Abstract
Biogas, produced in anaerobic digestion, is a sustainable alternative for generating energy from agro-industrial and municipal waste. Information from the microbiota active in the process expands the possibilities for technological innovation. In this study, taxonomic annotations, and functional prediction of the microbial community of the inoculum of two processes were carried out: an industrial unit (pilot-scale urban solid waste plant—IU) and a laboratory-scale reactor fed with swine and cattle waste (LS). The biochemical potential of biogas was obtained using tested inoculum with microcrystalline cellulose, obtaining 682 LN/kgVS (LSC—laboratory scale inoculum and microcrystalline cellulose), and 583 LN/kgVS (IUC—industrial unit inoculum and microcrystalline cellulose), which is equivalent to a recovery of 91.5% of total biogas to LSC. The phyla Synergistota and Firmicutes were more abundant in LS/LSC. In the IU/IUC (treatment of restaurant waste and customs seizures), there was a greater microbiological variety and a predominance of the Bacteroidota, Cloacimonadota, Firmicutes and Caldatribacteriota. The genus Methanosaeta predominated in the process, and it was possible to infer the genes (K01895, K00193 and K00625) related to acetoclastic pathway, as well as endoglucanases that are involved in the metabolism of cellulose (LSC). Terpenoids, polyketides, cofactors, and vitamin metabolism were higher in reactors that received different substrates (IU; IUC). The taxonomic and functional differences revealed the importance of determining the microbiota in the analysis of the potential of an inoculum, combined with the use of microcrystalline cellulose, which can provide optimization information in the production of clean energy.
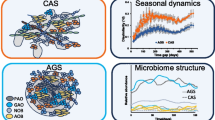
Graphical Abstract








Similar content being viewed by others
Data availability
The data used to support the findings of this study is available upon request.
References
Amin FR, Khalid H, El-Mashad HM et al (2021) Functions of bacteria and archaea participating in the bioconversion of organic waste for methane production. Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2020.143007
Andrews S, Lindenbaum P, Howard B, Ewels P (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/
APHA, AWWA, WEF (2017) Standard Methods for the Examination of Water and Wastewater. Washington, DC.
Bedoya K, Hoyos O, Zurek E et al (2020) Annual microbial community dynamics in a full-scale anaerobic sludge digester from a wastewater treatment plant in Colombia. Sci Total Environ 726:138479. https://doi.org/10.1016/j.scitotenv.2020.138479
Bolyen E, Rideout JR, Dillon MR, Bokulich NA et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Bonugli-Santos RC, Marteres TJ, Luiz FN et al (2022) Prolonged acetogenic phase and biological succession during anaerobic digestion using swine manure. Folia Microbiol (praha). https://doi.org/10.1007/s12223-021-00937-2
Brasil 2021 Ministério de Minas e Energia. Conselho nacional de política energética - CNPE. Resoluçãoo n° 7, de 20 de abril de 2021. https://www.in.gov.br/en/web/dou/-/despacho-do-presidente-da-republica-320067170
Calabrò PS, Fazzino F, Pangallo D (2021) How does separate collection efficiency influence biological stability of commingled Italian municipal solid waste? Sustain Chem Pharm. https://doi.org/10.1016/j.scp.2020.100355
Calderon AG, Duan H, Chen X et al (2021) Enhancing anaerobic digestion using free nitrous acid: Identifying the optimal pre-treatment condition in continuous operation. Water Res 205:117694. https://doi.org/10.1016/j.watres.2021.117694
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Camargo FP, Sakamoto IK, Duarte ICS et al (2021) Metataxonomic characterization of bacterial and archaeal community involved in hydrogen and methane production from citrus peel waste (Citrus sinensis L. Osbeck) in batch reactors. Biomass and Bioenergy. https://doi.org/10.1016/j.biombioe.2021.106091
Caporaso JG, Lauber CL, Walters WA et al (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Chang Bejarano A, Champagne P (2022) Optimization of biogas production during start-up with electrode-assisted anaerobic digestion. Chemosphere 302:134739. https://doi.org/10.1016/j.chemosphere.2022.134739
Chen L, Meng X, Zhou G et al (2022) Effects of organic loading rates on the anaerobic co-digestion of fresh vinegar residue and pig manure: Focus on the performance and microbial communities. Biochem Eng J 183:108441. https://doi.org/10.1016/j.bej.2022.108441
Chernicharo CAL (2007) Biological wastewater treatment: anaerobic reactors. IWA Publishing, London
Christou ML, Vasileiadis S, Kalamaras SD et al (2021) Ammonia-induced inhibition of manure-based continuous biomethanation process under different organic loading rates and associated microbial community dynamics. Bioresour Technol 320:124323. https://doi.org/10.1016/j.biortech.2020.124323
CIBiogas 2021 Panorama do Biogás no Brasil 2021. Foz do Iguaçu.
