Abstract
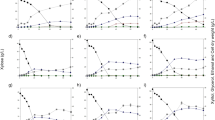
Aiming to broaden the base of knowledge about wild yeasts, four new indigenous strains were isolated from corn residues, and phylogenetic-tree assemblings on ITS and LSU regions indicated they belong to Meyerozyma caribbica. Yeasts were cultivated under full- and micro-aerobiosis, starting with low or high cell-density inoculum, in synthetic medium or corn hydrolysate containing glucose and/or xylose. Cells were able to assimilate both monosaccharides, albeit by different metabolic routes (fermentative or respiratory). They grew faster in glucose, with lag phases ~ 10 h shorter than in xylose. The hexose exhaustion occurred between 24 and 34 h, while xylose was entirely consumed in the last few hours of cultivation (44–48 h). In batch fermentation in synthetic medium with high cell density, under full-aerobiosis, 18–20 g glucose l−1 were exhausted in 4–6 h, with a production of 6.5–7.0 g ethanol l−1. In the xylose medium, cells needed > 12 h to consume the carbohydrate, and instead of ethanol, cells released 4.4–6.4 g l−1 xylitol. Under micro-aerobiosis, yeasts were unable to assimilate xylose, and glucose was more slowly consumed, although the ethanol yield was the theoretical maximum. When inoculated into the hydrolysate, cells needed 4–6 h to deplete glucose, and xylose had a maximum consumption of 57%. Considering that the hydrolysate contained ~ 3 g l−1 acetic acid, it probably has impaired sugar metabolism. Thus, this study increases the fund of knowledge regarding indigenous yeasts and reveals the biotechnological potential of these strains.
Graphical abstract






Similar content being viewed by others
Data availability
Not applicable.
Code availability
Not applicable.
References
Alves SL Jr, Herberts RA, Hollatz C, Miletti LC, Stambuk BU (2007) Maltose and maltotriose active transport and fermentation by Saccharomyces cerevisiae. J Am Soc Brew Chem 65:99–104. https://doi.org/10.1094/ASBCJ-2007-0411-01
Alves SL Jr, Müller C, Bonatto C, Scapini T, Camargo AF, Fongaro G, Treichel H (2019) Bioprospection of enzymes and microorganisms in insects to improve second-generation ethanol production. Ind Biotechnol 15:336–349. https://doi.org/10.1089/ind.2019.0019
Artifon W, Bonatto C, Bordin ER, Bazoti SF, Dervanoski A, Alves SL Jr, Treichel H (2018) Bioethanol production from hydrolyzed lignocellulosic after detoxification via adsorption with activated carbon and dried air stripping. Front Bioeng Biotechnol 6:107. https://doi.org/10.3389/fbioe.2018.00107
Barrilli ÉT, Tadioto V, Milani LM, Deoti JR, Fogolari O, Müller C, Barros KO, Rosa CA, Santos AA, Stambuk BU, Treichel H, Alves SL Jr (2020) Biochemical analysis of cellobiose catabolism in Candida pseudointermedia strains isolated from rotten wood. Arch Microbiol 202:1729–1739. https://doi.org/10.1007/s00203-020-01884-1
Bautista-Rosales PU, Calderon-Santoyo M, Servín-Villegas R, Ochoa-Álvarez NA, Ragazzo-Sánchez JÁ (2013) Action mechanisms of the yeast Meyerozyma caribbica for the control of the phytopathogen Colletotrichum gloeosporioides in mangoes. Biol Control 65:293–301. https://doi.org/10.1016/j.biocontrol.2013.03.010
Bellissimi E, Van Dijken JP, Pronk JT, Van Maris AJA (2009) Effects of acetic acid on the kinetics of xylose fermentation byan engineered, xylose-isomerasebased Saccharomyces cerevisiae strain. FEMS Yeast Res 9:358–364. https://doi.org/10.1111/j.1567-1364.2009.00487.x
