Abstract
A healthy life is essential for a happy society, however it is a fact that seemingly invisible diseases plague our families and people suffer. The thyroid disease falls in such a category. Thyroid disorders are long-term and with carefully handled illnesses, people with thyroid disorders may also live stable and normal lives. Thyroid diagnosis, particularly for an inexperienced clinician, is a difficult proposal. Many researchers have established various methods for the diagnosis of the disease and several models for disease prediction have been developed. As with several other domains, machine learning approaches to modelling health care problems is gaining popularity. This study aims at providing solutions towards such a thyroid disease prediction. Dimension reduction techniques are applied, and reduced dimension data input to classifiers. Also, data augmentation is applied so as to be able to generate sufficient data for deep neural network model. Classifier prediction is compared to other similar researches. Real life dataset for thyroid disease has been used, and experiments conducted in distributed environment. Our proposed two stage approach gives a maximum accuracy of 99.95% which is very good as compared to existing techniques. We have shown that dimension reduction and data augmentation can be used very efficiently for achieving high accuracy of disease prediction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Machine learning has become an important part of human lives that provides smart and affordable solutions to various problems. As such, healthcare is catching the attention of many researchers, as society relies upon healthy and performing individuals for its balanced functioning. It is obvious that a diseased person would spend much of his time in fretting about his health, thus leaving very little productive time left to complete the assigned duties, let alone perform well. This is an uncalled for situation. For instance, a lady sitting down on her desk, trying to sort out a coding problem, feels agitated with throbbing pulse which wants to beat out of her heart. Or, another person, say an accountant, trying to complete the balance sheet for a client, feels feverish, and delirious. Clearly, people involved in both of these examples are not in a position to complete their tasks to the best of their capabilities. The reason being—they might be suffering from a thyroid disorder, called hyperthyroidism. Some may feel drowsy and lethargic, which is a case of hypothyroidism. The thyroid malfunction is one of the common diseases affecting people from all age groups. The disease is not dangerous as other diseases like heart disease and cancer, but it may be the cause of other diseases with severe complications.
To our rescue, some very dedicated researchers have been putting in the very best of their efforts in modelling these disease prediction problems using statistical techniques, and now machine learning and deep learning techniques. Data mining and machine learning techniques can be used to identify thyroid disease. This method both reduces misdiagnoses due to human mistakes and allows for efficient use of time. However, most data mining and machine learning approaches require marked training data. For greater accuracy, the volume of data is critical. There are other issues which researchers have to face while performing research related to health care data, such as authorization for collecting data, privacy and secrecy concerns etc. In spite of all this, researchers are motivated to explore in terms of analyzing the data, its exploratory analysis, preprocessing, dimension reduction, data augmentation, and so on. The aim of this research is to provide a modelling solution to the prediction of thyroid disease so that society can benefit from the research advancements of computational techniques. We have applied dimension reduction techniques, and the output of these techniques is input into two classifiers. The comparative analysis shows the efficacy of our approach. To make the size of data large enough for buiding a dep neural network model, we have applied data augmentation.
The main objectives of the present research work are as follows:
-
Preprocessing of data
-
Apply dimension reduction and data augmentation techniques
-
Build classifiers in a distributed environment
-
Perform comparative analysis
The rest of the paper is organized as follows: Sect. 2 presents the literature review. Section 3 presents the methodology. Section 4 describes the experiments, and Sect. 5 represents the results and exploratory analysis. Finally, Sect. 6 concludes the paper.