Douglas GM, Maffei VJ, Zaneveld JR et al (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38:685–688. https://doi.org/10.1038/s41587-020-0548-6
Gadaleta G, De Gisi S, Picuno C et al (2022) Energy recovery options for the management of cellulose-based bio-plastics and mixed municipal solid waste. Biomass and Bioenergy 166:106628. https://doi.org/10.1016/j.biombioe.2022.106628
Gallardo-Altamirano MJ, Maza-Márquez P, Montemurro N et al (2021) Insights into the removal of pharmaceutically active compounds from sewage sludge by two-stage mesophilic anaerobic digestion. Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2021.147869
Hahnke S, Langer T, Koeck DE, Klocke M (2016) Description of Proteiniphilum saccharofermentans sp. nov., Petrimonas mucosa sp. nov. and Fermentimonas caenicola gen. nov., sp. nov., isolated from mesophilic laboratory-scale biogas reactors, and emended description of the genus Proteiniphilum. Int J Syst Evol Microbiol 66:1466–1475. https://doi.org/10.1099/ijsem.0.000902
Hammer Ø, Harper DAT, Ryan PD (2009) PAST–palaeontological statistics. Palaeontol Electron 4:1–9
Hu F, Zhang S, Wang X et al (2022) Investigating the role of different materials supplementation in anaerobic digestion of kitchen waste: performance and microbial community dynamics. Biochem Eng J 184:108490. https://doi.org/10.1016/j.bej.2022.108490
Iltchenco J, Peruzzo V, Magrini FE et al (2021) Microbiota profile in mesophilic biodigestion of sugarcane vinasse in batch reactors. Water Sci Technol 84:2028–2039. https://doi.org/10.2166/wst.2021.375
Ji Y, Liu P, Conrad R (2018) Response of fermenting bacterial and methanogenic archaeal communities in paddy soil to progressing rice straw degradation. Soil Biol Biochem 124:70–80. https://doi.org/10.1016/j.soilbio.2018.05.029
Jiang X, Lyu Q, Bi L et al (2022) Improvement of sewage sludge anaerobic digestion through synergistic effect combined trace elements enhancer with enzyme pretreatment and microbial community response. Chemosphere 286:131356. https://doi.org/10.1016/j.chemosphere.2021.131356
Jurgutis L, Slepetiene A, Volungevicius J, Amaleviciute-Volunge K (2020) Biogas production from chicken manure at different organic loading rates in a mesophilic full scale anaerobic digestion plant. Biomass and Bioenergy 141:105693. https://doi.org/10.1016/j.biombioe.2020.105693
Kanehisa M, Sato Y, Kawashima M et al (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44:457–462. https://doi.org/10.1093/nar/gkv1070
Koch K, Lippert T, Drewes JE (2017) The role of inoculum’s origin on the methane yield of different substrates in biochemical methane potential (BMP) tests. Bioresour Technol 243:457–463. https://doi.org/10.1016/j.biortech.2017.06.142
Kunz A, Steinmetz RLR, Amaral AC (2022) Fundamentos da digestão anaeróbia, purificação do biogás, uso e tratamento do digestato, 2a Edição. Sbera, Embrapa Suínos e Aves, Brasil
Kurade MB, Saha S, Salama ES et al (2019) Acetoclastic methanogenesis led by Methanosarcina in anaerobic co-digestion of fats, oil and grease for enhanced production of methane. Bioresour Technol 272:351–359. https://doi.org/10.1016/j.biortech.2018.10.047
Lee J, Koo T, Yulisa A, Hwang S (2019) Magnetite as an enhancer in methanogenic degradation of volatile fatty acids under ammonia-stressed condition. J Environ Manage 241:418–426. https://doi.org/10.1016/j.jenvman.2019.04.038