Bonatto C, Camargo AF, Scapini T, Stefanski FS, Alves SL, Müller C, Fongaro G, Treichel H (2020a) Biomass to bioenergy research: current and future trends for biofuels. In: Gupta VK, Treichel H, Kuhad RC, Rodriguez-Cout S (eds) Recent developments in bioenergy research, 1st edn. Elsevier, Amsterdam, pp 1–17
Bonatto C, Venturin B, Mayer DA, Bazoti SF, de Oliveira D, Alves SL Jr, Treichel H (2020b) Experimental data and modelling of 2G ethanol production by Wickerhamomyces sp. UFFS-CE-3.1.2. Renew Energ 145:2445–2450. https://doi.org/10.1016/j.renene.2019.08.010
Brito EHS, Fontenelle ROS, Brilhante RSN, Cordeiro RA, Monteiro AJ, Sidrim JJC, Rocha MFG (2009) The anatomical distribution and antimicrobial susceptibility of yeast species isolated from healthy dogs. Vet J 182:320–326. https://doi.org/10.1016/j.tvjl.2008.07.001
Cadete RM, Rosa CA (2018) The yeasts of the genus Spathaspora: potential candidates for second-generation biofuel production. Yeast 35:191–199. https://doi.org/10.1002/yea.3279
Cadete RM, Melo MA, Dussán KJ, Rodrigues RCLB, Silva SS, Zilli JE, Vital MJS, Gomes FCO, Lachance M-A, Rosa CA (2012) Diversity and physiological characterization of D-xylose-fermenting yeasts isolated from the Brazilian Amazonian Forest. PLoS ONE 7:e43135. https://doi.org/10.1371/journal.pone.0043135
Cadete RM, de las Heras AM, Sandström AG, Ferreira C, Gírio F, Gorwa-Grauslund M-F, Rosa CA, Fonseca C (2016) Exploring xylose metabolism in Spathaspora species: XYL1.2 from Spathaspora passalidarum as the key for efficient anaerobic xylose fermentation in metabolic engineered Saccharomyces cerevisiae. Biotechnol Biofuels 9:167. https://doi.org/10.1186/s13068-016-0570-6
Cadete RM, Melo-Cheab MA, Dussan KJ, Rodrigues RCLB, da Silva SS, Gomes FCO, Rosa CA (2017) Production of bioethanol in sugarcane bagasse hemicellulosic hydrolysate by Scheffersomyces parashehatae, Scheffersomyces illinoinensis and Spathaspora arborariae isolated from Brazilian ecosystems. J Appl Microbiol 123:1203–1213. https://doi.org/10.1111/jam.13559
Carneiro CVGC, Silva FCP, Almeida JRM (2019) Xylitol production: identification and comparison of new producing yeasts. Microorganisms 7:484. https://doi.org/10.3390/microorganisms7110484
Casey E, Sedlak M, Ho NWY, Mosier NS (2010) Effect of acetic acid and pH on the cofermentation of glucose and xylose to ethanol by a genetically engineered strain of Saccharomyces cerevisiae. FEMS Yeast Res 10:385–393. https://doi.org/10.1111/j.1567-1364.2010.00623.x
Dall Cortivo PR, Hickert LR, Hector R, Ayub MAZ (2018) Fermentation of oat and soybean hull hydrolysates into ethanol and xylitol by recombinant industrial strains of Saccharomyces cerevisiae under diverse oxygen environments. Ind Crop Prod 113:10–18. https://doi.org/10.1016/j.indcrop.2018.01.010
De Marco L, Epis S, Capone A, Martin E, Bozic J, Crotti E, Ricci I, Sassera D (2018) The genomes of four Meyerozyma caribbica isolates and novel insights into the Meyerozyma guilliermondii species complex. G3 Genes Genom Genet 8:755–759. https://doi.org/10.1534/g3.117.300316
Dionísio SR, Santoro DCJ, Bonan CIDG, Soares LB, Biazi LE, Rabelo SC, Ienczak JL (2021) Second-generation ethanol process for integral use of hemicellulosic and cellulosic hydrolysates from diluted sulfuric acid pretreatment of sugarcane bagasse. Fuel 304:121290. https://doi.org/10.1016/j.fuel.2021.121290
