Abstract
Natural pristine environments including cold habitats are thought to be the potent reservoirs of antibiotic-resistant genes and have been recurrently reported in polar glaciers’ native bacteria, nevertheless, their abundance among the non-polar glaciers’ inhabitant bacteria is mostly uncharted. Herein we evaluated antibiotic resistance profile, abundance of antibiotic-resistant genes plus class 1, 2, and 3 integron integrases in 65 culturable bacterial isolates retrieved from a non-polar glacier. The 16S rRNA gene sequencing analysis identified predominantly Gram-negative 43 (66.15%) and Gram-positive 22 (33.84%) isolates. Among the Gram-negative bacteria, Gammaproteobacteria were dominant (62.79%), followed by Betaproteobacteria (18.60%) and Alphaproteobacteria (9.30%), whereas Phyla Actinobacteria (50%) and Firmicutes (40.90%) were predominant among Gram-positive. The Kirby Bauer disc diffusion method evaluated significant antibiotic resistance among the isolates. PCR amplification revealed phylum Proteobacteria predominantly carrying 21 disparate antibiotic-resistant genes like; blaAmpC 6 (100%), blaVIM-1, blaSHV and blaDHA 5 (100%) each, blaOXA-1 1 (100%), blaCMY-4 4 (100%), followed by Actinobacteria 14, Firmicutes 13 and Bacteroidetes 11. Tested isolates were negative for blaKPC, qnrA, vanA, ermA, ermB, intl2, and intl3. Predominant Gram-negative isolates had higher MAR index values, compared to Gram-positive. Alignment of protein homology sequences of antibiotic-resistant genes with references revealed amino acid variations in blaNDM-1, blaOXA-1, blaSHV, mecA, aac(6)-Ib3, tetA, tetB, sul2, qnrB, gyrA, and intI1. Promising antibiotic-resistant bacteria, harbored with numerous antibiotic-resistant genes and class 1 integron integrase with some amino acid variations detected, accentuating the mandatory focus to evaluate the intricate transcriptome analysis of glaciated bacteria conferring antibiotic resistance.
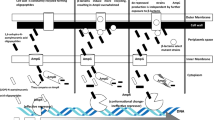
Graphical abstract







Similar content being viewed by others
References
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8(4):251–259. https://doi.org/10.1038/nrmicro2312
Allen HK, Moe LA, Rodbumrer J, Gaarder A, Handelsman J (2009) Functional metagenomics reveals diverse β-lactamases in a remote alaskan soil. ISME J 3(2):243–251. https://doi.org/10.1038/ismej.2008.86
Amato P, Hennebelle R, Magand O, Sancelme M, Delort AM, Barbante C, Boutron C, Ferrari C (2007) Bacterial characterization of the snow cover at Spitzberg, Svalbard. FEMS Microbiol Ecol 59(2):255–264. https://doi.org/10.1111/j.1574-6941.2006.00198.x
Anesio AM, Laybourn-Parry J (2012) Glaciers and ice sheets as a biome. Trends Ecol Evol 27(4):219–225. https://doi.org/10.1016/j.tree.2011.09.012
Anesio AM, Lutz S, Chrismas NA, Benning LG (2017) The microbiome of glaciers and ice sheets. npj Biofilms and Microbiomes 3(1):1–1. https://doi.org/10.1038/s41522-017-0019-0
Baghel VS, Tripathi RD, Ramteke PW, Gopal K, Dwivedi S, Jain RK, Rai UN, Singh SN (2005) Psychrotrophic proteolytic bacteria from cold environment of Gangotri glacier, western Himalaya, India. Enzyme Microb Technol 36(5–6):654–659. https://doi.org/10.1016/j.enzmictec.2004.09.005
Ball MM, Gómez W, Magallanes X, Rosales R, Melfo A, Yarzábal LA (2014) Bacteria recovered from a high-altitude, tropical glacier in venezuelan Andes. World J Microbiol Biotechnol 30(3):931–941. https://doi.org/10.1007/s11274-013-1511-1
Bhatia M, Sharp M, Foght J (2006) Distinct bacterial communities exist beneath a high Arctic polythermal glacier. Appl Environ Microbiol 72(9):5838–5845. https://doi.org/10.1128/AEM.00595-06