Li C, He P, Hao L et al (2022a) Diverse acetate-oxidizing syntrophs contributing to biogas production from food waste in full-scale anaerobic digesters in China. Renew Energy 193:240–250. https://doi.org/10.1016/j.renene.2022.04.143
Li MT, Rao L, Wang L et al (2022) Bioaugmentation with syntrophic volatile fatty acids-oxidizing consortia to alleviate the ammonia inhibition in continuously anaerobic digestion of municipal sludge. Chemosphere 288:132389. https://doi.org/10.1016/j.chemosphere.2021.132389
Li Y, Chen Z, Peng Y et al (2022) Deeper insights into the effects of substrate to inoculum ratio selection on the relationship of kinetic parameters, microbial communities, and key metabolic pathways during the anaerobic digestion of food waste. Water Res 217:118440. https://doi.org/10.1016/j.watres.2022.118440
Li Y, Jing Z, Pan J et al (2022) Multi-omics joint analysis of the effect of temperature on microbial communities, metabolism, and genetics in full-scale biogas reactors with food waste. Renew Sustain Energy Rev 160:112261. https://doi.org/10.1016/j.rser.2022.112261
Lima MA, Mendes LFR, Mothé GA et al (2020) Renewable energy in reducing greenhouse gas emissions: reaching the goals of the Paris agreement in Brazil. Environ Dev 33:100504. https://doi.org/10.1016/j.envdev.2020.100504
Lu J, Jia Z, Wang P et al (2022) Restoration of acidified dry anaerobic digestion of food waste: bioaugmentation of butyric acid-resistant microbes. J Environ Chem Eng 10:106935. https://doi.org/10.1016/j.jece.2021.106935
Magrini FE, de Almeida GM, da Maia Soares D et al (2020) Effect of different heat treatments of inoculum on the production of hydrogen and volatile fatty acids by dark fermentation of sugarcane vinasse. Biomass Convers Bioref. https://doi.org/10.1007/s13399-020-00687-0
McNally CP, Eng A, Noecker C et al (2018) BURRITO: an interactive multi-omic tool for visualizing taxa-function relationships in microbiome data. Front Microbiol 9:1–11. https://doi.org/10.3389/fmicb.2018.00365
Mlaik N, Karray F, Sayadi S et al (2022) Semi-continuous anaerobic digestion of the organic fraction of municipal solid waste: digester performance and microbial population dynamics. J Environ Chem Eng 10:107941. https://doi.org/10.1016/j.jece.2022.107941
Müller B, Sun L, Westerholm M, Schnürer A (2016) Bacterial community composition and fhs profiles of low- and high-ammonia biogas digesters reveal novel syntrophic acetate-oxidising bacteria. Biotechnol Biofuels 9:1–18. https://doi.org/10.1186/s13068-016-0454-9
Ottoni JR, Bernal SPF, Marteres TJ et al (2022) Cultured and uncultured microbial community associated with biogas production in anaerobic digestion processes. Arch Microbiol 204:1–13. https://doi.org/10.1007/s00203-022-02819-8
Paesi S, Schiavenin A, Almeida LG et al (2022) Comparison of two different kinds of seed sludge and characterization of microorganisms producing hydrogen and soluble metabolites from raw glycerol. Brazilian J Chem Eng 39:387–402. https://doi.org/10.1007/s43153-021-00212-4
Palomo-Briones R, de Montoya-Rosales J, J, Razo-Flores E, (2021) Advances towards the understanding of microbial communities in dark fermentation of enzymatic hydrolysates: diversity, structure and hydrogen production performance. Int J Hydrogen Energy 46:27459–27472. https://doi.org/10.1016/j.ijhydene.2021.06.016
Pellera FM, Gidarakos E (2016) Effect of substrate to inoculum ratio and inoculum type on the biochemical methane potential of solid agroindustrial waste. J Environ Chem Eng 4:3217–3229. https://doi.org/10.1016/j.jece.2016.05.026