Eliodório KP, Cunha GCG, Müller C, Lucaroni AC, Giudici R, Walker GM, Alves SL Jr, Basso TO (2019) Advances in yeast alcoholic fermentations for the production of bioethanol, beer and wine. In: Gadd GM, Sariaslani S (eds) Advances in applied microbiology, vol 109. Elsevier, Amsterdam, pp 61–119
Ferreira D, Nobre A, Silva ML, Faria-Oliveira F, Tulha J, Ferreira C, Lucas C (2013) XYLH encodes a xylose/H+ symporter from the highly related yeast species Debaryomyces fabryi and Debaryomyces hansenii. FEMS Yeast Res 13:585–596. https://doi.org/10.1111/1567-1364.12061
Gadagkar SR, Rosenberg MS, Kumar S (2005) Inferring species phylogenies from multiple genes: concatenated sequence tree versus consensus gene tree. J Exp Zool 304B:64–74. https://doi.org/10.1002/jez.b.21026
Guamán-Burneo MC, Dussán KJ, Cadete RM, Cheab MAM, Portero P, Carvajal-Barriga EJ, da Silva SS, Rosa CA (2015) Xylitol production by yeasts isolated from rotting wood in the Galápagos Islands, Ecuador, and description of Cyberlindnera galapagoensis f.a., sp. nov. Antonie Van Leeuwenhoek 108:919–931. https://doi.org/10.1007/s10482-015-0546-8
Hahn-Hägerdal B, Karhumaa K, Larsson CU, Gorwa-Grauslund M, Görgens J, van Zyl WH (2005) Role of cultivation media in the development of yeast strains for large scale industrial use. Microb Cell Fact 4:31. https://doi.org/10.1186/1475-2859-4-31
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Jan FG, Hamayun M, Hussain A, Iqbal A, Jan G, Khan SA, Khan H, Lee I-J (2019) A promising growth promoting Meyerozyma caribbica from Solanum xanthocarpum alleviated stress in maize plants. Biosc Rep. https://doi.org/10.1042/BSR20190290
Jönsson LJ, Martín C (2016) Pretreatment of lignocellulose: formation of inhibitory by-products and strategies for minimizing their effects. Bioresour Technol 199:103–112. https://doi.org/10.1016/j.biortech.2015.10.009
Khunnamwong P, Jindamorakot S, Limtong S (2018) Endophytic yeast diversity in leaf tissue of rice, corn and sugarcane cultivated in Thailand assessed by a culture-dependent approach. Fungal Biol 122:785–799. https://doi.org/10.1016/j.funbio.2018.04.006
Kim J-S, Baek J-H, Park N-H, Kim C (2015) Complete genome sequence of halophilic yeast Meyerozyma caribbica MG20W isolated from rhizosphere soil. Genome Announc 3:e00127-e215. https://doi.org/10.1128/genomeA.00127-15
Leong SL, Niba AT, Ny S, Olstorpe M (2012) Microbial populations during maize storage in Cameroon. Afr J Biotechnol 11:8692–8697. https://doi.org/10.5897/AJB12.108
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296. https://doi.org/10.1093/nar/gkab301
Lopes ML, Paulillo SCL, Godoy A, Cherubin RA, Lorenzi MS, Giometti FHC, Bernardino CD, Amorim Neto HB, Amorim HV (2016) Ethanol production in Brazil: a bridge between science and industry. Braz J Microbiol 47:64–76. https://doi.org/10.1016/j.bjm.2016.10.003
Martínez-Nieto P, Vanegas-Hoyos M, Zapata-Pineda M, Robles-Camargo J (2013) Hydrolysis of Eicchornia crassipes and Egeria densa for ethanol production by yeasts isolated from Colombian lake Fúquene. Int J Environ Ecol Eng 7:59–68. https://doi.org/10.5281/zenodo.1088798
Moremi ME, Van Rensburg ELJ, La Grange DC (2020) The improvement of bioethanol production by pentose-fermenting yeasts isolated from herbal preparations, the gut of dung beetles, and marula wine. Int J Microbiol 2020:5670936. https://doi.org/10.1155/2020/5670936