2 Literature Review
Researchers have been putting in efforts to integrate Information technology in healthcare not only in terms of applying machine learning techniques to healthcare data but also to devise techniques such as telemedicine, smart care taking platforms etc. In [1] Anwar et al. propose a model where telemedicine technology could be helpful where there is a shortage of medical specialists or doctors. In their paper titled “Child Temperature Monitoring System” [2], the authors provide a smart way to protect the infant from sudden infant death syndrome. This technique is a novel concept that will help parents and care taker to know their newborn better, especially because the infant is helpless in sharing. Anwar and Prasad in [3] claim that following a critical care plan is vital for chronic diseases. More so, if the patient has some disability. This can be achieved by integration of Information and Communication Technologies (ICT) and focal health care business models. This definitely is a step towards a better workforce. In [4] Koren et al. claim that sensor data is subject to several sources of faults and errors, which may further lead to imprecise or even incorrect and misleading answers. So data collected from wearable sensors need to be analyzed to confirm that they are correct and relevant. Only then this data can be included in a formal Electronic Health Record. The collection of health care data using sensors in large amounts has further motivated researchers to carry on these studies effectively using big data platforms [4,5,6,7,8,9]. MapReduce [10, 11], a distributable and scalable parallel processing framework, is used for data processing in healthcare. Deep learning approaches have been applied for the prediction of violent incidents by patients [12].
Now we discuss some research work specific to thyroid disease prediction. The thyroid gland produces hormones for the regulation of metabolism, and they are of three types: triiodothyronine (T3), thyroxin (T4), and thyroid-stimulating hormone (TSH). If these hormones are produced in excess, it is hyperthyroidism, and if in less, it is hypothyroidism. Some symptoms, in addition to those cited in our example earlier, are intolerance to cold, muscle ache, cramps, constipation, weight gain, or loss. Researchers in [13] have applied neural network models to diagnose thyroid disease. In [14] Alqurashi and Wang worked upon a thyroid dataset with five features using various ensemble clustering methods. Akbas et al. in [15] studied the detection of thyroid cancer using multiple approaches. Other researchers have applied K-Nearest neighbour, Support vector machine, Neural fuzzy methods, random forest tree, extra tree for studying this disease data [16, 17]. Dhyan Chandra Yadav in [18] proposed the prediction of thyroid diseases using a decision tree ensemble approach. In [19], the researchers developed a Computer-aided Diagnosis system using PCA and extreme learning techniques to predict thyroid diseases. The experiments show that a maximum accuracy of 98.1% was obtained. Prasan Kumar Sahu in [20] proposed a cloud-enabled big data framework to provide a healthcare solution. The proposed technique deals with structured and unpackaged data generated by healthcare systems and by the use of wearable body sensors; and results show 98% accuracy in predicting disease using correlation analysis.
In [21], the research was conducted on different patients.TSH has been shown to be related to the value of lipid levels or cholesterol levels. Lipid values increased in patients after the level of TSH decreased. In [22], the researchers have developed the hybrid architecture system using rough data sets theory and machine learning algorithms to predict thyroid diseases. In [23], Zhiwen Yu applied a semi-supervised classifier ensemble approach and inspected the trouble of managing high-dimensional datasets with constrained categorized samples. Nyirenda [24] used a statistical approach to find the relationship between thyroid and vascular disease. Research has found that a patient suffering from thyroid disease is more prone to vascular disease. Significant mortality in patients with thyroid disease due to vascular disease is observed at a later stage. Raghuraman et al. in [25] performed comparative thyroid disease diagnosis using Machine learning techniques—Support Vector Machine (SVM), Multiple Linear Regression and Decision Trees, and the highest accuracy of 97.97% was obtained by the decision tree model. Dharamrajan et al. in [26] applied Support Vector machine (SVM) and Decition tree classifier for thyroid prediction, and obtained an accuracy of 97.35 using decision trees.
3 Methodology
Data set and methods are discussed in this section.The thyroid disease dataset consists of 3152 cases, 23 characteristics and finally a class to predict whether the individual is ill or not We present techniques and experimental set-up used for this task. The work flow of our work begins with preprocessing of data, then applying dimension reduction and data augmentation techniques. After this, classifiers are implemented in a distributed environment, and finally comparative analysis is performed. We begin by describing the three dimension reduction techniques. Dimensionality reduction is a method for obtaining the information with lesser number of dimensions from a high dimensional feature space. In machine learning it is very important for the better classification, regression, presentation and visualization of data to reduce the high-dimensional data collection. It is also helpful to better understand the associations between the data. This allows us to identify the intrinsic dimensionality and generalization of the dataset. Since volume of data is a critical issue in healthcare, data augmentation is applied to synthetically generate data so as to develop deep learning models which are said to be data hungry.