Perman E, Schnürer A, Björn A, Moestedt J (2022) Serial anaerobic digestion improves protein degradation and biogas production from mixed food waste. Biomass and Bioenergy 161:106478. https://doi.org/10.1016/j.biombioe.2022.106478
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Rehman M, Iqbal A, Chang C et al (2019) Anaerobic digestion. Water Environ Res 91:1253–1271
Shamurad B, Sallis P, Petropoulos E et al (2020) Stable biogas production from single-stage anaerobic digestion of food waste. Appl Energy. https://doi.org/10.1016/j.apenergy.2020.114609
Steinmetz RLR, Mezzari MP, da Silva MLB et al (2016) Enrichment and acclimation of an anaerobic mesophilic microorganism’s inoculum for standardization of BMP assays. Bioresour Technol 219:21–28. https://doi.org/10.1016/j.biortech.2016.07.031
Stiborova H, Strejcek M, Musilova L et al (2020) Diversity and phylogenetic composition of bacterial communities and their association with anthropogenic pollutants in sewage sludge. Chemosphere 238:124629. https://doi.org/10.1016/j.chemosphere.2019.124629
Tápparo DC, Cândido D, Steinmetz RLR et al (2021) Swine manure biogas production improvement using pre-treatment strategies: lab-scale studies and full-scale application. Bioresour Technol Rep. https://doi.org/10.1016/j.biteb.2021.100716
Tomazetto G, Hahnke S, Wibberg D et al (2018) Proteiniphilum saccharofermentans str. M3/6T isolated from a laboratory biogas reactor is versatile in polysaccharide and oligopeptide utilization as deduced from genome-based metabolic reconstructions. Biotechnol Rep 18:e00254. https://doi.org/10.1016/j.btre.2018.e00254
Town JR, Dumonceaux TJ (2016) Laboratory-scale bioaugmentation relieves acetate accumulation and stimulates methane production in stalled anaerobic digesters. Appl Microbiol Biotechnol 100:1009–1017. https://doi.org/10.1007/s00253-015-7058-3
Value MF, Value F, Sample I, et al (2021) Determination of FOS/TAC Value in Biogas Reactors. 2014–2015
VDI 4630 (2006) Fermentation of organic materials Characterisation of the substrate, sampling, collection of material data, fermentation tests
Wang Y, Qian PY (2009) Conservative fragments in bacterial 16S rRNA genes and primer design for 16S ribosomal DNA amplicons in metagenomic studies. PLoS ONE. https://doi.org/10.1371/journal.pone.0007401
Wang S, Hou X, Su H (2017) Exploration of the relationship between biogas production and microbial community under high salinity conditions. Sci Rep 7:1–10. https://doi.org/10.1038/s41598-017-01298-y
Werner JJ, Knights D, Garcia ML et al (2011) Bacterial community structures are unique and resilient in full-scale bioenergy systems. Proc Natl Acad Sci USA 108:4158–4163. https://doi.org/10.1073/pnas.1015676108
Westerholm M, Müller B, Singh A et al (2018) Detection of novel syntrophic acetate-oxidizing bacteria from biogas processes by continuous acetate enrichment approaches. Microb Biotechnol 11:680–693. https://doi.org/10.1111/1751-7915.13035
Xiao B, Tang X, Zhang W et al (2022) Effects of rice straw ratio on mesophilic and thermophilic anaerobic co-digestion of swine manure and rice straw mixture. Energy 239:122021. https://doi.org/10.1016/j.energy.2021.122021
Yang S, Chen Z, Wen Q (2021) Impacts of biochar on anaerobic digestion of swine manure: methanogenesis and antibiotic resistance genes dissemination. Bioresour Technol 324:124679. https://doi.org/10.1016/j.biortech.2021.124679
