 535–540 (2011). https://doi.org/10.1007/s11295-010-0353-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-010-0353-3