3.1 Principal Component Analysis
Principal component analysis (PCA) is an uncontrolled linear transformation technology commonly used in many fields, mainly for extracting functions and reducing dimensionality. Other common PCA applications include data processing, bonded signals de-noising, genome data analysis, and bioinformatics gene expression levels.PCA allows us to classify data trends based on feature-to-feature correlations. In short, PCA seeks to find the highest-dimensional data range directions and projects them into an equivalent or lesser new subspace than the first.
3.2 Singular Value Decomposition
The Singular Value Decomposition (SVD) of matrices provides us with singular vectors which are of reduced dimension, and may be used for classification very effectively. This is specially so for data matrices which are usually rectangular in nature, and eigenvalue decomposition is not possible. For symmetric matrices, the Spectral Theorem holds, which says that there is a basis of eigenvectors and every eigenvalue is real. The spectral theorem also provides a canonical decomposition, called the spectral decomposition, eigenvalue decomposition, or eigendecomposition, of the underlying vector space on which the operator acts. We now briefly explain the correlation between the spectral decomposition and the SVD. The matrix \(A A^{T}\) is of dimension \(m x m\), a symmetric and positive definite matrix. Thus, \(A^{T} A = VE_{1} V^{T} \) and the \(V\) matrix comprises of the eigen vectors of \(A^{T} A\). These vectors are orthogonal and in \(n\) dimensions. \(E_{1}\) is a diagonal matrix comprising of eigen values of \(A^{T} A\). Similar logic holds true for \(A A^{T} = U E_{2} U^{T}\), and the \(U\) matrix comprises of the eigen vectors of \(A A^{T}\). These vectors are orthogonal and in \(m\) dimensions. \(E_{2}\) is a diagonal matrix comprising of eigen values of \(A A^{T}\). The Singular Value Decomposition of \(A\) uses the \(U\) and the \(V\) which have been introduced earlier to be eigen vectors of \(A A^{T}\) and \(A^{T} A\). The factorization of a rectangular matrix \(A\) (of \(m\) rows and \(n\) columns) into its Singular Value Decomposition is \(A = U \Sigma V^{T}\), such that the columns of \(U\) are the left Singular vectors in \(m\) dimensions and columns of \(V\) are the right Singular vectors in \(n\) dimensions, the matrix ∑ is a diagonal matrix where the numbers on the diagonal are non-negative and are called Singular values. It is interesting how these singular values play an important role in reducing the number of effective dimensions. In our research work, for each class, we applied the Singular Value Decomposition and found the \(U\) singular vectors for non-zero singular values. These \(U_{i}\) were used for classification purpose. Further, we iterated through the number of singular values which were optimally required to perform the classification operation.
3.3 Decision Tree
Decision tree methods build a choice model based on real data attribute values. Decisions are taken for a particular record in tree structures before a prediction is selected. Data for category and regression problems are trained on decisions. Decision trees are always quick and right and offer explainable solutions. A decision tree is a tree design, where each inner node (non-leaf node) is a test attribute and each branch is a test result. The leaf nodes are the class nodes The objective is a model based on the input variables, which will estimate the value of the destination variable. In our work, decision trees have been used to identify the features in the order of decreasing importance.
3.4 Building Classifiers
After feature reduction, the K-Nearest Neighbour (KNN) and Neural Network (NN) classifiers are built. We present the outline of algorithms for implementing feature reduction and classification. Algorithm 1 depicts the pseudocode for SVD with KNN classifier. Step 1 loads the dataset in Resilient Distributed Datasets (RDD). Step 2 does the preprocessing and normalization of the dataset. Step 3 deals with splitting the dataset into training (80%) and testing data(20%). Testing data is broadcasted in each slave to receive only one copy of testing data (Step 4). SVD is applied to the training data and U left singular vectors are obtained, representing the training data (Step 7–8). Further, Euclidean distance between U and test data is calculated, and distances are collected at master. Then we apply the KNN classifier. (Step 9–12).


