Abstract
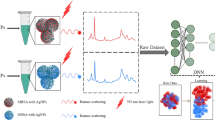
Bacterial species within the Acinetobacter baumannii-calcoaceticus (Acb) complex are very similar and are difficult to discriminate. Misidentification of these species in human infection may lead to severe consequences in clinical settings. Therefore, it is important to accurately discriminate these pathogens within the Acb complex. Raman spectroscopy is a simple method that has been widely studied for bacterial identification with high similarities. In this study, we combined surfaced-enhanced Raman spectroscopy (SERS) with a set of machine learning algorithms for identifying species within the Acb complex. According to the results, the support vector machine (SVM) model achieved the best prediction accuracy at 98.33% with a fivefold cross-validation rate of 96.73%. Taken together, this study confirms that the SERS-SVM method provides a convenient way to discriminate between A. baumannii, Acinetobacter pittii, and Acinetobacter nosocomialis in the Acb complex, which shows an application potential for species identification of Acinetobacter baumannii-calcoaceticus complex in clinical settings in near future.



Similar content being viewed by others
Data availability
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation. No datasets were generated or analysed during the current study.
References
Ahuja T, Ghosh A, Mondal S, Basuri P, Jenifer SK, Srikrishnarka P et al (2019) Ambient electrospray deposition Raman spectroscopy (AESD RS) using soft landed preformed silver nanoparticles for rapid and sensitive analysis. Analyst 144(24):7412–7420
Bagudo AI, Obande GA, Harun A, Singh KKB (2020) Advances in automated techniques to identify Acinetobacter calcoaceticus–Acinetobacter baumannii complex. Asian Biomed 14(5):177–186
Bernards AT, Toorn J, Boven CPA, Dijkshoorn L (1996) Evaluation of the ability of a commercial system to identify Acinetobacter genomic species. Eur J Clin Microbiol Infect Dis 15(4):303–308
Britton K, Dalterio R, Nelson W, Britt D, Sperry JF (1988) Ultraviolet resonance Raman spectra of Escherichia coli with 222.5–251.0 nm pulsed laser excitation. Appl Spectrosc 42(5):782–788
Ho C-S, Jean N, Hogan CA, Blackmon L, Jeffrey SS, Holodniy M et al (2019) Rapid identification of pathogenic bacteria using Raman spectroscopy and deep learning. Nat Commun 10(1):4927
Jayan H, Pu H, Sun D-W (2022) Analyzing macromolecular composition of E. coli O157: H7 using Raman-stable isotope probing. Spectrochim Acta A 276:121217
Kang HM, Yun KW, Choi EH (2023) Molecular epidemiology of Acinetobacter baumannii complex causing invasive infections in Korean children during 2001–2020. Ann Clin Microbiol Antimicrob. https://doi.org/10.1186/s12941-023-00581-3
Kim J, Chin Y-W (2023) Antimicrobial agent against methicillin-resistant Staphylococcus aureus biofilm monitored using Raman spectroscopy. Pharmaceutics 15(7):1937
Kim S, Kim MH, Lee WI, Kang SY, Jeon YL (2018) Misidentification of Acinetobacter baumannii as Alcaligenes faecalis by VITEK 2 system; case report. Lab Med 49(1):e14–e17
Lin Y-C, Sheng W-H, Chang S-C, Wang J-T, Chen Y-C, Wu R-J et al (2008) Application of a microsphere-based array for rapid identification of Acinetobacter spp. with distinct antimicrobial susceptibilities. J Clin Microbiol 46(2):612–617
Liu PY-F, Wu W-L (1997) Use of different PCR-based DNA fingerprinting techniques and pulsed-field gel electrophoresis to investigate the epidemiology of Acinetobacter calcoaceticus-Acinetobacter baumannii complex. Diagn Microbiol Infect Dis 29(1):19–28
Liu W, Tang J-W, Lyu J-W, Wang J-J, Pan Y-C, Shi X-Y et al (2022) Discrimination between carbapenem-resistant and carbapenem-sensitive Klebsiella pneumoniae strains through computational analysis of surface-enhanced Raman spectra: a pilot study. Microbiol Spectr. https://doi.org/10.1128/spectrum.02409-21
Liu W, Tang J-W, Mou J-Y, Lyu J-W, Di Y-W, Liao Y-L et al (2023) Rapid discrimination of Shigella spp. and Escherichia coli via label-free surface enhanced Raman spectroscopy coupled with machine learning algorithms. Front Microbiol 14:1101357
Lu X, Rasco BA, Jabal JM, Aston DE, Lin M, Konkel ME et al (2011) Investigating antibacterial effects of garlic (Allium sativum) concentrate and garlic-derived organosulfur compounds on Campylobacter jejuni by using Fourier transform infrared spectroscopy, Raman spectroscopy, and electron microscopy. Appl Environ Microbiol 77(15):5257–5269
Lu W, Li H, Qiu H, Wang L, Feng J, Fu YV (2023) Identification of pathogens and detection of antibiotic susceptibility at single-cell resolution by Raman spectroscopy combined with machine learning. Front Microbiol 13:1076965
Lynch JP, Clark NM, Zhanel GG (2022) Infections due to Acinetobacter baumannii–calcoaceticus complex: escalation of antimicrobial resistance and evolving treatment options. Semin Respir Crit Care Med 43(01):97–124
Lyu J-W, Zhang XD, Tang J-W, Zhao Y-H, Liu S-L, Zhao Y et al (2023) Rapid prediction of multidrug-resistant Klebsiella pneumoniae through deep learning analysis of SERS spectra. Microbiol Spectr. https://doi.org/10.1128/spectrum.04126-22