Blair J, Webber MA, Baylay AJ, Ogbolu DO, Piddock LJ (2015) Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol 13(1):42–51. https://doi.org/10.1038/nrmicro3380
Boetius A, Anesio AM, Deming JW, Mikucki JA, Rapp JZ (2015) Microbial ecology of the cryosphere: sea ice and glacial habitats. Nat Rev Microbiol 13(11):677–690. https://doi.org/10.1038/nrmicro3522
Cabello FC (2006) Heavy use of prophylactic antibiotics in aquaculture: a growing problem for human and animal health and for the environment. Environ Microbiol 8(7):1137–1144. https://doi.org/10.1111/j.1462-2920.2006.01054.x
Cambray G, Guerout AM, Mazel D (2010) Integrons. Annu Rev Genet 44::41 – 66 https://doi.org/10.1146/annurev-genet-102209-163504
Casanueva A, Tuffin M, Cary C, Cowan DA (2010) Molecular adaptations to psychrophily: the impact of ‘omic’technologies. Trends Microbiol 18(8):374–381. https://doi.org/10.1016/j.tim.2010.05.002
Chaturvedi P, Reddy GS, Shivaji S (2005) Dyadobacter hamtensis sp. nov., from Hamta glacier, located in the Himalayas, India. Int J Syst Evol Microbiol 55(5):2113–2117. https://doi.org/10.1099/ijs.0.63806-0
Chaturvedi P, Shivaji S (2006) Exiguobacterium indicum sp. nov., a psychrophilic bacterium from the Hamta glacier of the Himalayan Mountain ranges of India. Int J Syst Evol Microbiol 56(12):2765–2770. https://doi.org/10.1099/ijs.0.64508-0
Chee-Sanford JC, Aminov RI, Krapac IJ, Garrigues-Jeanjean N, Mackie RI (2001) Occurrence and diversity of tetracycline resistance genes in lagoons and groundwater underlying two swine production facilities. Appl Environ Microbiol 67(4):1494–1502. https://doi.org/10.1128/AEM.67.4.1494-1502.2001
Christner BC, Mosley-Thompson E, Thompson LG, Zagorodnov V, Reeve JN (2002) Isolation and Identification of Bacteria from Ancient and Modern Ice Cores. Patagonian Icefields 9–15. Springer, Boston, MA. https://doi.org/10.1007/978-1-4615-0645-4_2
Catalog (2018) Clinical & Laboratory standards institute. https://clsi.org/media/2042/catalog2018_web-unlinked
Cowan DA, Casanueva A, Stafford W (2007) Ecology and biodiversity of cold-adapted microorganisms. Physiol Biochem Extremophiles 117 – 32. https://doi.org/10.1128/9781555815813.ch9
D’Costa VM, King CE, Kalan L, Morar M, Sung WW, Schwarz C, Froese D, Zazula G, Calmels F, Debruyne R, Golding GB (2011) Antibiotic resistance is ancient. Nature 477(7365):457–461. https://doi.org/10.1038/nature10388
D’Costa VM, McGrann KM, Hughes DW, Wright GD (2006) Sampling the antibiotic resistome. Sci 311(5759):374–377. https://doi.org/10.1126/science.1120800
De Souza MJ, Nair S, Bharathi L, Chandramohan D (2006) Metal and antibiotic-resistance in psychrotrophic bacteria from Antarctic Marine waters Ecotoxicol 15(4):379 – 84. https://doi.org/10.1007/s10646-006-0068-2
Dong H, Zhang G, Jiang H, Yu B, Chapman LR, Lucas CR, Fields MW (2006) Microbial diversity in sediments of saline Qinghai Lake, China: linking geochemical controls to microbial ecology. Microb Ecol 51(1):65–82. https://doi.org/10.1007/s00248-005-0228-6
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MO, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogens. Sci 337(6098):1107–1111. https://doi.org/10.1126/science.1220
Furushita M, Shiba T, Maeda T, Yahata M, Kaneoka A, Takahashi Y, Torii K, Hasegawa T, Ohta M (2003) Similarity of tetracycline resistance genes isolated from fish farm bacteria to those from clinical isolates. Appl Environ Microbiol 69(9):5336–5342. https://doi.org/10.1128/AEM.69.9.5336-5342.2003
Gibbs SG, Green CF, Tarwater PM, Mota LC, Mena KD, Scarpino PV (2006) Isolation of antibiotic-resistant bacteria from the air plume downwind of a swine confined or concentrated animal feeding operation. Environ Health Perspect 114(7):1032–1037. https://doi.org/10.1289/ehp.8910