Algorithm 2 represents the steps for the feature reduction technique with a Neural network classifier. Steps 1–2 are the same as in Algorithm 1. Feature reduction technique DT or PCA is applied on RDD Dataset and data is split in the same manner as in Step-3 in Algorithm 1. The model is prepared by applying the neural network classifier on training data. Further, the model is tested on the testing data to predict the accuracy score. (Step 5–7).
3.5 Data Augmentation and Deep Learning
For applying the data augmentation, we created the 10,000 samples using Gaussian distribution. The ratio of class 0(non-thyroid) and class 1(thyroid) is 91:9 in the original dataset. The mean and standard deviation of the features have been calculated for each class label. So, we created 900 samples of class 1 and 9100 samples of class 0 using \(Gaussian\left( {mu,sigma} \right) + random\left( \lambda \right)\), where \(mu\) represents the mean of each feature and \(sigma\) denotes the standard deviation of each sample and noise term is added with a random number \({ }\lambda \in \left( { - 0.1,0.1} \right).\) We created 20% samples of 10,000 samples for validation purposes.
3.6 Data Pre-processing and Normalization
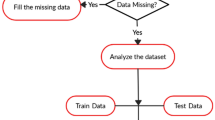
Information pre-handling addresses the primary assignment in data mining procedures. It includes cleaning, extraction, and change of information into a reasonable arrangement for machine execution. Crude information contains missing data and invalid data. It prompts a debacle in the forecast with machine learning. Categorical variables, consisting of categorical values are replaced by 0 and 1. For example, Male and females are replaced by 1 and 0. Normalization is a very important task in the deep learning task. It involves the standardization of the data.
4 Experiments
The experimental setup used for this research work had five Personal Computers: a single Master Node and four Worker Nodes. Every computer was identical and had this specification: 8 GB RAM(DDR3), Intel Core i7 Processor (5th Gen), and a 1 TB Hard disk. The operating system that has been used is Linux Ubuntu-18.04 with Apache Spark-2.4.3. Python Language is used in the Spark platform.
All experiments were conducted in a distributed environment, on the Spark platform. The data was loaded using \({\text{Data}} = {\text{sc}}.{\text{textfile}}\left( {{\text{file}}} \right)\). Then, we performed preprocessing and removed missing values. The null values are replaced by 0. Then the data was normalized. The data set was split into 80% and 20% training and testing ratios. The test data was broadcast to all worker nodes using, \({\text{testdata}} = {\text{ sc}}.{\text{broadcast}}\left( {{\text{testdata}}} \right)\). The training data was split into the worker nodes using \({\text{rdd}} = {\text{sc}}.{\text{parallelize}}\left( {{\text{train}}} \right)\). The row matrix from the \({\text{rdd}}\) was created using \({\text{mat }} = {\text{RowMatrix}}\left( {{\text{rdd}}} \right).\) After this, the dimension reduction techniques were applied and reduced dimensions fed into the classifiers. All this is executed on worker nodes, and then the distance computation for the test data is done for classification purposes. The master node collects all the distance values and predicts the class label corresponding to the minimum distance. Finally, the accuracy score is calculated.
For the K Nearest neighbour classifier, the values of K were taken to be as 3, 5, 7, 9, and the best results have been reported. The number of features for the input layer is 22 features for PCA-NN, 12 features for SVD—NN and 5 features for DT-NN classifier. In the neural network model, 10 neurons were present in two hidden layers, and sigmoid activation function was used. This is implemented on the Spark platform with \({\text{block Size}} = 128\), \({\text{seed value}} = 1234,\) and activation function is sigmoid.