Nagarajan A, Thulasinathan B, Arivalagan P, Alagarsamy A, Muthuramalingam JB, Thangarasu SD, Thangavel K (2021) Particle size influence on the composition of sugars in corncob hemicellulose hydrolysate for xylose fermentation by Meyerozyma caribbica. Bioresour Technol 340:125677. https://doi.org/10.1016/j.biortech.2021.125677
Romi W, Keisam S, Ahmed G, Jeyaram K (2014) Reliable differentiation of Meyerozyma guilliermondii from Meyerozyma caribbica by internal transcribed spacer restriction fingerprinting. BMC Microbiol 14:52. https://doi.org/10.1186/1471-2180-14-52
Saloheimo A, Rauta J, Stasyk OV, Sibirny AA, Penttilä M, Ruohonen L (2007) Xylose transport studies with xylose-utilizing Saccharomyces cerevisiae strains expressing heterologous and homologous permeases. Appl Microbiol Biotechnol 74:1041–1052. https://doi.org/10.1007/s00253-006-0747-1
Santos RO, Cadete RM, Badotti F, Mouro A, Wallheim DO, Gomes FC, Stambuk BU, Lachance MA, Rosa CA (2011) Candida queiroziae sp. nov., a cellobiose-fermenting yeast species isolated from rotting wood in Atlantic rain forest. Antonie Van Leeuwenhoek 99:635–642. https://doi.org/10.1007/s10482-010-9536-z
Santos RM, Nogueira FCS, Brasil AA, Carvalho PC, Leprevost FV, Domont GB, Eleutherio ECA (2017) Quantitative proteomic analysis of the Saccharomyces cerevisiae industrial strains CAT-1 and PE-2. J Proteomics 151:114–121. https://doi.org/10.1016/j.jprot.2016.08.020
Shin M, Kim J, Ye S, Kim S, Jeong D, Lee DY, Kim JN, Jin Y-S, Kim KH, Kim SR (2019) Comparative global metabolite profiling of xylose-fermenting Saccharomyces cerevisiae SR8 and Scheffersomyces stipitis. Appl Microbiol Biotechnol 103:5435–5446. https://doi.org/10.1007/s00253-019-09829-5
Stambuk BU, Eleutherio ECA, Florez-Pardo LM, Souto-Maior AM, Bom EPS (2008) Brazilian potential for biomass ethanol: challenge of using hexose and pentose co-fermenting yeast strains. J Sci Ind Res 67:918–926
Suh S-O, Houseknecht JL, Gujjari P, Zhou JJ (2013) Scheffersomyces parashehatae f.a., sp. nov., Scheffersomyces xylosifermentans f.a., sp. nov., Candida broadrunensis sp. nov. and Candida manassasensis sp. nov., novel yeasts associated with wood-ingesting insects, and their ecological and biofuel implications. Int J Syst Evol Microbiol 63:4330–4339. https://doi.org/10.1099/ijs.0.053009-0
Sukpipat W, Komeda H, Prasertsan P, Asano Y (2017) Purification and characterization of xylitol dehydrogenase with L-arabitol dehydrogenase activity from the newly isolated pentose-fermenting yeast Meyerozyma caribbica 5XY2. J Biosci Bioeng 123:20–27. https://doi.org/10.1016/j.jbiosc.2016.07.011
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Thanh VN, Van Dyk MS, Wingfield MJ (2002) Debaryomyces mycophilus sp. nov., a siderophore-dependent yeast isolated from woodlice. FEMS Yeast Res 2:415–427. https://doi.org/10.1016/S1567-1356(02)00132-0
Trichez D, Steindorff AS, Soares CEVF, Formighieri EF, Almeida JRM (2019) Physiological and comparative genomic analysis of new isolated yeasts Spathaspora sp. JA1 and Meyerozyma caribbica JA9 reveal insights into xylitol production. FEMS Yeast Res 19:foz034. https://doi.org/10.1093/femsyr/foz034
Urbina H, Blackwell M (2012) Multilocus phylogenetic study of the Scheffersomyces yeast clade and characterization of the N-terminal region of xylose |reductase gene. PLoS ONE 7:e39128. https://doi.org/10.1371/journal.pone.0039128
Urbina H, Schuster J, Blackwell M (2013) The gut of Guatemalan passalid beetles: a habitat colonized by cellobiose- and xylose-fermenting yeasts. Fungal Ecol 6:339–355. https://doi.org/10.1016/j.funeco.2013.06.005