Gibson Molly K, Forsberg Kevin J, Dantas G (2015) Improved annotation of antibiotic resistance determinants reveals microbial resistomes cluster by ecology. ISME J 9(1):207–216. https://doi.org/10.1038/ismej.2014.106
Gilichinsky DA, Wilson GS, Friedmann EI, McKay CP, Sletten RS, Rivkina EM, Vishnivetskaya TA, Erokhina LG, Ivanushkina NE, Kochkina GA, Shcherbakova VA (2007) Microbial populations in Antarctic permafrost: biodiversity, state, age, and implication for astrobiology. Astrobiology 7(2):275–311. https://doi.org/10.1089/ast.2006.0012
Glaring MA, Vester JK, Lylloff JE, Abu Al-Soud W, Sørensen SJ, Stougaard P (2015) Microbial diversity in a permanently cold and alkaline environment in Greenland. PloS one 10(4):e0124863. https://doi.org/10.1371/journal.pone.0124863
Goñi-Urriza M, Capdepuy M, Arpin C, Raymond N, Caumette P, Quentin C (2000) Impact of an urban effluent on antibiotic resistance of riverine Enterobacteriaceae and Aeromonas spp. Appl Environ Microbiol 66(1):125–132. https://doi.org/10.1128/AEM.66.1.125-132.2000
Gupta GN, Srivastava S, Khare SK, Prakash V (2014) Extremophiles: an overview of microorganism from extreme environment. Int J Agri Environ Biotechnol 7(2):371. https://doi.org/10.5958/2230-732X.2014.00258.7
Hall BG, Barlow M (2004) Evolution of the serine β-lactamases: past, present and future. Drug Resist Updates 7(2):111–123. https://doi.org/10.1016/j.drup.2004.02.003
Hawkey PM (2008) The growing burden of antimicrobial resistance. J Antimicrob Chemother 62(suppl1):i1–9. https://doi.org/10.1093/jac/dkn241
James E, Wong CM (2015) Antibiotic resistance among bacteria from Antarctic and Tropics. Trans Sci Tech 2:16–20. https://doi.org/10.15242/iicbe.c0815032
Junge K, Christner B, Staley JT (2011) Diversity of psychrophilic bacteria from sea ice-and glacial ice communities. Extremophiles Handb. https://doi.org/10.1007/978-4-431-53898-1_39
Kamekura M (1998) Diversity of extremely halophilic bacteria. Extremophiles 2(3):289–295. https://doi.org/10.1007/s007920050071
Kaser G (1999) A review of the modern fluctuations of tropical glaciers. Glob Planet Change 22(1–4):93–103. https://doi.org/10.1016/S0921-8181(99)00028-4
Kashuba E, Dmitriev AA, Kamal SM, Melefors O, Griva G, Römling U, Ernberg I, Kashuba V, Brouchkov A (2017) Ancient permafrost staphylococci carry antibiotic resistance genes. Microb Ecol Health Dis 28(1):1345574. https://doi.org/10.1080/16512235.2017.1345574
Kato C, Li L, Nogi Y, Nakamura Y, Tamaoka J, Horikoshi K (1998) Extremely barophilic bacteria isolated from the Mariana Trench, Challenger Deep, at a depth of 11,000 meters. Appl Environ Microbiol 64(4):1510–1513. https://doi.org/10.1128/AEM.64.4.1510-1513.1998
Kirby-Bauer A (1996) Antimicrobial sensitivity testing by agar diffusion method. J Clin Pathol 44:493
Kishore KH, Begum Z, Pathan AA, Shivaji S (2010) Paenibacillus glacialis sp. nov., isolated from the Kafni glacier of the Himalayas, India. Int J Syst Evol Microbiol 60(8):1909–1913. https://doi.org/10.1099/ijs.0.015271-0
Krumperman PH (1983) Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl Environ Microbiol 46(1):165–170. https://doi.org/10.1128/aem.46.1.165-170.1983
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547. https://doi.org/10.1093/molbev/msy096
Lanoil B, Skidmore M, Priscu JC, Han S, Foo W, Vogel SW, Tulaczyk S, Engelhardt H (2009) Bacteria beneath the West Antarctic ice sheet. Environ Microbiol 11(3):609–615. https://doi.org/10.1111/j.1462-2920.2008.01831.x
Levy SB, Marshall B (2004) Antibacterial resistance worldwide: causes, challenges and responses. Nat Med 10(12):S122–S129. https://doi.org/10.1038/nm1145
Lighthart B, Shaffer BT (1995) Airborne bacteria in the atmospheric surface layer: temporal distribution above a grass seed field. Appl Environ Microbiol 61(4):1492–1496. https://doi.org/10.1128/aem.61.4.1492-1496.1995