For Prediction with augmented data and deep neural network, our experiment used the two hidden layers with 16 neurons and one input layer with 23 inputs with activation function Rectified Linear Unit. The output layer has one neuron with an activation function sigmoid. In this experiment, we set the batch size = 64, a number of epochs = 100, and an experiment was conducted to validate the 20% data of the entire dataset. Figure 1 shows the architecture of deep learning neural network.
5 Results
All experiments were conducted in a distributed environment on the Spark platform. The dimension reduction techniques were applied, and then the features identified by these techniques were input into the classifiers. For the K Nearest neighbor classifier, the values of K are taken to be as 3, 5, 7, 9, and the best results have been reported. In the neural network, we set the parameters as, Maximum iteration = 100 and number of layers = [no of features, 10, 10, 2]. After 100 iterations, the error did not converge. The odd values of k in KNN had been taken into consideration because of the majority of voting classifiers take these values, and is also available as the options to find the best value of k in the python libraries.
5.1 Dataset Description and Exploratory Analysis
Table 1 shows the dataset for thyroid disease, composed of 3152 instances, 23 features, and class [27]. The thyroid dataset aims to predict whether the person is suffering from sickness-euthyroid disease or not.
The names and description of various Features is given in Table 2.
The distribution of the classes of thyroid dataset is shown as (Table 3):
Next, we find the importance of each feature using Gini index, as given in Fig. 2 and further, the correlation between different features is presented in Fig. 3.
5.2 Comparative Analysis of Classifier Performance
From the values in Table 4, it can be seen that as a dimension reduction technique, the singular value decomposition performs better than principal components analysis, while the decision tree is better than singular value decomposition. The best accuracy of 98.70% is obtained by the decision tree dimension reduction technique, which selects five features and the neural network classifier. Note that the values of F1-score, precision, and recall are also the best. The same is displayed in Fig. 4 plot. Table 5 shows the total run time of different classifiers. It shows that the Neural network classifier takes a little higher time than the K-NN classifier.
In Table 6 we present the results of the deep neural network model built with augmented data. From the values in Table 6, it can be seen that we got the highest parameters score than the earlier results in Table 5. Note that the values of F1-score, precision, and recall are also the best.
Figure 5 shows that accuracy varies almost as much as training and testing. It reached a maximum of 99.95% at its peak of testing data.
Figure 6 shows the loss between training and validation data. Initially, the loss of training data is high and then gets reduced to a loss of validation data in 100 epochs.
Finally, in Table 7 and Fig. 7 we give a comparison of our model performance with other researchers. Ioniţă and Ioniţă, in their work in [28] apply Naive bayes, Decision tree, Multilayer perceptron, and Radial basis function network. Tyagi et al. [29] also use a decision trees along with artificial neural networks for the classification of the thyroid datasets.
Sivasakthivel et al. [30] apply different kinds of decision tree classifiers for the same purpose. Li-Na Li in [19] developed a Computer-aided Diagnosis system using PCA and extreme learning techniques to predict thyroid diseases, and a maximum accuracy of 98.1% was obtained. Prasan Kumar Sahu in [20] proposed a cloud-enabled big data framework to provide a healthcare solution and results show 98% accuracy in predicting disease using correlation analysis. Raghuraman et al. in [25] performed comparative thyroid disease diagnosis using Machine learning techniques—Support Vector Machine (SVM), Multiple Linear Regression and Decision Trees, and the highest accuracy of 97.97% was obtained by the decision tree model. Dharamrajan et al. in [26] applied Support Vector machine (SVM) and Decition tree classifier for thyroid prediction, and obtained an accuracy of 97.35 using decision trees. Finally our two proposed techniques, the first with feature reduction shows an accuracy of 98.7% while the second with data augmentation technique gives an accuracy of 99.95%, and outperform all the others.