Varize CS, Cadete RM, Lopes LD, Christofoleti-Furlan RM, Lachance M-A, Rosa CA, Basso LC (2018) Spathaspora piracicabensis f. a., sp. nov., a D-xylose- fermenting yeast species isolated from rotting wood in Brazil. Antonie Van Leeuwenhoek 111:525–531. https://doi.org/10.1007/s10482-017-0974-8
Vaz ABM, Rosa LH, Vieira MLA, de Garcia V, Brandão LR, Teixeira LCRS, Moliné M, Libkind D, van Broock M, Rosa CA (2011) The diversity, extracellular enzymatic activities and photoprotective compounds of yeasts isolated in Antarctica. Braz J Microbiol 42:937–947. https://doi.org/10.1590/S1517-83822011000300012
Weusthuis RA, Visser W, Pronk JT, Scheffers WA, van Dijken JP (1994) Effects of oxygen limitation on sugar metabolism in yeasts: a continuous-culture study of the Kluyver effect. Microbiology 140:703–715. https://doi.org/10.1099/00221287-140-4-703
White TJ, Bruns TD, Lee SB, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand DH, Shinsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yurkov AM, Dlauchy D, Péter G (2017) Meyerozyma amylolytica sp. nov. from temperate deciduous trees and the transfer of five Candida species to the genus Meyerozyma. Int J Syst Evol Microbiol 67:3977–3981. https://doi.org/10.1099/ijsem.0.002232
Zaky AS, French CE, Tucker GA, Du C (2020) Improving the productivity of bioethanol production using marine yeast and seawater-based media. Biomass Bioenerg 139:105615. https://doi.org/10.1016/j.biombioe.2020.105615
Acknowledgements
This work was supported in part by grants and fellowships from the Brazilian National Council for Scientific and Technological Development—CNPq (Grant Numbers 454215/2014-2, 303735/2018-0, 305258/2018-4 and 308389/2019-0), the Santa Catarina State Research and Innovation Support Foundation—FAPESC (Grant Number 749/2016; T.O. 2016TR2188), and the Research Promotion Program from Federal University of Fronteira Sul—UFFS (Grant Numbers PES-2019-0638 and PES-2020-0213). VT, LMM, ETB, LB, and AD are grateful to the Support Program for Scientific and Technological Initiation from Federal University of Fronteira Sul (PRO-ICT/UFFS), and CM to the Coordination for the Improvement of Higher Education Personnel (CAPES/PNPD, No. 88887.352933/2019-00), for scholarships.
Author information
Authors and Affiliations
Contributions
VT, LMM, EVT and CWB performed the yeast isolation, the cellular growths and the batch fermentations. LB and AD performed the pretreatment and the hydrolysis of corn biomass. RH sequenced the ITS region of the strains. OF carried out the HPLC analysis. CM analyzed the data and contributed new methods. GMM, JPB, HT, BUS and SLAJ participated in designing the study and provided financial support. SLAJ also assembled the phylogenetic trees and wrote the manuscript, which was revised and approved by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
The authors declare that this study is in accordance with the ethical standards of our institutions, and did not require the approval of any ethics committee in our country to be carried out.
Informed consent
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tadioto, V., Milani, L.M., Barrilli, É.T. et al. Analysis of glucose and xylose metabolism in new indigenous Meyerozyma caribbica strains isolated from corn residues. World J Microbiol Biotechnol 38, 35 (2022). https://doi.org/10.1007/s11274-021-03221-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-021-03221-0