Liu Y, Yao T, Jiao N, Kang S, Xu B, Zeng Y, Huang S, Liu X (2009) Bacterial diversity in the snow over Tibetan Plateau Glaciers. Extremophiles 13(3):411–423. https://doi.org/10.1007/s00792-009-0227-5
Lo Giudice A, Bruni V, Michaud L (2007) Characterization of Antarctic psychrotrophic bacteria with antibacterial activities against terrestrial microorganisms. J Basic Microbiol 47(6):496–505. https://doi.org/10.1002/jobm.200700227
Lo Giudice A, Casella P, Bruni V, Michaud L (2013) Response of bacterial isolates from Antarctic shallow sediments towards heavy metals, antibiotics and polychlorinated biphenyls. Ecotoxicology 22(2):240–250. https://doi.org/10.1007/s10646-012-1020-2
Lutz S, Anesio AM, Edwards A, Benning LG (2015) Microbial diversity on Icelandic glaciers and ice caps. Front Microbiol 2015;6:307. https://doi.org/10.3389/fmicb.2015.00307
MacElroy RD (1974) Some comments on the evolution of extremophiles. BioSystems 6(1):74–75. https://doi.org/10.1016/0303-2647(74)90026-4
Makowska N, Zawierucha K, Nadobna P, Piątek-Bajan K, Krajewska A, Szwedyk J, Iwasieczko P, Mokracka J, Koczura R (2020) Occurrence of integrons and antibiotic resistance genes in cryoconite and ice of Svalbard, Greenland, and the Caucasus glaciers. Sci Total Environ 716:137022. https://doi.org/10.1016/j.scitotenv.2020.137022
Malik A, Çelik EK, Bohn C, Böckelmann U, Knobel K, Grohmann E (2008) Detection of conjugative plasmids and antibiotic resistance genes in anthropogenic soils from Germany and India. FEMS Microbiol Lett 279(2):207–216. https://doi.org/10.1111/j.1574-6968.2007.01030.x
Männistö MK, Häggblom MM (2006) Characterization of psychrotolerant heterotrophic bacteria from finnish Lapland. Sys Appl Microbiol 29(3):229–243. https://doi.org/10.1016/j.syapm.2005.09.001
Margesin R, Miteva V (2011) Diversity and ecology of psychrophilic microorganisms. Res Microbiol 162(3):346–361. https://doi.org/10.1016/j.resmic.2010.12.004
Martínez JL (2008) Antibiotics and antibiotic resistance genes in natural environments. Sci 321(5887):365–367. https://doi.org/10.1126/science.1159483
McCann CM, Christgen B, Roberts JA, Su JQ, Arnold KE, Gray ND, Zhu YG, Graham DW (2019) Understanding drivers of antibiotic resistance genes in high Arctic soil ecosystems. Environ Int 125:497–504. https://doi.org/10.1016/j.envint.2019.01.034
Miller RV, Gammon K, Day MJ (2009) Antibiotic resistance among bacteria isolated from seawater and penguin fecal samples collected near Palmer Station, Antarctica. Can J Microbiol 55(1):37–45. https://doi.org/10.1139/W08-119
Montross SN, Skidmore M, Tranter M, Kivimäki AL, Parkes RJ (2013) A microbial driver of chemical weathering in glaciated systems. Geology 41(2):215–218. https://doi.org/10.1130/G33572.1
Morita RY (1975) Psychrophilic bacteria. Bacteriol Rev 39(2):144–167. https://doi.org/10.1128/br.39.2.144-167.1975
Mueller DR, Pollard WH (2004) Gradient analysis of cryoconite ecosystems from two polar glaciers. Polar Biol 27(2):66–74. https://doi.org/10.1007/s00300-003-0580-2
Musilova M, Tranter M, Bennett SA, Wadham J, Anesio AM (2015) Stable microbial community composition on the Greenland ice sheet. Front Microbiol 6:193. https://doi.org/10.3389/fmicb.2015.00193
Nakamura Y, Asada C, Sawada T (2003) Production of antibacterial violet pigment by psychrotropic bacterium RT102 strain. Biotechnol Bioprocess Eng 8(1):37–40. https://doi.org/10.1007/BF02932896
Parnell J, McMahon S (2016) Physical and chemical controls on habitats for life in the deep subsurface beneath continents and ice. Philos Trans R Soc A 374(2059):20140293. https://doi.org/10.1098/rsta.2014.0293