6 Conclusion
An enormous growth has been observed in medical expert systems in recent years, and the systems available are now sufficiently developed to be targeted in practice. In order to provide patient care more efficiently, however, expert systems will gradually be incorporated into hospital information systems. For treatments like the production and design of vaccinations, medical data are essential. The dataset is collected in the medical application through the testing of the patient's response to a particular medicine or the collection of medical tests to diagnose a certain medical condition. Thyroidism is specially hard to determine because symptoms can easily be confused with other symptoms. Therapy can regulate dysfunction by early diagnosis of thyroid disease. A modeling solution for the prediction of the thyroid disease is suggested in this study to allow society to enjoy the research progress of computer techniques. The thyroid disease dataset consists of 3152 cases, 23 characteristics and finally a class to predict whether the individual is ill or not. The techniques for dimension reduction and data augmentation are used and used as input to two classifiers. The detailed results of experiments are presented in Tables 4, 5 and 6. A comparative analysis with the study of other researchers in Table 7 shows that our techniques of feature reduction and data augmentation pareform really well with accuracy of 98.7% and 99.95%. As part of our ongoing work, we aim to apply deep learning models for prediction of complex life threatening diseases.
References
Anwar, S., Prasad, R., Chowdhary, B. S., et al. (2019). A telemedicine platform for disaster management and emergency care. Wireless Personal Communications, 106, 191–204. https://doi.org/10.1007/s11277-019-06273-6
Prasad, S., & Prasad, R. (2020). Child temperature monitoring system. Wireless Personal Communications, 115, 711–723. https://doi.org/10.1007/s11277-020-07595-6
Anwar, S., & Prasad, R. (2020). Connections of chronic diseases and socio-dynamic cues for integrating ICT with care plan adherence. Wireless Personal Communications, 113, 1567–1578. https://doi.org/10.1007/s11277-020-07299-x
Koren, A., Jurčević, M., & Prasad, R. (2020). Comparison of data-driven models for cleaning eHealth sensor data: Use case on ECG signal. Wireless Personal Communications, 114, 1501–1517. https://doi.org/10.1007/s11277-020-07435-7
Islam, S. M. R., Kwak, D., Kabir, M. H., Hossain, M., & Kwak, K.-S. (2015). The internet of things for health care: A comprehensive survey. IEEE Access, 3, 678–708.
Raghupathi, W., & Raghupathi, V. (2014). Big data analytics in healthcare: Promise and potential. Health Information Science and Systems, 2(1), 1–10.
Yu, Z., et al. (2016). Incremental semi-supervised clustering ensemble for high dimensional data clustering. IEEE Transactions on Knowledge and Data Engineering, 28(3), 701–714.
Rallapalli, S., Gondkar, R. R., & Ketavarapu, U. P. K. (2016). Impact of processing and analyzing healthcare big data on cloud computing environment by implementing hadoop cluster. Procedia Computer Science, 85, 16–22.
Wang, S., Chang, X., Li, X., Long, G., Yao, L., & Sheng, Q. Z. (2016). Diagnosis code assignment using sparsity-based disease correlation embedding. IEEE Transactions on Knowledge and Data Engineering, 28(12), 3191–3202.
Huang, T., Lan, L., Fang, X., An, P., Min, J., & Wang, F. (2015). Promises and challenges of big data computing in health sciences. Big Data Research, 2(1), 2–11.
Dean, J., & Ghemawat, S. (2008). MapReduce: Simplified data processing on large clusters. Communications of the ACM, 51(1), 107–113.
Menger, V., Scheepers, F., & Spruit, M. (2018). Comparing deep learning and classical machine learning approaches, for predicting inpatient violence incidents from clinical text. Applied Sciences, 8, 981.
Ozyilmaz, L., Yildirim T. (2002). Diagnosis of thyroid disease using artificial neural network methods. In Proceedings of the 9th international conference on neural information processing, 2002. ICONIP '02., Singapore, pp. 2033–2036 vol.4, doi: https://doi.org/10.1109/ICONIP.2002.1199031.
Alqurashi, T., & Wang, W. (2019). Clustering ensemble method. International Journal of Machine Learning and Cybernetics, 10(6), 1227–1246.