Yang S, Xue W, Liu P et al (2022) Revealing the methanogenic pathways for anaerobic digestion of key components in food waste: performance, microbial community, and implications. Bioresour Technol 347:126340. https://doi.org/10.1016/j.biortech.2021.126340
Yong TW, Sharma P, Tian H et al (2023) Startup performance and microbial communities of a decentralized anaerobic digestion of food waste. Chemosphere. https://doi.org/10.1016/j.chemosphere.2023.137937
Zhan Y, Cao X, Xiao Y et al (2022) Start-up of co-digestion of poultry litter and wheat straw in anaerobic sequencing batch reactor by gradually increasing organic loading rate: methane production and microbial community analysis. Bioresour Technol 354:127232. https://doi.org/10.1016/j.biortech.2022.127232
Zhang W, Wang X, Xing W et al (2021) Links between synergistic effects and microbial community characteristics of anaerobic co-digestion of food waste, cattle manure and corn straw. Bioresour Technol 329:124919. https://doi.org/10.1016/j.biortech.2021.124919
Zhang Y, Zhang L, Yu N et al (2022) Enhancing the resistance to H2S toxicity during anaerobic digestion of low-strength wastewater through granular activated carbon (GAC) addition. J Hazard Mater 430:128473. https://doi.org/10.1016/j.jhazmat.2022.128473
Zhu K, Zhang L, Wang X et al (2021) Inhibition of norfloxacin on anaerobic digestion: focusing on the recoverability and shifted microbial communities. Sci Total Environ 752:141733. https://doi.org/10.1016/j.scitotenv.2020.141733
Zhu R, Wang DH, Zheng Y et al (2022) Understanding the mechanisms behind micro-aeration to enhance anaerobic digestion of corn straw. Fuel. https://doi.org/10.1016/j.fuel.2022.123604
Zhuravleva EA, Shekhurdina SV, Kotova IB et al (2022) Effects of various materials used to promote the direct interspecies electron transfer on anaerobic digestion of low-concentration swine manure. Sci Total Environ 839:156073. https://doi.org/10.1016/j.scitotenv.2022.156073
Ziels RM, Beck DAC, Stensel HD (2017) Long-chain fatty acid feeding frequency in anaerobic codigestion impacts syntrophic community structure and biokinetics. Water Res 117:218–229. https://doi.org/10.1016/j.watres.2017.03.060
Acknowledgements
We thank the International Center for Renewable Energies—Biogás supported by Itaipu Binacional (supported by Itaipu Technological Park Foundation) and EDITAL 105/2020/PRPPG-Priority Latin America and the Caribbean.
Funding
This work was supported by EDITAL 105/2020/PRPPG-Priority Latin America and the Caribbean.
Author information
Authors and Affiliations
Contributions
FNL analyze the production of biogas and wrote the paper; MRZP analyzed the data and wrote the paper; FEV analyzed the microbiological data and wrote the paper; JG bioinformatics analysis; JGS analyzes of the results and wrote the paper; RFM carried out the experiments and performed analyses; SP wrote the paper and supervised the research. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Luiz, F.N., Passarini, M.R.Z., Magrini, F.E. et al. Metataxonomic characterization of the microbial community involved in the production of biogas with microcrystalline cellulose in pilot and laboratory scale. World J Microbiol Biotechnol 39, 184 (2023). https://doi.org/10.1007/s11274-023-03573-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03573-9