Perron GG, Whyte L, Turnbaugh PJ, Goordial J, Hanage WP, Dantas G, Desai MM (2015) Functional characterization of bacteria isolated from ancient arctic soil exposes diverse resistance mechanisms to modern antibiotics. PloS one 10(3):e0069533. https://doi.org/10.1371/journal.pone.0069533
Rafiq M, Hayat M, Anesio AM, Jamil SU, Hassan N, Shah AA, Hasan F (2017) Recovery of metallo-tolerant and antibiotic resistant psychrophilic bacteria from Siachen glacier. Pakistan PloS one 12(7):e0178180. https://doi.org/10.1371/journal.pone.0178180
Rafiq M, Hayat M, Hassan N, Ibrar M, Haleem A, Rehman M, Ahmad F, Shah AA, Hasan F (2016) Characterization of antibacterial compounds produced by psychrotrophic Alcaligenes faecalis HTP6 isolated from Passu Glacier, Pakistan. Int J Biosci 8(5):122–135. https://doi.org/10.12692/ijb/8.5.122-135
Rafiq M, Hayat M, Zada S, Sajjad W, Hassan N, Hasan F (2019) Geochemistry and bacterial recovery from Hindu Kush Range glacier and their potential for metal resistance and antibiotic production. Geomicrobiol J 36(4):326–338. https://doi.org/10.1080/01490451.2018.1551947
Reddy GS, Prabagaran SR, Shivaji S (2008) Leifsonia pindariensis sp. nov., isolated from the Pindari glacier of the indian Himalayas, and emended description of the genus Leifsonia. Int J Syst Evol Microbiol 58(9):2229–2234. https://doi.org/10.1099/ijs.0.65715-0
Reddy GS, Pradhan S, Manorama R, Shivaji S (2010) Cryobacterium Pindariense sp. nov., a psychrophilic bacterium from a himalayan glacier. Int J Syst Evol Microbiol 60:866–870. https://doi.org/10.1099/ijs.0.011775-0
Rothschild LJ, Mancinelli RL (2001) Life in extreme environments. Nature 409(6823):1092–1101. https://doi.org/10.1038/35059215
Russell AD (2003) Biocide use and antibiotic resistance: the relevance of laboratory findings to clinical and environmental situations. Lancet Infect Dis 3(12):794–803. https://doi.org/10.1016/S1473-3099(03)00833-8
Sakamoto T, Murata N (2002) Regulation of the desaturation of fatty acids and its role in tolerance to cold and salt stress. Curr Opi Microbiol 5(2):206–210. https://doi.org/10.1016/S1369-5274(02)00306-5
Segawa T, Takeuchi N, Rivera A, Yamada A, Yoshimura Y, Barcaza G, Shinbori K, Motoyama H, Kohshima S, Ushida K (2013) Distribution of antibiotic resistance genes in glacier environments. Environ Microbiol Rep 5(1):127–134. https://doi.org/10.1111/1758-2229.12011
Segawa T, Takeuchi N, Ushida K, Kanda H, Kohshima S (2010) Altitudinal changes in a bacterial community on Gulkana Glacier in Alaska. Microb Environ 25(3):171–182. https://doi.org/10.1264/jsme2.ME10119
Shen JP, Li ZM, Hu HW, Zeng J, Zhang LM, Du S, He JZ (2019) Distribution and succession feature of antibiotic resistance genes along a soil development chronosequence in Urumqi No. 1 Glacier of China. Front Microbiol 1569. https://doi.org/10.3389/fmicb.2019.01569
Shen L, Yao T, Xu B, Wang H, Jiao N, Kang S, Liu X, Liu Y (2012) Variation of culturable bacteria along depth in the East Rongbuk ice core. Mt Everest Geosci Front 3(3):327–334. https://doi.org/10.1016/j.gsf.2011.12.013
Sherpa MT, Najar IN, Das S, Thakur N (2020) Distribution of antibiotic and metal resistance genes in two glaciers of North Sikkim, India. Ecotoxicol Environ Saf 203:111037. https://doi.org/10.1016/j.ecoenv.2020.111037
Shivaji S, Chaturvedi P, Reddy GS, Suresh K (2005) Pedobacter himalayensis sp. nov., from the Hamta glacier located in the himalayan mountain ranges of India. Int J Syst Evo Microbiol 55(3):1083–1088. https://doi.org/10.1099/ijs.0.63532-0
Shivaji S, Madhu S, Singh S (2011) Extracellular synthesis of antibacterial silver nanoparticles using psychrophilic bacteria. Process Biochem 46(9):1800–1807. https://doi.org/10.1016/j.procbio.2011.06.008