Akbas, A., Turhal, U., Babur, S., & Avci, C. (2013). Performance improvement with combining multiple approaches to diagnosis of thyroid cancer. Engineering, 5(10), 264–267.
Awasthi, A. K., Antony, A. (2018). An intelligent system for thyroid disease classification and diagnosis. In 2018 Second international conference on inventive communication and computational technologies(ICICCT). IEEE, pp 1261–1264.
Azar, A. T., Hassanien, A. E., Kim, T. H. (2012). Expert system based on neural-fuzzy rules for thyroid diseases diagnosis. In Computer applications for bio-technology, multimedia, and Ubiquitous City. Springer, Berlin, Heidelberg pp. 94–105.
Yadav, D. C., & Pal, S. (2020). Prediction of thyroid disease using decision tree ensemble method. Human-Intelligent Systems Integration, 2, 89–95.
Li, L.-N., Ouyang, J.-H., Chen, H.-L., & Liu, D.-Y. (2012). A computer aided diagnosis system for thyroid disease using extreme learning machine. Journal of Medical Systems, 36(5), 3327–3337.
Sahoo, P. K., Mohapatra, S. K., & Wu, S.-L. (2016). Analyzing healthcare big data with prediction for future health condition. IEEE Access, 4, 9786–9799.
Canaris, G. J., Manowitz, N. R., Mayor, G., & Ridgway, E. C. (2000). The Colorado thyroid disease prevalence study. Archives of Internal Medicine, 160(4), 526–534.
Prasad, V., Srinivasa Rao, T., & Surendra Prasad Babu, M. (2016). Thyroid disease diagnosis via hybrid architecture composing rough data sets theory and machine learning algorithms. Soft Computing, 20(3), 1179–1189.
Yu, Z., Zhang, Y., You, J., Philip Chen, C. L., Wong, H.-S., Han, G., & Zhang, J. (2017). Adaptive semi-supervised classifier ensemble for high dimensional data classification. IEEE Transactions on Cybernetics, 49, 366–379.
Nyirenda, M. J., Clark, D. N., Finlayson, A. R., Read, J., Elders, A., Bain, M., Fox, K. A. A., & Toft, A. D. (2005). Thyroid disease and increased cardiovascular risk. Thyroid, 15(7), 718–724.
Raghuraman, M. T., Sailatha, E., Gunasekaran, S. (2019). Efficient thyroid disease prediction and comparative study using machine learning algorithms. International Journal of Information and Computing Science. 6(6), 617–624.
Dharmarajan, K., Balasree, K., Arunachalam, A. S., & Abirmai, K. (2020). Thyroid disease classification using decision tree and SVM. Indian Journal of Public Health Research & Development, 11(03), 229–234.
UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science
Ioniţă, I., & Ioniţă, L. (2016). Prediction of thyroid disease using data mining techniques. BRAIN. Broad Research in Artificial Intelligence and Neuroscience, 7(3), 115–124.
Tyagi, A., Mehra, R., Saxena, A. (2018). Interactive thyroid disease prediction system using machine learning technique. In 2018 Fifth international conference on parallel, distributed and grid computing (PDGC). IEEE, pp 689–693.
Sivasakthivel, A., & Shrivakshan, G. T. (2017). A comparative study of diag-nosing thyroid diseases using classification algorithm. International Journals of Advanced Research in Computer Science and Software Engineering, 7(8), 181. ISSN: 2277-128X.
Acknowledgements
The authors acknowledge the anonymous reviewers for their insightful comments which helped in preparing the paper in its present form.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by RJ, VB and AM. The first draft of the manuscript was written by VB and all authors commented on previous versions of the manuscript. The revised version has been prepared by the contribution by all authors. All authors read and approved the final revised manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jha, R., Bhattacharjee, V. & Mustafi, A. Increasing the Prediction Accuracy for Thyroid Disease: A Step Towards Better Health for Society. Wireless Pers Commun 122, 1921–1938 (2022). https://doi.org/10.1007/s11277-021-08974-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11277-021-08974-3