Milani ES, Hasani A, Varschochi M, Sadeghi J, Memar MY, Hasani A (2021) Biocide resistance in Acinetobacter baumannii: appraising the mechanisms. J Hosp Infect 117:135–146
Nocera FP, Attili A-R, De Martino L (2021) Acinetobacter baumannii: its clinical significance in human and veterinary medicine. Pathogens 10(2):127
Seifert H, Gerner-Smidt P (1995) Comparison of ribotyping and pulsed-field gel electrophoresis for molecular typing of Acinetobacter isolates. J Clin Microbiol 33(5):1402–1407
Sharma P, Das A, Rao KH, Forssberg K (2003) Surface characterization of Acidithiobacillus ferrooxidans cells grown under different conditions. Hydrometallurgy 71(1–2):285–292
Sousa C, Botelho J, Silva L, Grosso F, Nemec A, Lopes J et al (2014a) MALDI-TOF MS and chemometric based identification of the Acinetobacter calcoaceticus-Acinetobacter baumannii complex species. Int J Med Microbiol 304(5–6):669–677
Sousa C, Silva L, Grosso F, Nemec A, Lopes J, Peixe L (2014b) Discrimination of the Acinetobacter calcoaceticus–Acinetobacter baumannii complex species by Fourier transform infrared spectroscopy. Eur J Clin Microbiol Infect Dis 33(8):1345–1353
Tang J-W, Liu Q-H, Yin X-C, Pan Y-C, Wen P-B, Liu X et al (2021) Comparative analysis of machine learning algorithms on surface enhanced Raman spectra of clinical Staphylococcus species. Front Microbiol 12:696921
Tang J-W, Li J-Q, Yin X-C, Xu W-W, Pan Y-C, Liu Q-H et al (2022) Rapid discrimination of clinically important pathogens through machine learning analysis of surface enhanced Raman spectra. Front Microbiol 13:843417
Tang J-W, Lyu J-W, Lai J-X, Zhang X-D, Du Y-G, Zhang X-Q et al (2023) Determination of Shigella spp. via label-free SERS spectra coupled with deep learning. Microchem J 189:108539
Teixeira AM, Nemec A, Sousa C (2019) Differentiation of taxonomically closely related species of the genus Acinetobacter using Raman spectroscopy and chemometrics. Molecules 24(1):168
Ugolotti E, Marco ED, Bandettini R, Tripodi G, Biassoni R (2016) The whole genome sequencing of Acinetobacter-calcoaceticus-baumannii complex strains involved in suspected outbreak in an Intensive Care Unit of a pediatric hospital. J Hosp Adm 5(6):81
Usman M, Tang J-W, Li F, Lai J-X, Liu Q-H, Liu W et al (2023) Recent advances in surface enhanced Raman spectroscopy for bacterial pathogen identifications. J Adv Res 51:91–107
Vila J, Marcos MA, Jimenez de Anta MT (1996) A comparative study of different PCR-based DNA fingerprinting techniques for typing of the Acinetobacter calcoaceticus-A. baumannii complex. J Med Microbiol 44(6):482–489
Wang L, Liu W, Tang J-W, Wang J-J, Liu Q-H, Wen P-B et al (2021) Applications of Raman spectroscopy in bacterial infections: principles, advantages, and shortcomings. Front Microbiol. https://doi.org/10.3389/fmicb.2021.683580
Wang L, Tang J-W, Li F, Usman M, Wu C-Y, Liu Q-H et al (2022) Identification of bacterial pathogens at genus and species levels through combination of Raman spectrometry and deep-learning algorithms. Microbiol Spectr. https://doi.org/10.1128/spectrum.02580-22
Whiteway C, Breine A, Philippe C, Van der Henst C (2022) Acinetobacter baumannii. Trends Microbiol 30(2):199–200
Xu J, Wu D, Li Y, Xu J, Gao Z, Song Y-Y et al (2018) Plasmon-triggered hot-spot excitation on SERS substrates for bacterial inactivation and in situ monitoring. ACS Appl Mater Interfaces 10(30):25219–25227
Yamada S, Kurotani A, Chikayama E, Kikuchi J (2020) Signal deconvolution and noise factor analysis based on a combination of time–frequency analysis and probabilistic sparse matrix factorization. Int J Mol Sci 21(8):2978
Zhang L-Y, Tian B, Huang Y-H, Gu B, Ju P, Luo Y et al (2023) Classification and prediction of Klebsiella pneumoniae strains with different MLST allelic profiles via SERS spectral analysis. PeerJ 11:e16161
Acknowledgements
We thank the anonymous reviewers for their thoughtful comments that greatly improve the quality of the manuscript.
Funding
This study was supported by Guangdong Basic and Applied Basic Research Foundation [Grant No. 2022A1515220023], Talent Start-Up Funding of Guangdong Provincial People’s Hospital [Grant No. KY012023293], and Research Funding for Guangdong Provincial Medical Technology [Grant No. B2021010].
Author information
Authors and Affiliations
Contributions
L.W., S.L.L. and J.W.T. conceived and designed the experiments. L.W. provided platform and resources, and contributed to project administration and student supervision. X.S.X., L.F.Y., and Y.F.L. carried out the experimental and computational investigations. Q.Y., Y.T.S., J.C., and X.R.W. contributed to the data analysis and visualization. All the authors wrote and revised the manuscript. All the authors approved the submitted version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiong, XS., Yao, LF., Luo, YF. et al. Differentiation of closely-related species within Acinetobacter baumannii-calcoaceticus complex via Raman spectroscopy: a comparative machine learning analysis. World J Microbiol Biotechnol 40, 146 (2024). https://doi.org/10.1007/s11274-024-03948-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-024-03948-6