Stibal M, Tranter M, Benning LG, Řehák J (2008) Microbial primary production on an Arctic glacier is insignificant in comparison with allochthonous organic carbon input. Environ Microbiol 10(8):2172–2178. https://doi.org/10.1111/j.1462-2920.2008.01620.x
Takeuchi N, Kohshima S (2004) A snow algal community on Tyndall Glacier in the Southern Patagonia Icefield, Chile. Arct Antarct Alp Res 36(1):92–99. https://doi.org/10.1657/1523-0430.2004.036.0092:ASACOT.2.0.CO;2
Takeuchi N, Uetake J, Fujita K, Aizen VB, Nikitin SD (2006) A snow algal community on Akkem Glacier in the russian Altai Mountains. Ann Glaciol 43:378–384. https://doi.org/10.3189/172756406781812113
Tomova I, Stoilova-Disheva M, Lazarkevich I, Vasileva-Tonkova E (2015) Antimicrobial activity and resistance to heavy metals and antibiotics of heterotrophic bacteria isolated from sediment and soil samples collected from two Antarctic islands. Front Life Sci 8(4):348–357. https://doi.org/10.1080/21553769.2015.1044130
Ushida K, Segawa T, Kohshima S, Takeuchi N, Fukui K, Li Z, Kanda H (2010) Application of real-time PCR array to the multiple detection of antibiotic resistant genes in glacier ice samples. J Gen Appl Microbiol 56(1):43–52. https://doi.org/10.2323/jgam.56.43
Van Elsas JD, Bailey MJ (2002) The ecology of transfer of mobile genetic elements. FEMS Microbiol Ecol 42(2):187–197. https://doi.org/10.1111/j.1574-6941.2002.tb01008.x
Van Goethem MW, Pierneef R, Bezuidt OK, Van De Peer Y, Cowan DA, Makhalanyane TP (2018) A reservoir of ‘historical’antibiotic resistance genes in remote pristine Antarctic soils. Microbiome 6(1):1–2. https://doi.org/10.1186/s40168-018-0424-5
Whitman WB, Coleman DC, Wiebe WJ (1998) Prokaryotes: the unseen majority. Proc Natl Acad Sci 95(12):6578–6583. https://doi.org/10.1073/pnas.95.12.6578
Wright GD (2010) Antibiotic resistance in the environment: a link to the clinic? Curr Opin Microbiol 13(5):589–594. https://doi.org/10.1016/j.mib.2010.08.005
Zhang S, Yang G, Hou S, Zhang T, Li Z, Liang F (2018) Distribution of ARGs and MGEs among glacial soil, permafrost, and sediment using metagenomic analysis. Environ Pollut 234:339–346. https://doi.org/10.1016/j.envpol.2017.11.031
Zhang DC, Brouchkov A, Griva G, Schinner F, Margesin R (2013) Isolation and characterization of bacteria from ancient siberian permafrost sediment. Biology 2(1):85–106. https://doi.org/10.3390/biology2010085
Zhang S, Yang G, Wang Y, Hou S (2010) Abundance and community of snow bacteria from three glaciers in the Tibetan Plateau. J Environ Sci 22(9):1418–1424. https://doi.org/10.1016/S1001-0742(09)60269-2
Zhang XX, Zhang T, Fang HH (2009) Antibiotic resistance genes in water environment. Appl Microbiol Biotechnol 82(3):397–414. https://doi.org/10.1007/s00253-008-1829-z
Zhang S, Hou S, Ma X, Qin D, Chen T (2007) Culturable bacteria in Himalayan glacial ice in response to atmospheric circulation. Biogeosciences 4(1):1–9. https://doi.org/10.5194/bg-4-1-2007
Acknowledgements
The authors did not receive support from any organization for the submitted work.
Author information
Authors and Affiliations
Contributions
SN, FH, MR and ILP conceived and designed study. SN and WQB performed research. MR, FH and AAS analyzed data. SN and WQB wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
All authors have read and approved the final version.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nawaz, S., Rafiq, M., Pepper, I.L. et al. Prevalence and abundance of antibiotic-resistant genes in culturable bacteria inhabiting a non-polar passu glacier, karakorum mountains range, Pakistan. World J Microbiol Biotechnol 39, 94 (2023). https://doi.org/10.1007/s11274-023-03532-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03532-4